Abstract
While most of our tissues appear static, in fact, cell motion comprises an important facet of all life forms, whether in single or multicellular organisms. Amoeboid cells navigate their environment seeking nutrients, whereas collectively, streams of cells move past and through evolving tissue in the development of complex organisms. Cell motion is powered by dynamic changes in the structural proteins (actin) that make up the cytoskeleton, and regulated by a circuit of signaling proteins (GTPases) that control the cytoskeleton growth, disassembly, and active contraction. Interesting mathematical questions we have explored include (1) How do GTPases spontaneously redistribute inside a cell? How does this determine the emergent polarization and directed motion of a cell? (2) How does feedback between actin and these regulatory proteins create dynamic spatial patterns (such as waves) in the cell? (3) How do properties of single cells scale up to cell populations and multicellular tissues given interactions (adhesive, mechanical) between cells? Here I survey mathematical models studied in my group to address such questions. We use reaction-diffusion systems to model GTPase spatiotemporal phenomena in both detailed and toy models (for analytic clarity). We simulate single and multiple cells to visualize model predictions and study emergent patterns of behavior. Finally, we work with experimental biologists to address data-driven questions about specific cell types and conditions.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
1 Introduction: Motile Cells and Their Inner Workings
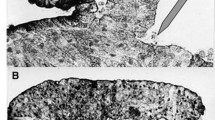
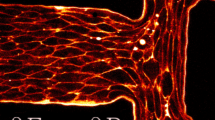
Many types of cells are endowed with the ability to move purposefully. As an example, neutrophils, shown in Fig. 1a, are white blood cells that make up part of our immune system, in charge of patrolling tissues for pathogens or sites of injury. The motion of unicellular organisms such as bacteria, while interesting in its own right, is governed by distinct mechanisms that will not be discussed here.
Cell motility and cell polarization: from biology to mathematical model: a A white blood cell (neutrophil) moving between red blood cells (disk-shaped objects) from a 1950s movie clip by David Rogers. The 1D band represents a transect of the cell from front to back. We are concerned with how the cell breaks symmetry and polarizes to define such a front-back axis. b, c Sketch of a cell in top-down b and side c views, indicating the same 1D axis. d In our mathematical model, we aim to explain how regulatory proteins in the cell (called GTPases) spontaneously polarize and form hot spots of activity that define the front and back of the cell. e In our abstract “wave-pinning” model, this same process is depicted as a 1D pattern-formation event, with a wave that stalls to produced a polarized distribution
In a movie dating to the 1950s’ David Rogers (then at Vanderbilt University) captured the amoeboid movements of a neutrophil as it navigates between red blood cells (disk shaped objects in Fig. 1a). In this movie, which can be seen on a popular YouTube site, we see a crawling cell, with dynamic shape—a broad front that pushes outwards, and a thin tail that is pulled along as the cell moves. Figure 1b, c are two projections of cell shape (top down in (b) and side view in (c)) that we later utilize in modeling cell polarization.
It is worth pointing out the sizes and timescales that concern us here. In contrast to some papers (e.g. Prof. Marsha Berger’s whose work describes geological size scales and timescales of hours and days [1]), here we deal with the micro-world of cells, whose diameter is on the order of 10–30 \(\upmu \)m. The time-scale of relevance is on the order of seconds. As summarized in Table 1, the process of cell polarization, which defines the front and back of the cell and specifies its direction of motion, take place over seconds across the tiny cell diameter. Also noteworthy is the fact that the production of new copies of proteins (i.e. protein synthesis) does not suffice to explain how protein activity becomes concentrated at some parts of a cell, since synthesis takes hour(s), while the response times of a cell to stimuli that polarize it is known to take only seconds for fast-moving cells like neutrophils.
Here the purpose is to explain an important first step in cell motility: the symmetry breaking that creates a front and a back in the cell (Fig. 1d), namely the polarization of the cell. But before embarking on the mathematics that describes this process, we first discuss the important cellular components that are involved.
1.1 Actin Powers Cell Motility
Unlike plants and bacteria, animal cells have no tough outer cell wall. They are enclosed in a lipid membrane that envelopes the interior, which in turn includes the fluid cytosol and many organelles. Most organelles, including the cell’s nucleus are not directly involved in powering cellular motion.
Without some structural components, the cell would be essentially a bag of fluids. An internal “skeleton” (called the cytoskeleton) is formed by a meshwork of filamentous actin (F-actin), a dynamic biopolymer protein structure that is assembled at what becomes the cell front. The polymerization of actin leads to protrusion of the cell front [23]. Meanwhile, in association with the motor protein myosin, contraction of actomyosin leads to retraction of the rear portion of the cell [33], Fig. 2a.
