Abstract
The human lateral geniculate body (LGB) with its six sickle shaped layers (lam) represents the principal thalamic relay nucleus for the visual system. Cytoarchitectonic analysis serves as the groundtruth for multimodal approaches and studies exploring its function. This technique, however, requires experienced knowledge about human neuroanatomy and is costly in terms of time. Here we mapped the six layers of the LGB manually in serial, histological sections of the BigBrain, a high-resolution model of the human brain, whereby their extent was manually labeled in every 30th section in both hemispheres. These maps were then used to train a deep learning algorithm in order to predict the borders on sections in-between these sections. These delineations needed to be performed in 1 µm scans of the tissue sections, for which no exact cross-section alignment is available. Due to the size and number of analyzed sections, this requires to employ high-performance computing. Based on the serial section delineations, high-resolution 3D reconstruction was performed at 20 µm isotropic resolution of the BigBrain model. The 3D reconstruction shows the shape of the human LGB and its sublayers for the first time at cellular precision. It represents a use case to study other complex structures, to visualize their shape and relationship to neighboring structures. Finally, our results could provide reference data of the LGB for modeling and simulation to investigate the dynamics of signal transduction in the visual system.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
- Lateral geniculate body (LGB)
- Corpus geniculatum laterale (CGL)
- BigBrain
- Deep learning
- 3D reconstruction
- Cytoarchitecture
1 Introduction
The lateral geniculate body (LGB, lat. Corpus geniculatum laterale, from now on LGB) plays a key role in visual perception. Together with the medial geniculate body, which is involved in auditive processing, both nuclei constitute the metathalamus. The LGB is located on the ventral surface of the brain. It mainly receives connections from the retina via the optic tract, but also from layer 6 of the visual cortex and the reticular nucleus of the thalamus [1]. The most prominent efferent projections reach the primary visual cortex, i.e. Brodmann’s area 17 (or area V1, or hOc1 [2]; via the optic radiation (Fig. 1 top image) [1]. The human LGB consists of six layers. The two most ventrally located layers (layers 1 and 2) consist of larger neurons and are known as magnocellular, while layers 3 to 6 are parvocellular layers (Fig. 1 bottom image). Koniocellular neurons are located in between those laminae.
The human visual pathway (top image). The location of the lateral geniculate body (LGB) is marked by the black rectangle. Six layers of the LGB (bottom image) describing contralateral projections depicted in green and ipsilateral projections in red. Created with BioRender.com (Color figure online)
Layers 2, 3 and 5 receive fibers from the ipsilateral eye, whereas layers 1, 4 and 6 receive fibers from the contralateral eye [3]. I.e., each layer receives information from one eye only. Later on, the information will be merged to be processed and interpreted as a binocular image in the visual cortex [4]. Approximately 80% of the retinal information derive from midget ganglion cells and are transferred to the parvocellular neurons in the LGB in layers 3 to 6. These small neurons are specialized in object and detail recognition due to their ability of generating a high spatial resolution and red-green color vision [4]. Midget cells are characterized as small, color-sensitive slow adapting cells from the retina, contrary to parasol cells. Retinal parasol cells send impulses to the bigger magnocellular layers 1 and 2, which are functional for time resolution and hence for the perception of position and movement [5]. Non-Midget-non-Parasol ganglion cells from the retina project to koniocellular neurons of the LGB, which further project to the primary visual cortex, similarly to the parvo- and magnocellular systems. Since the LGB transfers retinal information directly to the primary visual cortex via the optic radiation, it is also defined as first-order relay [1]. The koniocellular neurons most probably play a role in color perception [4]. The middle part of the LGB in coronal sections is called the hilum, and the taperings at the medial and lateral ends of the LGB are called the medial and lateral horn of the LGB, respectively [6].
