Abstract
This chapter presents information about the development and use of Electronic Health Record (EHR) Databases. There are petabytes of untapped research data hoarded within hospitals worldwide. There is enormous potential in the secondary analysis of this clinical data, leveraging data already collected in everyday medical practice, we could gain insight into the clinical decision-making process and it’s impact on patient outcomes. In this chapter we outline a high-level overview of some of the important considerations when building clinical research databases.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
FormalPara Learning ObjectivesThe main objectives include:
-
Understand the differences between traditional disease information repositories and EHR databases, and why they are useful
-
Review examples of current EHR clinical databases
-
Learn the necessary steps to develop an EHR clinical database
1 Background
1.1 Introduction to Clinical Databases
Health care information has traditionally been presented in “disease repositories”―a listing of manually collected disease specific information, often stored as aggregate registries. More recently, clinical databases have been developed, resulting in new ways to present, understand, and use health care data. Databases are defined as sets of routinely collected information, organized so it can be easily accessed, manipulated, and updated. Different from disease repositories, these new clinical databases are characterized by heterogeneous patient-level data, automatically gathered from the EHRs. They include many high-resolution variables originating from a large number of patients, thus allowing researchers to study both clinical interactions and decisions for a wide range of disease processes.
Two important phenomena accelerated the evolution of traditional disease repositories into new clinical databases. The first one is the global adoption of EHRs, in which paper-based systems are transformed into digital ones. Although the primary purpose of EHRs is not data collection, their implementation allows health systems to automatically gather large amounts of data (Bailly et al. 2018). Recognizing the enormous potential of the secondary analysis of these data for initiatives from quality improvement to treatment personalization, health and research institutions have started to leverage these novel clinical databases. The second phenomenon that supported the development of clinical databases is the extraordinary expansion of computational power that allowed the development of the necessary infrastructure to store vast amounts of diverse data, and the capacity to process it in a reasonable timeframe. These events enabled the emergence of the field of data science and machine learning. This new knowledge has been made accessible to a large and global audience through new massive open online courses (MOOCs), spurring substantial interest in analysis of large amounts of health data and the opportunity to crowdsource new machine learning techniques through available open source programming tools. (Sanchez-Pinto et al. 2018).
1.2 Goals for Database Creation
The main goal of creating a healthcare database is to put clinical information in a format that can be intuitively explored and rapidly processed, allowing researchers to extract valuable knowledge from the data. In a traditional database, there are relational structures built into store the data which guarantee consistency of the relationships between its entities (e.g. between patient and hospital visit). These structures are commonly referred to as “data models”, and consist of the definition of tables, fields, and requirements for that database. When developing such models, it is essential to capture meaningful representation of the concepts and processes we want to study. This can be a challenge in health care because there are many different actors, and faithfully representing their relationships is crucial to understand what is occurring and also to achieve relevant and reliable research conclusions. Another critical step when creating and maintaining a clinical database is incorporating data quality and security, so it can be appropriately and reliably used in secondary data analysis.
1.3 Examples of Clinical Databases Worldwide
1.3.1 Medical Information Mart for Intensive Care (MIMIC)
The Medical Information Mart for Intensive Care (MIMIC) (Johnson et al. 2016) is one of the most popular and widely used open access clinical databases worldwide. Launched in 2003, MIMIC originated from a partnership between the Massachusetts Institute of Technology (MIT) Laboratory for Computational Physiology, Philips Medical Systems, and Beth Israel Deaconess Medical Center, with funding from the National Institute of Biomedical Imaging and Bioengineering. It is currently in its third version and has de-identified data from 40,000 medical and surgical patients admitted to the Beth Israel Deaconess Medical Center (BIDMC). Originally created with the aim of leveraging machine learning in the healthcare setting to build advanced ICU patient monitoring and decision support systems, MIMIC’s main goal is to improve the efficiency, accuracy, and timeliness of clinical decision-making for ICU patients.
MIMIC has been used for many clinical studies from independent researchers (Aboelsoud et al. 2018; Johnson et al. 2018; Komorowski et al. 2018; Sandfort et al. 2018; Serpa Neto et al. 2018; Waudby-Smith et al. 2018; Block et al. 2018; Collins et al. 2014; Computing NCfB 2018; Deliberato et al. 2018; Dernoncourt et al. 2017; Desautels et al. 2016; Desautels et al. 2017; Farhan et al. 2016; Feng et al. 2018; Fleurence et al. 2014; Ghassemi et al. 2014; Johnson et al. 2016). Since its first version, MIMIC allowed researchers to freely access the data, after registering, completing a preliminary course on human research, and abiding by a data use agreement to avoid the potential misuse of clinical data. This has been one of the main reasons for its popularity in the clinical research community, along with the enormous quantity of diverse information for all patients in MIMIC, making complex cross-evaluating studies feasible. Another important feature for researchers is that individual patient consent has been waived by BIDMC’s Institutional Review Board, an essential and challenging prerequisite to allow for a clinical database to go public in the real world.
In addition to clinical data extracted from the EHR such as demographics, diagnoses, lab values, vital signs, events, and medications, there is a subset of patients with bedside monitor waveforms from ECG, EEG, and vital sign tracings that are stored in flat binary files with text header descriptors. MIMIC also maintains documentation of data structure and a public GitHub repository for researchers interested in working with the database. As result, new users can benefit from the work of others by accessing the available code, and are encouraged to contribute their own work, thereby strengthening and furthering the impact of MIMIC.