Due to the abundance of actin monomers at excess concentration in every cell, actin assembly would be an explosive process were it not tightly controlled by many interacting regulatory cellular proteins. Many of those proteins, discovered and characterized experimentally over the last decades [27, 34], interact with actin to make it branch, to cut or cap its growing ends, to sequester or to recycle its monomeric subunits. Other proteins play the role of master-regulators that control the components of the cytoskeleton [30].
1.2 GTPases Are Master Regulators
One important class of proteins that regulate the cytoskeleton is the class of Rho GTPases, among which Rac and Rho are well known [3]. In the schematic Fig. 2, GTPases are shown to promote the assembly of filamentous actin, and the activity of myosin contraction. The GTPase Rac does the former, while the GTPase Rho enables the latter. Hence, if we can explain how Rac and Rho activities concentrate at one or another part of the cell, we can also explain the localizations of a front and rear cellular axis, and hence cell polarization. This then, is the main focus of our approach.
Schematic diagram of the cell’s motility machinery: a Actin filaments (F-actin), represented as blue curves, assemble at what becomes the cell front. Actin polymerization leads to protrusion at the front edge of the cell. In the cell rear, myosin motors (not shown) associate with F-actin to contract and pull up the “tail”. Proteins in the class known as Rho GTPases are master regulators. These proteins control where and when actin assembly and myosin contraction take place. GTPases play an essential role in cell polarization. b Each GTPase has an active and an inactive state, modeled by the variables u, v. Only when bound to the cell membrane (shown in yellow) is the GTPase active. A, I denote rates of activation and inactivation
Interestingly, proteins in the family of Rho GTPases have a curious life-cycle. They occur in active and inactive forms, with only the active forms exerting the effects mentioned above [8]. Moreover, the active forms are always bound to the fatty membrane that forms the outer cell envelop (shown in yellow in Fig. 2). Hence, the small GTPases spend their cellular lives shuttling between the cell membrane (where part of their structure gets embedded when active) and the cell interior (where they are entirely inactive). This basic idea is illustrated in Fig. 2b. The GTPases act as cellular switches that are “ON” when active and “OFF” otherwise.
A natural question one could ask, is what is the functional purpose of the GTPase cycling between the cell’s membrane and the cell’s interior? As we shall see, mathematics may have something to contribute towards answering such questions. A second question is what property of the cellular machinery account for the spontaneous polarization of the cell? That is, how do GTPases redistribute so that their levels of activity differ between the front and rear of a cell [2].
2 Mathematical Models
In our earliest works on cell polarization, we attempted to account for many known features of the GTPase activity and their crosstalk and interactions [6, 18, 20]. Such models were largely computational, as it was a challenge to analyse them mathematically. It was clear that more basic model variants would be useful for mathematical progress to be feasible.
As described in Mori et al. [24, 25], we simplify a very complicated cellular process to allow for mathematical tractability. We thereby hope to identify key elements that allow for spontaneous cell polarization. First, we consider just one GTPase (say Rac), rather than the entire network (Cdc42, Rac and Rho). We ask which biological attributes account for spontaneous symmetry breaking and polar pattern formation. To investigate this, we construct the following mathematical model.
We define u(t), v(t) to be the concentrations of the active and inactive forms of the GTPase. Then, based on the schematic diagram in Fig. 2b, it follows that
This is not yet enough, since spatial distribution is a vital aspect. Hence, we require a spatial variable, and need to account for the localization of each of u, v. To do so, we also need to define the geometry of interest.
As argued earlier, and noted in Fig. 1, to explain symmetry breaking for polarization, a 1D model along the front-back axis suffices. And while the detailed residence of the proteins on the membrane or cell interior is important, it proves helpful to simplify this too, in the steps shown in Fig. 3. In that figure, we first idealize the cell as a thin sheet of uniform thickness, surrounded top and bottom by a membrane (yellow outline). Zooming in on a small portion of the cell, we might see active (red) and inactive (black) copies of the GTPase associated with the membrane or the fluid cell interior. We homogenize these compartments, treating both u and v as dependent variables on a 1D spatial domain \(0 \le x \le L\) where L is the cell diameter. We do however, take into account the very different rates of diffusion of a protein in the membrane (\(D_u \approx 0.01{-} 0.1\, \upmu \text {m}^2/s\)) versus the fluid cell interior (\(D_v\approx 10\, \upmu \text {m}^2/s\)) [28]. As we shall see, this huge disparity in diffusion plays a significant role.