Lesions in the LGB can affect the function of the visual pathway. For example, patients suffering from multiple sclerosis or Alzheimer’s disease show a general volume loss of the LGB, indicating in the latter case a significant relation between degeneration of the LGB and amyloid-β pathology [7, 8]. After loss of visual experience due to an injury of the primary visual cortex but with an intact LGB, certain visual information processing still exists. This phenomenon discovered by functional magnetic resonance imaging (fMRI) is called blindsight [9].
This work aims to map the LGB and its layers at microscopical resolution, to provide reference data for studies targeting the LGB in the living human brain, to develop a use case enabling the combination of expert annotations based on cytoarchitectonic criteria in a subset of sections with deep learning in order to increase the number of delineated sections. A 3D reconstruction of the human LGB in both hemispheres of the BigBrain dataset was computed and visualized [10].
2 Materials and Methods
2.1 Histology
The BigBrain (65-year-old male) comes from the body donor program of the Anatomical Institute of Düsseldorf in accordance to legal and ethical requirements. Prior to histological processing MRI images were taken (1.5 T, Siemens Medical Systems GmbH, Erlangen, Germany) in order to provide a reference volume for subsequent image registration. Histological processing was previously described in detail [10, 11]. In short, the brain was fixed in 4% buffered formalin, embedded in paraffin and cut in coronal plane, each Sect. 20 µm thick. Every section – of total 7404 – was mounted and silver stained for cell bodies. The sections were digitized using a TISSUEscope™ LE120 Scanner (Huron Digital Pathology) [10]. The spatial resolution of the images is 1 µm in-plane, the average size of the images is 10 GByte per section.
2.2 Manual Analysis and Reference Mapping of Histological Sections
High-resolution images were analyzed and manually delineated using SectionTracer, an online tool written in JavaScript [12]. Borders of the LGB and its six layers were traced in 16 sections per hemisphere, i.e. every 30th image at a distance of 600 µm.
The human LGB is surrounded by white matter, i.e. fiber tracts, and could therefore be clearly distinguished from its neighboring structures: the thalamus was found dorso-medially from the LGB while the medial geniculate body was located medially. At rostral levels the LGB was completely surrounded by white matter.
The borders of the six layers of the LGB were identified based on differences in the cytoarchitecture (Fig. 2). Magno- and parvocellular layers were mainly distinguished according to their cell-size and -density. The pale koniocellular neurons served as main indicators for the borders between the different layers of the LGB. Wherever these criteria were not sufficient, i.e. where two parvocellular layers were not separated by a koniocellular lamina, cytoarchitectonic criteria such as size, shape, density and distribution of neurons were applied. Borders were drawn where the cytoarchitectonic pattern changed. Layers were numbered according to their position, starting from the most ventral layer 1 at the brain surface, increasing to the uppermost dorsal layer 6.
The volumes of the LGB and its layers were measured using Cavalieri’s principle. A shrinkage factor of 1,931 was used for calculations in order to consider the shrinkage occurring during histological processing, the fixation, respectively [13].
The layers were then 3D reconstructed and transferred to the BigBrain space, which has a spatial resolution of 20 µm isotropic [12]. Results were visualized using the software ParaView [14, 15].
2.3 Training of the Deep-Learning Algorithm to Predict Missing Delineations
A convolutional neural network (CNN) was trained to classify each image pixel according to each lamina of the LGB. Network architecture and training procedure were based on Schiffer et al. [16], as they have been used successfully to aid mapping of cytoarchitectonic areas in several cortical brain regions [17,18,19,20].
The workflow uses a U-shape architecture for brain area segmentation with two encoder branches capturing the input data at different spatial scales. Training and prediction was controlled by the web-based interface of the tool, and performed remotely on the supercomputer JURECA [21] at Jülich Supercomputing Centre (JSC). The training time on the HPC system was around 70 min.
