Abstract
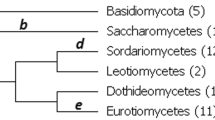
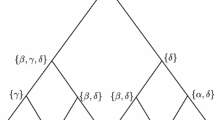
Phylogenetics deals with the task of reconstructing all the ancestral relations among a set of lineages. For any given phylogenetic tree that explains their evolution, subject to the Kimura Three Parameters Model, Hadamard Conjugation is an equation that relates its tree weights (within a matrix Q) to its substitution patterns distribution (within a matrix P) on leaves. In practice, the latter matrix is approximated via the DNA sequence alignment. Some challenges have to be faced in the process such as filling the gaps when they exist. Throughout this manuscript, it is provided fully explanation of how the matrix P can be approximated. The authors contribute to the scientific community with one library running on Maple to lead with these tasks. We conclude this work with a fully developed example focused on three spieces of orchids of the genus Lophiarella distributed in southwestern Mexico and northwestern Mesoamerica for which we obtained the matrix P.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Chor, B., Hendy, M.D., Snir, S.: Maximum likelihood jukes-cantor triplets: analytic solutions. Mol. Biol. Evol. 23(3), 626–632 (2006). https://doi.org/10.1093/molbev/msj069
McBee, C.D., Penttila, T.: Remarks on Hadamard conjugation and combinatorial phylogenetics. Australas. J. Comb. 66(2), 177–191 (2016)
Álvarez, E.: Spectral Sequence Spectrum for an m nucleotide sequence alignment with or without GAPS (2021). https://doi.org/10.5281/zenodo.4587007
Carnevali, G., Cetzal-Ix, W., Balam, R.N., Leopardi, C., Romero-González, G.A.: A combined evidence phylogenetic re-circumscription and a taxonomic revision of lophiarella (Orchidaceae: Oncidiinae). Syst. Botany 38(1), 46–63 (2013). https://doi.org/10.1600/036364413X661926
Nixon, K.C., WinClada ver. 1.00. 08. Published by the author, Ithaca, NY (2002). http://www.cladistics.com/ aboutWinc.htm
Casanellas, M.: El modelo evolutivo de Kimura: un enlace entre el álgebra, la estadística y la biología. La Gaceta de la RSME. 2, 241–257 (2018)
Holder, M., Lewis, P.O.: Phylogeny estimation: traditional and Bayesian approaches. Nat. Rev. 4, 275–284 (2003). https://doi.org/10.1038/nrg1044
Simmons, M.P., Ochoterena, H.: Gaps as characters in sequence-based phylogenetic analyses. Syst. Biol. 49, 369–381 (2000). https://doi.org/10.1093/sysbio/49.2.369
Hendy, M.D., Charleston, M.A.: Hadamard conjugation: a versatile tool for modelling nucleotide sequence evolution. New Zealand J. Botany. 31, 231–237 (1993). https://doi.org/10.1080/0028825X.1993.10419500
Hendy, M.D., Penny, D., Steel, M.A.: A discrete Fourier analysis for evolutionary trees. Proc. Natl. Acad. Sci. USA 91, 3339–3343 (1994). https://doi.org/10.1073%2Fpnas.91.8.3339
Hendy, M.D., Snir, S.: Hadamard conjugation for the kimura 3ST model: combinatorial proof using path sets. IEEE/ACM Trans. Comput. Biol. Bioinf. 5(3), 461–471 (2008). https://doi.org/10.1109/TCBB.2007.70227
Steel, M.A., Hendy, M.D., Székely, L.A., Erdös, P.L.: Spectral analysis and a closest tree method for genetic sequences. Appl. Math. Lett. 5(6), 63–67 (1992). https://doi.org/10.1016/0893-9659(92)90016-3
Kimura, M.: Estimation of evolutionary sequences between homologous nucleotide sequences. Proc. Nat. Acad. Sci., USA 78, 454–458 (1981). https://doi.org/10.1073/pnas.78.1.454
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Appendix
Appendix
Voucher information and GenBank accesion numbers on nrITS DNA for taxa used in Sect. 3:
-
Lophiarella flavovirens, Mexico, Colima, Carnevali 7270 (CICY), JQ319734;
-
Lophiarella microchila, Mexico, Chiapas, Carnevali 7643 (CICY), JQ319735;
-
Lophiarella splendida, Guatemala, Carnevali 7232 (CICY), JQ319736.
Nucleotide sequences, as they were used on Maple, are:


Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
González, E.Á., Balam-Narváez, R. (2021). Computation of the Observed Spectral Sequence Spectrum for Nucleotide Sequence Alignments. In: Corless, R.M., Gerhard, J., Kotsireas, I.S. (eds) Maple in Mathematics Education and Research. MC 2020. Communications in Computer and Information Science, vol 1414. Springer, Cham. https://doi.org/10.1007/978-3-030-81698-8_3
Download citation
DOI: https://doi.org/10.1007/978-3-030-81698-8_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-81697-1
Online ISBN: 978-3-030-81698-8
eBook Packages: Computer ScienceComputer Science (R0)