The model becomes
In principle, the rates of activation and inactivation A, I, are not merely constant. If they were, then Eq.(1) would be linear in u, v, and would have fairly uninteresting steady state solutions. Some nonlinearity is essential, and this also requires feedback—something that can only depend on levels of active proteins. (Recall that the inactive GTPases do not participate in any interactions.) We have considered models where many other proteins influence each of the state transitions [14, 18, 21], and in that case, the model would expand in complexity,
Such examples, considered in the context of biological experiments, are briefly discussed further on, but mathematically, they are harder to analyze.
Our ultimate purpose, mathematically, is to strip away such complexity and focus on the most elementary example, where a single GTPase polarizes on its own. To do so, we considered the version
with feedback exclusively in the activation rate A(u) and a constant rate of inactivation I. This specific choice is somewhat arbitrary, as shown in [18], since it is possible to obtain essentially the same behaviour with nonlinearity introduced by assuming that \(I=I(u)\) with A constant, or by other variants where both A and I depend on u. The biological interpretation is somewhat different, since distinct proteins in cells play the role of activating (GEFs) and inactivating (GAPS) the GTPases. In the case of constant I, we can rescale time, so that \(I=1\). Altogether, then, the single-GTPase system consists of the pair of PDEs
with
where b is the basal rate of activation and \(\gamma \) is an additional rate of activation depicting positive feedback from u to its own activation. The constant \(n \ge 2\) is the so-called “Hill coefficient”. Larger values of n result in sharper switching between states.
We also assume Neumann boundary conditions, namely,
This signifies that no material leaks out of the ends of the 1D domain, i.e. that the cell ends are sealed.
Notably, on the timescale of interest (a few seconds), no protein is made or lost, it is merely exchanged between the active and inactive states (see Table 1). This is captured by the model, since it is easy to see that the total amount of protein in the domain is conserved, that is,
As shown in [24, 25], the following properties are necessary and sufficient to ensure that a unimodal pattern (depicting a polarized distribution) will exist as a nonuniform steady state of the model:
-
1.
There is some range of values \(v_1 \le v \le v_2\) for which the function f(u, v) has three roots, \(u_a< u_m < u_b\). (We refer to this range of v as the bistable regime.)
-
2.
Of these three roots, the outer two (\(u_a, u_b\)) are stable fixed points of the spatially homogeneous variant of (4).
-
3.
For some value, \(v^*\) in \(v_1 \le v \le v_2\), there is a change in the sign of the integral
$$ \int \limits _{u_a}^{u_b} f(u,v) du. $$ -
4.
The rates of diffusion of u and v are sufficiently different: \(D_u \ll D_v\).
It is interesting to contrast the system (4) with a related one consisting of (4a), (4c) and (4d) but with \(v\equiv \) constant, that is, with a single bistable reaction-diffusion equation in one variable, u. The latter is known to sustain traveling wave solutions, as shown in Fig. 4a. In contrast, the two-variable system (4a)–(4d) leads to waves that decelerate and stop inside the domain (once the sign condition above is satisfied) as demonstrated in Fig. 4b. We refer to this behaviour as “wave-pinning”. We see that Fig. 4a fails to explain polarization, because the cell diameter is eventually uniformly active. Figure 4b is consistent with polarization, since the two ends of the domain develop distinct levels of activity as time goes by. In this sense, wave-pinning is a simple caricature of cell polarization.
Travelling waves versus wave-pinning: a A single reaction-diffusion equation (4a) (for constant v) with kinetics of type (4c) is known to sustain traveling wave solutions for u(x, t). b In contrast, the system of Eqs. (4a)–(4d) with conservation and distinct rates of diffusion (\(D_u \ll D_v\)) results in waves that stop inside the domain, a phenomenon we termed “wave-pinning”
2.1 How Wave-Pinning Works
Full details of the analysis of such dynamics are described in [25]. Here it suffices to briefly mention the key asymptotic analysis ideas used in establishing the result.
The system (4) is rescaled to exploit the existence of a small parameter
where r is a typical kinetics rate constant with units of 1/time (e.g., \(r=\gamma \)). We then examine the short and intermediate time-scales of the rescaled system.
On a short time-scale (\(t_s = t/\epsilon \)), it can be shown that to leading order, at various sites in the domain, u approaches its steady state values \(u_a, u_b\). This means that the domain is “carved up” into plateaus of high and of low activity levels u separated by transition layers between them.
To make progress, we consider the case of a single interface separating a low and a high plateau. Let the position of the interface be \(\phi (t)\). We go on to seek the intermediate time scale behaviour. We construct an inner and an outer solution next to the transition layer and show that, to leading order, the variable v is roughly spatially constant on the two sides of the interface \(v \approx V_0(t)\), while it is depleted in time as u evolves.