After training, the CNN was applied to automatically classify layers of LGB in sections without annotations. Automatically created annotations were quality-checked to exclude misclassified sections. Annotations were then non-linearly transformed [22] into the 3D reconstructed BigBrain space [10]. Previously excluded sections were replaced by interpolation [23]. Finally, the marching cubes algorithm [24] was applied to extract a surface mesh for each layer of LGB. This 3D reconstruction step is not part of the tool provided by [16], but it follows directly the experimental protocol for the BigBrain dataset described therein.
3 Results
3.1 Cytoarchitectonic Mapping Based on Expert Annotations and Deep Learning
The analysis of the LGB allowed to identify six layers, partially different in shape and size (Fig. 2). The two ventral layers contained magnocellular neurons of triangular and multiform shapes and differed significantly in the cytoarchitecture from the parvocellular layers 3–6. Layers 1 and 2 were thinner and more elongated; contained round and oval shaped neurons. Layers 3 and 6 were most prominent and were reached over a larger distance than the other two layers of the parvocellular part. Layer 5 was the shortest and least developed layer with respect to its mediolateral extent. On most sections, the layers were separated by a white koniocellular line, characterized by a low cell density. On a few sections, layers 1 and 4 as well as 4 and 6 were connected. The same was true for layers 2 and 3 as well as 3 and 5, where the koniocellular lines were lacking (Fig. 2a, black arrow). Although layers 1 and 2 were the thinnest structures, the magnocellular neurons are clearly visible, due to the magnocellular neurons (Fig. 2a, inset). Layers 3 to 6, on the other hand, were composed of smaller and more densely packed cells. The neurons on the lateral and medial horn of the LGB in each layer were loosely packed, while the neurons at the hilum were denser.
In total, 13 sections were labeled by an expert in the left hemisphere and 11 sections in the right hemisphere, with a distance of 0.6 mm. The LGB of the left hemisphere was found and processed by the CNN on 366 sections (rostro-caudal extent 7.3 mm) and on 293 coronal sections (extent of 5.9 mm) of the right hemisphere. For comparison between the expert annotation and the prediction of the CNN see Fig. 2a and b.
3.2 High-Resolution 3D Reconstruction
The exact location of the human LGB is depicted in the BigBrain model in Fig. 3a. A high-resolution 3D reconstruction was performed to get a deeper insight into the complex shape of the human LGB. The prominent hilum and the elongated lateral horn are characteristic for the shape of the human LGB (Fig. 3b). In more detail, the dorsal surface of the LGB is mainly covered by layers 4, 5 and 6. Layer 4 is more prominent at the hilum and the medial horn (Fig. 3b), while layer 3 is most prominent at the lateral horn (Fig. 3c). Most parts of the ventral surface of the LGB are covered by layer 1 medially and by layer 2 laterally (Fig. 3c).
3.3 Volumes of Layers
The total volumes of the LGB, after consideration of the shrinkage factor, are 120.5 mm3 on the left, and 113.2 mm3 on the right hemisphere (Table 1). The parvocellular layers are bigger than the magnocellular layers. The total volume of the LGB is 141.2 on the left, and 132.2 mm3 on the right hemisphere. It contains the six layers, but also koniocellular layers and blood vessels, which are not included in the volumes of the single layers.
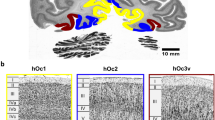
Shape and borders of the six layers (labeled by different colors and numbers) of the human lateral geniculate body (LGB) on coronal sections. Upper row shows a comparison between manual expert mapping (a) and the prediction of the CNN (b). The location of the LGB on the coronal section is shown in (c). One caudal (d) and one rostral section (e) indicate different shapes of the LGB at different levels of sectioning. Scale bars: a–b, c–e 1 mm, c 20 mm. Scalebar magnification in a: 50 µm.
3D reconstruction of the lateral geniculate body (LGB). (a) Localization of the left lateral geniculate body (LGB) in the BigBrain model. (b-c) Surface of the reconstructed left and right LGB revealing its specific shape and its different layers. (b) dorso-caudal view (c) ventro-rostral view. Abbreviations: d = dorsal, l = lateral, m = medial, v = ventral.