Using well-known analysis for wave-speed, we construct the speed of the wave, finding it to be described by a ratio of two integrals
Here \(u_a, u_b\) depend on \(V_0(t)\), and \(I_2\) is a strictly positive integral. We argue that the wave stops when the numerator vanishes, which is guaranteed to happen at some point by Condition 3, a Maxwell condition. Indeed, once v is depleted sufficiently, to the level \(v^*\), the integral in the numerator vanishes. Details and discussion of the steps appear in [25]. Regimes of polarization are shown in (Fig. 5).
Regimes of wave-pinning: Wave-pinning, which represents cell polarization, depends on a balance between the total amount of GTPase (5) and the size of the small parameter \(\epsilon = D_u/ (r L^2)\). If the total amount is too small, the wave of activity collapses, whereas if it is too large, the wave sweeps across the entire domain, and a net homogenous state results. Polarization can also be lost in several ways (1) If the cell size decreases too much, and hence \(\epsilon \) increases, the system leaves the polarization regime. (2) If cell size increases so that the mean total GTPase becomes too “diluted”, polarization can also be lost. Image credit: Alexandra JIlkine
Intuitively, the result can be explained as follows: at the transition zone, the high u plateau activates an adjoining site by virtue of local diffusion and positive feedback. The spread of u, however, is at the expense of the inactive form v, which gets depleted as the wave of activity spreads. Once v is sufficiently depleted, the spread of the activity wave can no longer be sustained. At that point, the wave freezes.
It is also interesting to note that the fast diffusion of v means that it acts as a “global messenger” in the sense that it rapidly stores domain-wide information about the level of activity in the cell. Hence, local activation (of u by itself) and global depletion (of v) synergize to produce the polarization of activity in the domain.
3 Recent Work: Analysis, Simulation, and Contact with Experiments
The wave-pinning equations are merely a prototype of the dynamics of a protein in the small GTPase family. Related systems with greater levels of biological detail have also been explored [12, 14, 21]. Indeed insights by AFM Marée in [20] contributed to the understanding that led to the mathematical treatment of wave-pinning in [24, 25].
3.1 Analysis of Slow-Fast Reaction Diffusion Systems: LPA
While studying systems of reaction-diffusion equations (RDEs) for cell polarization, we have benefitted from a number of recent methods that result in shortcuts for quick diagnosis of pattern-formation regimes. Among these, the “Local Perturbation Analysis” (LPA) is a method to track local and global variables in RDEs using ODEs that approximate the fate of a small peak of activity (\(u_L\)). This method was invented by AFM Mareé and V Grieneisen [9, 36], and popularized in several papers [11, 12, 15]. It has helped us to identify approximate regimes where a nonuniform pattern could form by a finite perturbation of a spatially uniform state in a fast-slow reaction diffusion system.
Methods of analysis and simulations: a Local perturbation analysis (LPA), a shortcut bifurcation method has helped to detect regimes of patterning in slow-fast reaction-diffusion systems. Here we show an example of how the basal activation rate b influences potential regimes of wave-pinning and of Turing-type instability. See text and references [11, 12, 15] for details. b A number of methods have been used to simulate polarization in 2D deforming domains representing the “top-down” view of a cell (as in Fig. 1b). From top to bottom: A cellular-Potts model simulation by A. F. M. Mareé of a 2D deforming cell with an internal reaction-diffusion signaling circuit (and an implicit reaction-diffusion solver) that includes GTPases, interacting lipids, actin, and other components [21], the wave-pinning system (4) solved in an immersed-boundary method simulation by Ben Vanderlei [35], by the level set and moving boundary node method by Zajac [7], and using CompuCell3D by undergraduate summer research student Zachary Pellegrin
Figure 6 illustrates a typical LPA bifurcation result, and its interpretation. The method identifies the existence of a spatially uniform global branch (in black), and parameter regimes where this branch is stable (solid) or unstable (dot-dashed curve). Even when the global homogeneous steady state is stable, a polarized pattern can be established with large enough stimulus. The local variable \(u_L\) represents a thin local peak of active u. That peak could grow (and lead to a polar pattern) in the regime where the solid red curve is present. The LPA diagram demonstrates that a sufficiently large stimulus peak is needed, that its size has to exceed a threshold (dashed red curve), and that some parameter regimes allow for patterning in response to arbitrarily small stimuli (dot-dashed black curve). The latter regimes can be identified with Turing instabilities. The former regimes are not discoverable by the usual linear stability analysis (LSA) for Turing pattern formation, and are a helpful aspect of LPA that goes beyond LSA.
In our experience, solving the full PDEs with insights gained from LPA diagrams makes it easier to identify the interesting parameter regimes. Details of the method and its uses has been extensively described in [15]. Other useful shortcuts have included “sharp-switch” approximations (Hill functions replaced by piecewise constant functions), as in [12], and analysis of plateaus described in [36]. None of these replace the need for simulating the PDEs, but all of them help to gain familiarity with possible expected behaviours of the reaction-diffusion systems we have investigated. Most recently, Andreas Buttenschön has created full numerical bifurcation software for PDEs that permits much greater accuracy in tracking solution branches [4]. The software builds on state of the art well-conditioned collocation techniques to discretize functions and their operators. Solution branches are continued using a matrix-free Newton-Gauss method, for which rigorous convergence estimates are available.
3.2 Simulating the PDEs in Dynamic Cell-Shaped Domains
So far, analytic results were described in 1D domains that represent a cell transect. It is instructive to ask how the same systems behave in domains whose shape more closely relates to that of cells, and in particular, where the internal chemistry affects (and is affected by) the deforming cell. Based on the fact that cell fragments (radius \({\approx }\) 5–10 \(\mu \)m) without a nucleus, and with overall uniform thickness (\({\approx } 0.2\, \upmu \)m) are capable of motility, we take the liberty of reducing cell shape to its two-dimensional “top-down” projection shown in Fig. 1b, d. We solve the governing equations (4) or more detailed versions, in the 2D domain, and assume that the boundary of the domain is influenced by the local chemical activity level. For example, if u represents the level of activity of the GTPase Rac, it causes the boundary to be pushed outwards (via F-actin assembly), whereas Rho has the opposite effect (activating contraction via myosin).
A number of results obtained over the years by group members are illustrated in Fig. 6b. In general, we found that the simplest system to understand analytically (4), is not as robust computationally as other variants. Cross-talk between GTPases results in larger parameter regimes for polarization. As an example, models consisting of four PDEs that describe the mutual antagonism between Rac and Rho [12] lead to greater robustness in 2D computations. An even more detailed variant, that includes several GTPases (Rac, Rho, Cdc42), as well as their effects on actin assembly and myosin contraction was capable of realistic behaviour such as directed motility (chemotaxis) [20]. The addition of a layer of signaling lipids (phosphoinositides) also permitted a simulated cell to rapidly select one front despite conflicting or competing stimuli [21].
Simulating the reaction-diffusion systems for GTPase signaling in deforming domains also reveals that evolving domain shape and level curves of the chemical system influence one another: the zero-flux boundary conditions impose constraints on the level curves that also accelerate the dynamics of the chemical redistribution when the domain deforms. Such findings were discussed in detail in [21].
For practical reasons, it is harder to simulate the same systems in 3D. However, recent work by the group of Anotida Madzvamuse [5] has extended these results to a coupled bulk-surface wave-pinning computation in a 3D cell-shaped static domain.
3.3 Contact with Biological Experiments
While details are beyond the scope of this summary, it is worth noting several directions in which the mathematical modeling has contributed to understanding of experimental cell biology.
Willian Bement (U Wisconsin) studies the patterns of GTPases (Rho and Cdc42) that form spontaneously around sites of laser-inflicted wounds in frog eggs (Xenopus oocytes). The connectivity of these GTPases, and their crosstalk with proteins that activate or inactivate them (e.g. Abr) has been modeled by group members, including Cory Simon, Laura Liao, and William R Holmes. Combining models with experiments has helped to build an understanding of the biology [12, 13, 32].
The polarization of HeLa cells exposed to gradients that stimulate a graded response by the GTPase Rac were studied experimentally by Benjamin Lin, in the Lab of Andre Levchenko [19]. A model for Cdc42, Rac, and Rho, interacting with one another and with the phosphoinositides PIP, PIP\(_2\) and PIP\(_3\) explained the timing and strength of the response, and predicted results of experimental manipulations that affect parts of the crosstalk [14, 19].
Experiments have been carried out on melanoma cells grown on microfabricated surfaces that mimic the natural environment of cells (“extracellular matrix”). JinSeok Park, of the Levchenko Lab at Yale University found three typical motility phenotypes, including persistently polarized, random, and oscillatory front-back cycling, depending on levels of adhesion to the substrate, and manipulations that affect activities of the GTPases or their downstream targets. We were able to account for the observed phenotypes by a model for Rac-Rho mutual antagonism, weighted by signals from the extracellular matrix substrate [16, 26, 29].
4 Extending the Minimal Model
The wave-pinning model has been used as a nucleus from which we have expanded to larger circuits, and greater levels of biological detail. We showed that some properties of the system (4) is shared by a circuit of the mutually antagonistic GTPases Rac-Rho [12]. A notable common feature is the existence of parameter regimes in which several states coexist. These include states of uniformly low activity, uniformly high activity, or polarized levels of activity. Which of these develops then depends on initial conditions. A recent contribution [38] extends these findings to more general model variants.