4 Discussion and Conclusion
This study presents the first application for mapping a complex subcortical gray matter structure, the LGB, combining an expert-based and a deep-learning approach, resulting in a high-resolution 3D reconstruction of the LGB in the BigBrain template. It shows, for the first time, the shape of the individual six layers in 3D space while previous information was mainly obtained from 2D sections. The maps are publicly available in the EBRAINS multilevel human brain atlas (https://ebrains.eu/service/human-brain-atlas), where they can be explored in an interactive 3D viewer (https://interactive-viewer.apps.hbp.eu/saneUrl/BigBrain_LGB) and can be found under the DOI: https://doi.org/10.25493/33Z0-BX.
Due to the large size of the human brain, subcortical nuclei such as the LGB and cortical areas are found in tens to hundreds to thousands of sections, in dependence on the size of the structure. To map a structure manually in every section using traditional methods becomes impossible when structures are complex or large. As an alternative option, the extent of the structures can be interpolated. The drawback of this method is that images have to be 3D reconstructed before further processing [16]. Herein, we provide a use case using a semi-automated prediction of borders supported by deep learning. The algorithm learns from manually annotated borders and is able to use this knowledge for annotating the same area in previously unseen sections. Since it directly interprets the texture and brain topology, it is much more precise than a 3D interpolation in the reconstructed space and allows to use unregistered single sections.
Due to the large number of images and large size of the sections, training and application of the CNN was performed on the supercomputer system JURECA [18] at Jülich Supercomputing Centre (JSC). The use of high-performance computers becomes even more relevant when larger structures are being analyzed in whole brain sections. While computation in the LGB was completed in 2 h, comparable computations for the thalamus, for example, would take 50 h. Existing work on automatic classification of cytoarchitectonic cortical areas [16] also indicate that computational effort increases considerably when trying to automatically identify larger brain regions with more complex cytoarchitectonic and morphological properties.
Our findings of the total volume of the LGB, shrinkage factor included, is in line with previous findings. Andrews and colleagues reported mean LGB volumes of 121 mm3 for the right and 115 mm3 for the left hemisphere; the variance was quite high and ranged from 91.1 to 157 mm3 for both hemispheres [25]. Further investigations in additional brains in the future would help to better understand intersubject variability in terms of the size and/or shape of the LGB.
High-resolution mapping data of the LGB may open a broad field of applications. For example, routine Magnetic Resonance Imaging (MRI) studies often lack sufficient contrast and/or spatial resolution and could benefit from such atlas data. Current studies on the implementation of electrically stimulated prostheses in the visual cortex aim to restore part of the vision in blind people by multiple stimulations of electrodes to percept light [26, 27], where such maps could be applied in the future to increase the localization accuracy. The investigation of the pathomechanisms of diseases where the visual pathway is affected, such as Multiple Sclerosis or glaucoma could be supported by the maps [28, 29]. The maps provide input data for modelling and simulation of different types of neurons, neuronal pathways or networks using platforms like The Virtual Brain [30]. With respect to basic neuroscience, the visualization and differentiation of the layers is expected to contribute to a more in-depth analysis of information processing in visual pathways. The present approach provides a use case for application in other brain areas and brains of other species enabling a fast and detailed prediction of the extent of small and complex structures, including visualization and volume analyses, with a minimum of manual effort and time expenditure.