A hallmark of the kinetics we described above is the presence of bistability in some parameter regimes, i.e. the existence of two stable steady states separated by an unstable one. Such systems also display hysteresis, or a kind of history-dependence: slowly increasing a parameter results in a sudden appearance of a new steady state at some transition point, but to reverse the process, the same parameter has to be decreased much beyond the transition point. The addition of feedback from a third dynamic variable in such cases, is known to produce the possibility of oscillations.
Extensions of the minimal model: a The simplest basic wave-pinning model of Eq. (4) can produce a polarized pattern. b When the GTPase promotes assembly of F-actin, which then promotes GTPase inactivation, waves and other exotic dynamics can be observed, provided the negative feedback is on a slow time-scale [10, 22]. In a, b time increases along the vertical axis and space is on the horizontal axis. c Some GTPases cause the cell to spread (Rac) or to shrink (Rho), affecting cell tension. If the tension also affects GTPase activity, interesting dynamics are observed. Shown is a time sequence (left to right) of a “tissue” composed of \({\approx } 370\) cells, colour coded by their internal GTPase activity. The cell size is correlated to that activity, as described in [37]
We examined several cases of this type, motivated by biological observations. In one case, we studied feedback from F-actin to the inactivation of a GTPase, as observed, for example, in [31]. Assuming slow negative feedback from F-actin (to the inactivation of the GTPase), as shown in Fig. 7b leads to interesting dynamics of traveling waves and pulses in the domain [10, 22]. Feedback between the Rac-Rho circuit and the extracellular matrix also results in oscillations, as previously described [16]. More recently, we also modeled the interplay between mechanical tension in the cell and the activity of GTPases, as observed experimentally by [17]. Here we assumed that GTPase such as Rho and Rac can affect cell spreading, which changes the tension on the cell and feeds back to the activation of the GTPase. A typical circuit of this type is shown in Fig. 7c. As expected, such negative feedback is also consistent with regimes of oscillatory dynamics in individual cells, as demonstrated in [37]. Moreover, when cells with such behaviour are coupled to one another in 1D or in 2D (simulations in Fig. 7c), one observes waves of chemical activity coupled to cell-size changes as the “model tissue” undergoes the spatio-temporal dynamics so created.
5 Discussion
Cell biology presents an unlimited source of inspiring problems. The links between mathematics and cell biology are relatively recent, and not yet fully recognized. But the need for quantitative methods, computational platforms, and mathematical analysis of cellular phenomena promises to grow with time, presenting many opportunities for young applied mathematicians looking for problems to study.
Here I have mainly described a toy model that we constructed to help us understand cell polarization. The simplicity of the model made it mathematically tractable. Its analysis reveals several insights that were not a priori evident. First, with the right kind of positive feedback, we showed that a single GTPase could, on its own, lead to spontaneous polarization that explains cell directionality. In other words, it is not essential to have networks of such proteins to achieve this cellular process. Second, there is a functional purpose for the curious biology of GTPases: their cycling between membrane and cytosol is not a mere evolutionary artifact. We argue that this transition sets up the differences in diffusion between active and inactive GTPases—a difference that is crucial for polarization to be possible, according to our mathematical model.
The motivation of cell polarity led us to mathematics with a surprising twist, uncovering the phenomenon of decelerating waves and wave-pinning that were not widely recognized before in the literature on reaction-diffusion systems. From this standpoint, we could argue that biology inspires new mathematics. The efforts to understand models that were so developed also resulted in a variety of methods that ease the analysis, among them LPA. Extensions of the basic wave-pinning model led to variants with more exotic patterns and waves. These were investigated in various geometries, in single cells, and finally, in interacting groups of cells to identify causes for cell size fluctuations in a tissue and for a variety of emergent phenomena in single and collective cell motility. Finally, developing simple theoretical models and in parallel considering biologically-inspired detailed models are not mutually exclusive. Our experience in the former helps us with the later, and vice versa.
Many still-unanswered questions can be posed. Among these are some of the following: How does the internal GTPase state of a cell affect the outcome of interactions between cells, and how does contact between cells change their GTPase state? What are reasonable ways to model such cell-cell interactions leading to cell adhesion or cell separation? How is cell state coordinated in a multicellular tissue? What aspects of cell adhesion, mechanics, deformation, chemical secretion, and environmental topography (to name a few) affect and are affected by GTPase activities, and how should these be modelled? What methods of analysis can we develop to help with larger, more realistic models that have many interacting components? What aspects of 3D cell shape, and of cell motion in a 3D matrix lead to new phenomena, and what numerical methods should be developed to address such behaviours? Is there a compromise between large-scale computations and mathematical analysis in these more challenging scenarios? In conclusion, the motility and interactions of cells is a rich scientific area calling for investigation by applied mathematicians. Pattern formation inside living cells is merely one facet, while many other fundamental challenges are at hand.
References
Berger, M., Goodman, J.: Airburst-generated tsunamis. Pure Appl. Geophys. 175(4), 1525–1543 (2018)
Burridge, K., Doughman, R.: Front and back by Rho and Rac. Nat. Cell Biol. 8(8), 781 (2006)
Burridge, K., Wennerberg, K.: Rho and Rac take center stage. Cell 116(2), 167–179 (2004)
Buttenschön, A.: Reaction-diffusion bifurcation methods (in preparation) (2021)
Cusseddu, D., Edelstein-Keshet, L., Mackenzie, J.A., Portet, S., Madzvamuse, A.: A coupled bulk-surface model for cell polarisation. J. Theoret. Biol. 481, 119–135 (2019)
Dawes, A.T., Edelstein-Keshet, L.: Phosphoinositides and Rho proteins spatially regulate actin polymerization to initiate and maintain directed movement in a one-dimensional model of a motile cell. Biophys. J. 92(3), 744–768 (2007)
Edelstein-Keshet, L., Holmes, W.R., Zajac, M., Dutot, M.: From simple to detailed models for cell polarization. Philos. Trans. R. Soc. B Biol. Sci. 368(1629), 20130003 (2013)
Etienne-Manneville, S., Hall, A.: Rho GTPases in cell biology. Nature 420(6916), 629 (2002)
Grieneisen, V.: Dynamics of auxin patterning in plant morphogenesis. Ph.D. thesis. University of Utrecht (2009)
Holmes, W., Carlsson, A., Edelstein-Keshet, L.: Regimes of wave type patterning driven by refractory actin feedback: transition from static polarization to dynamic wave behaviour. Phys. Biol. 9(4), 046005 (2012)
Holmes, W.R.: An efficient, nonlinear stability analysis for detecting pattern formation in reaction diffusion systems. Bull. Math. Biol. 76(1), 157–183 (2014)
Holmes, W.R., Edelstein-Keshet, L.: Analysis of a minimal Rho-GTPase circuit regulating cell shape. Phys. Biol. 13, 046001 (2016)
Holmes, W.R., Liao, L., Bement, W., Edelstein-Keshet, L.: Modeling the roles of protein kinase C\(\beta \) and \(\eta \) in single-cell wound repair. Mol. Biol. Cell 26(22), 4100–4108 (2015)
Holmes, W.R., Lin, B., Levchenko, A., Edelstein-Keshet, L.: Modelling cell polarization driven by synthetic spatially graded Rac activation. PLoS Comput. Biol. 8(6), e1002366 (2012). https://doi.org/10.1371/journal.pcbi.1002366
Holmes, W.R., Mata, M.A., Edelstein-Keshet, L.: Local perturbation analysis: a computational tool for biophysical reaction-diffusion models. Biophys. J. 108(2), 230–236 (2015)
Holmes, W.R., Park, J., Levchenko, A., Edelstein-Keshet, L.: A mathematical model coupling polarity signaling to cell adhesion explains diverse cell migration patterns. PLoS Comput. Biol. 13(5), e1005524 (2017)
Houk, A.R., Jilkine, A., Mejean, C.O., Boltyanskiy, R., Dufresne, E.R., Angenent, S.B., Altschuler, S.J., Wu, L.F., Weiner, O.D.: Membrane tension maintains cell polarity by confining signals to the leading edge during neutrophil migration. Cell 148(1–2), 175–188 (2012)
Jilkine, A., Marée, A.F., Edelstein-Keshet, L.: Mathematical model for spatial segregation of the Rho-Family GTPases based on inhibitory crosstalk. Bull. Math. Biol. 69, 1943–1978 (2007)
Lin, B., Holmes, W.R., Wang, J., Ueno, T., Harwell, A., Edelstein-Keshet, L., Takanari Inoue, A.L.: Synthetic spatially graded Rac activation drives directed cell polarization and locomotion. PNAS 109(52), E3668–E3677 (2012)
Marée, A.F., Jilkine, A., Dawes, A., Grieneisen, V.A., Edelstein-Keshet, L.: Polarization and movement of keratocytes: a multiscale modelling approach. Bull. Math. Biol. 68, 1169–1211 (2006)
Marée, A.F.M., Grieneisen, V.A., Edelstein-Keshet, L.: How cells integrate complex stimuli: the effect of feedback from phosphoinositides and cell shape on cell polarization and motility. PLoS Comput. Biol. 8, e1002402 (2012)
Mata, M.A., Dutot, M., Edelstein-Keshet, L., Holmes, W.R.: A model for intracellular actin waves explored by nonlinear local perturbation analysis. J. Theoret. Biol. 334, 149–161 (2013)