References
Sherman, S.M., Guillery, R.W.: The role of the thalamus in the flow of information to the cortex. Philos Trans. R. Soc. B Biol. Sci. 357(1428), 1695–1708 (2002). https://doi.org/10.1098/rstb.2002.1161
Amunts, K., Malikovic, A., Mohlberg, H., Schormann, T., Zilles, K.: Brodmann’s areas 17 and 18 brought into stereotaxic space - where and how variable? Neuroimage 11(1), 66–84 (2000). https://doi.org/10.1006/nimg.1999.0516
Duggan, W.F.: Anatomy of the eye and orbit. Arch. Ophthalmol. 10(5), 723–724 (1933). https://doi.org/10.1001/archopht.1933.00830060147017
Eiber, C.D., et al.: Receptive field properties of koniocellular on/off neurons in the lateral geniculate nucleus of marmoset monkeys. J. Neurosci. 38(48), 10384–10398 (2018). https://doi.org/10.1523/JNEUROSCI.1679-18.2018
Purves, D.: Neuroscience, 5th edn. Sinauer Associates Inc., Sunderland (2012)
Prasad, S., Galetta, S.L.: Anatomy and physiology of the afferent visual system. Handbook of Clinical Neurology, vol. 102 (2011)
Papadopoulou, A., et al.: Damage of the lateral geniculate nucleus in MS. Neurology 92(19), e2240–e2249 (2019). https://doi.org/10.1212/WNL.0000000000007450
Erskine, D., et al.: Changes to the lateral geniculate nucleus in Alzheimer’s disease but not dementia with Lewy bodies. Neuropathol. Appl. Neurobiol. 42(4), 366–376 (2016). https://doi.org/10.1111/nan.12249
Schmid, M.C., et al.: Blindsight depends on the lateral geniculate nucleus. Nature 466(7304), 373–377 (2010). https://doi.org/10.1038/nature09179
Amunts, K., et al.: BigBrain: an ultrahigh-resolution 3D human brain model. Science (80-) 340(6139), 1472–1475 (2013). https://doi.org/10.1126/science.1235381
Amunts, K., Zilles, K.: Architectonic mapping of the human brain beyond Brodmann. Neuron 88(6), 1086–1107 (2015). https://doi.org/10.1016/j.neuron.2015.12.001
Amunts, K., Mohlberg, H., Bludau, S., Zilles, K.: Julich-Brain: a 3D probabilistic atlas of the human brain’s cytoarchitecture. Science (80-) 369(6506), 988–992 (2020). https://doi.org/10.1126/science.abb4588
Amunts, K., Schleicher, A., Zilles, K.: Cytoarchitecture of the cerebral cortex-more than localization. Neuroimage 37(4), 1061–1065 (2007). https://doi.org/10.1016/j.neuroimage.2007.02.037
Ahrens, J., Geveci, B., Law, C.: ParaView: an end-user tool for large data visualization. www.paraview.org. Accessed 11 Feb 2021
The ParaView Guide | ParaView. https://www.paraview.org/paraview-guide/. Accessed Feb 11 2021
Schiffer, C., et al.: Convolutional neural networks for cytoarchitectonic brain mapping at large scale, November 2020. http://arxiv.org/abs/2011.12857. Accessed 20 Dec 2020
EBRAINS - Ultrahigh resolution 3D cytoarchitectonic map of Area hOc1 (V1, 17, CalcS) created by a Deep-Learning assisted workflow. https://search.kg.ebrains.eu/instances/Dataset/696d6062-3b86-498f-9ca6-e4d67b433396. Accessed 6 Mar 2021
EBRAINS - Ultrahigh resolution 3D cytoarchitectonic map of Area hOc2 (V2, 18) created by a Deep-Learning assisted workflow. https://search.kg.ebrains.eu/instances/Dataset/63093617-9b72-45f5-88e6-f648ad05ae79. Accessed 6 Mar 2021
EBRAINS - Ultrahigh resolution 3D cytoarchitectonic map of Area hOc3v (LingG) created by a Deep-Learning assisted workflow. https://search.kg.ebrains.eu/instances/Dataset/f746514d-b79a-48e2-9c07-39f7c62459cf. Accessed 6 Mar 2021