Mogilner, A., Oster, G.: Cell motility driven by actin polymerization. Biophys. J. 71(6), 3030–3045 (1996)
Mori, Y., Jilkine, A., Edelstein-Keshet, L.: Wave-pinning and cell polarity from a bistable reaction-diffusion system. Biophys. J. 94(9), 3684–3697 (2008)
Mori, Y., Jilkine, A., Edelstein-Keshet, L.: Asymptotic and bifurcation analysis of wave-pinning in a reaction-diffusion model for cell polarization. SIAM J. Appl. Math. 71, 1401–1427 (2011)
Park, J., Holmes, W.R., Lee, S.H., Kim, H.N., Kim, D.H., Kwak, M.K., Wang, C.J., Edelstein-Keshet, L., Levchko, A.: A mechano-chemical feedback underlies co-existence of qualitatively distinct cell polarity patterns within diverse cell populations. PNAS 114(28), E5750–59 (2017)
Pollard, T.D., Blanchoin, L., Mullins, R.D.: Actin dynamics. J. Cell Sci. 114(1), 3 (2001)
Postma, M., Bosgraaf, L., Loovers, H.M., Van Haastert, P.J.: Chemotaxis: signalling modules join hands at front and tail. EMBO Rep. 5(1), 35–40 (2004)
Rens, E.G., Edelstein-Keshet, L.: Cellular tango: how extracellular matrix adhesion choreographs Rac-Rho signaling and cell movement. Phys. Biol. 18, 066005 (2021)
Ridley, A.J., Schwartz, M.A., Burridge, K., Firtel, R.A., Ginsberg, M.H., Borisy, G., Parsons, J.T., Horwitz, A.R.: Cell migration: integrating signals from front to back. Science 302(5651), 1704–1709 (2003)
Robin, F.B., Michaux, J.B., McFadden, W.M., Munro, E.M.: Excitable RhoA dynamics drive pulsed contractions in the early C. elegans embryo. BioRxiv, p. 076356 (2016)
Simon, C.M., Vaughan, E.M., Bement, W.M., Edelstein-Keshet, L.: Pattern formation of Rho GTPases in single cell wound healing. Mol. Biol. Cell 24(3), 421–432 (2013)
Small, J.V., Resch, G.P.: The comings and goings of actin: coupling protrusion and retraction in cell motility. Curr. Opin. Cell Biol. 17(5), 517–523 (2005)
Svitkina, T.M., Borisy, G.G.: Arp2/3 complex and actin depolymerizing factor/cofilin in dendritic organization and treadmilling of actin filament array in lamellipodia. J. Cell Biol. 145(5), 1009–1026 (1999)
Vanderlei, B., Feng, J.J., Edelstein-Keshet, L.: A computational model of cell polarization and motility coupling mechanics and biochemistry. Multiscale Model. Simul. 9(4), 1420–1443 (2011)
Walther, G.R., Marée, A.F., Edelstein-Keshet, L., Grieneisen, V.A.: Deterministic versus stochastic cell polarisation through wave-pinning. Bull. Math. Biol. 74(11), 2570–2599 (2012)
Zmurchok, C., Bhaskar, D., Edelstein-Keshet, L.: Coupling mechanical tension and GTPase signaling to generate cell and tissue dynamics. Phys. Biol. 15(4), 046004 (2018)
Zmurchok, C., Holmes, W.R.: Modeling cell shape diversity arising from complex Rho GTPase dynamics. bioRxiv, p. 561373 (2019)
Acknowledgements
LEK gratefully acknowledges the contributions of many group members to this research over the years. Among these, special thanks go to A. F. M. Mareé, Y. Mori, W. R. Holmes, A. Jilkine, A. T. Dawes, C. Zmurchok, A. Buttenschön and E. G. Rens. LEK is supported by a Discovery grant from the Natural Sciences and Engineering Research Council of Canada (NSERC).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2022 The Author(s)
About this paper
Cite this paper
Edelstein-Keshet, L. (2022). Pattern Formation Inside Living Cells. In: Chacón Rebollo, T., Donat, R., Higueras, I. (eds) Recent Advances in Industrial and Applied Mathematics. SEMA SIMAI Springer Series(), vol 1. Springer, Cham. https://doi.org/10.1007/978-3-030-86236-7_5
Download citation
DOI: https://doi.org/10.1007/978-3-030-86236-7_5
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-86235-0
Online ISBN: 978-3-030-86236-7
eBook Packages: Mathematics and StatisticsMathematics and Statistics (R0)