EBRAINS - Ultrahigh resolution 3D cytoarchitectonic map of Area hOc5 (LOC) created by a Deep-Learning assisted workflow. https://search.kg.ebrains.eu/instances/Dataset/ea8fb74b-0ecc-4801-9522-b4c2cb2a2a5c. Accessed 6 Mar 2021
Krause, D., Thörnig, P.: JURECA: modular supercomputer at Jülich Supercomputing Centre. J. Large-Scale Res. Facil. JLSRF 4, A132 (2018). https://doi.org/10.17815/jlsrf-4-121-1
Omidyeganeh, M., et al.: Non-linear registration of 1 μm histology sections into 3D 20 μm BigBrain space (2020)
Schober, M., Axer, M., Huysegoms, M., Schubert, N., Amunts, K., Dickscheid, T.: Morphing image masks for stacked histological sections using laplace’s equation. In: Tolxdorff, T., Deserno, T.M., Handels, H., Meinzer, H.-P. (eds.) Bildverarbeitung für die Medizin 2016. I, pp. 146–151. Springer, Heidelberg (2016). https://doi.org/10.1007/978-3-662-49465-3_27
Lewiner, T., Lopes, H., Ilson Vieira, A.W., Tavares, G.: Efficient implementation of Marching Cubes’ cases with topological guarantees. J. Graph. Tools 8(2), 1–15 (2003)
Andrews, T.J., Halpern, S.D., Purves, D.: Correlated size variations in human visual cortex, lateral geniculate nucleus, and optic tract. J. Neurosci. 17(8), 2859–2868 (1997). https://doi.org/10.1523/jneurosci.17-08-02859.1997
Chen, X., Wang, F., Fernandez, E., Roelfsema, P.R.: Shape perception via a high-channel-count neuroprosthesis in monkey visual cortex. Science 370(6521), 1191–1196 (2020). https://doi.org/10.1126/science.abd7435
Mirochnik, R.M., Pezaris, J.S.: Contemporary approaches to visual prostheses. Mil. Med. Res. 6(1), 1–9 (2019). https://doi.org/10.1186/s40779-019-0206-9
Nuzzi, R., Dallorto, L., Rolle, T.: Changes of visual pathway and brain connectivity in glaucoma: a systematic review. Front. Neurosci. 12(May), 363 (2018). https://doi.org/10.3389/fnins.2018.00363
Sepulcre, J., et al.: Contribution of white matter lesions to gray matter atrophy in multiple sclerosis evidence from voxel-based analysis of T1 lesions in the visual pathway. Arch. Neurol. 66(2), 173–179 (2009). https://doi.org/10.1001/archneurol.2008.562
Schirner, M., et al.: Brain modelling as a service: the virtual brain on EBRAINS, February 2021. http://arxiv.org/abs/2102.05888. Accessed 1 June 2021
Acknowledgements
This project has received funding from the European Union’s Horizon 2020 Framework Programme for Research and Innovation under the Specific Grant Agreement No. 945539 (Human Brain Project SGA3). This work was also funded by Helmholtz Association’s Initiative and Networking Fund through the Helmholtz International BigBrain Analytics and Learning Laboratory (HIBALL) under the Helmholtz International Lab grant agreement InterLabs-0015. Computing time was granted through JARA on the supercomputer JURECA at Jülich Supercomputing Centre (JSC).
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2021 The Author(s)
About this paper
Cite this paper
Brandstetter, A., Bolakhrif, N., Schiffer, C., Dickscheid, T., Mohlberg, H., Amunts, K. (2021). Deep Learning-Supported Cytoarchitectonic Mapping of the Human Lateral Geniculate Body in the BigBrain. In: Amunts, K., Grandinetti, L., Lippert, T., Petkov, N. (eds) Brain-Inspired Computing. BrainComp 2019. Lecture Notes in Computer Science(), vol 12339. Springer, Cham. https://doi.org/10.1007/978-3-030-82427-3_2
Download citation
DOI: https://doi.org/10.1007/978-3-030-82427-3_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-82426-6
Online ISBN: 978-3-030-82427-3
eBook Packages: Computer ScienceComputer Science (R0)