1.3.2 eICU Collaborative Research Database (eICU-CRD)
Another example of an open-access database is the eICU Collaborative Research Database (eICU-CRD) (Pollard et al. 2018). This project is derived from a critical care telehealth initiative by Philips® Healthcare. The eICU-CRD was made freely available by the same team as MIMIC and features a distinct patient pool originating from 208 ICUs across the U.S. from 2014 to 2015. As a result, MIMIC and eICU-CRD are independent yet complementary. Similar to MIMIC, the main objective of the project is to boost collaboration in secondary analysis of electronic health records, through the creation of openly available repositories.
1.3.3 Other Databases for Collaborative Research
There are other clinical databases that can be used for collaborative research, although access is more restricted, and data tend to be more general and less granular than the clinical information available in MIMIC or eICU-CRD. One example is PCORnet (Collins et al. 2014), a patient-centered clinical research project that aims to build a national research network, linked by a common data platform and embedded in clinical care delivery systems (Collins et al. 2014; Fleurence et al. 2014). This network aims to provide enough data for studies of rare or uncommon clinical entities, that have been difficult to conduct with the “classical” model. Medical record collections from over 60 million patients allow for large-scale observational and interventional trials to be accomplished more easily (Block et al. 2018). Access to the data can be requested through their web platform “Front Door” and is granted with a case-by-case policy depending on the project.
Other initiatives aim to create common data models, enabling the construction of multiple databases using a common ontology, so that data from each source means the same thing. The Observational Medical Outcomes Partnership (OMOP) and i2b2 have been established using this concept and aim to translate healthcare concepts to a common language in order to facilitate the sharing of meaningful data across the compatible databases. OMOP is managed by a network called Observational Health Data Science and Informatics (OHDSI), a multi-stakeholder, interdisciplinary collaborative network that spans over 600 million patients. A list of databases ported to their model can be found at their website (Observational Health Data Sciences and Informatics (OHDSI) 2018; OMOP CDM 2018). The i2b2 tranSMART Foundation (Computing NCfB. i2b2 (Informatics for Integrating Biology and the Bedside) 2018) is a member-driven non-profit foundation with an open-source/open-data strategy. It provides an open-source data model similar to OMOP, and a list of databases can be found at their website (Computing NCfB 2018). Both OMOP and i2b2 have open source software tools to manage and access the data, and a very active community of users with forums where relevant information and implementation tips can be found.
2 Exercise: Steps for Building an EHR Database for Research
2.1 Putting Together the Right Team
One of the most important steps at the start of any successful project is putting together the right team. Bringing together the range of skilled professionals with the required skills is essential when building an EHR database. One key role is that of a clinician with the knowledge to understand and decipher the highly specialized data collected in the EHR, especially because these data are often poorly organized within the EHR. Clinicians also have an important role in assessing the accuracy of the resulting database and working with data scientists to optimize its usability for targeted end-users.
Another critical member for the team is someone with substantial knowledge in data architecture, who can ensure consistency while modeling the highly complex data from EHRs. This person needs to work closely with the clinicians and data scientists to achieve a high quality, functional clinical database.
2.2 The Six Steps to Building an EHR Database
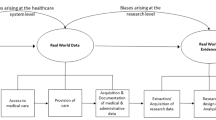
Once the multidisciplinary team has been formed, the next step is familiarizing everyone with the steps for building the database. This is important because the process of building a clinical database is iterative and continuous, as developers work to guarantee data quality and security. The six main stages for developing a clinical database are summarized in Fig. 4.1.
2.2.1 Step 1: Access and Data Model
At the start of the project, it can be helpful to acquire some resources to help with some of the laborious tasks that are inherent in the process of building clinical databases. For instance, if the clinicians building the database work at the hospital where the data is generated, obtaining access to a real time copy of the EHR source database (Step 1, Fig. 4.1) can facilitate mapping. In this scenario, clinicians can use one of their current patients to search for the information that the data architects are trying to map in the EHR system. This helps locate the data to be mapped in the source database. It also helps validate the mapping, by comparing the current reported values after the information is found. This resource is extremely valuable in the assessment of data consistency, since most of the data found in these source databases are used only for maintenance of system functionalities and have no clinical value, thus confusing the mapping process. Although obtaining real time copy of databases may be useful, it can be difficult to do in resource limited settings. In such cases, looking for other ways using the available computational infrastructure in order to acquire the data in a faster time frame is recommended, as any highly available data is valuable in building the database and creating a data-driven environment.
In addition to working with a copy of the EHR source database, the database development team needs to combine their skills in data architecture with their knowledge about the targeted uses of the database in order to find a data model that would fit all the stakeholders’ requirements (Step 1, Fig. 4.1). Balancing these needs is difficult, but critically important at this stage. While modeling all data to fit clinical or analytical mindsets might be desired, creating a model using high granularity and resolution data causes some limitations. Additionally, if conducting multicenter studies is one of the goals, the adoption of open-source health data models, or converging to a data model that can be used by prospective partners might be taken into consideration, as the use of common data models not only facilitates those studies, but also improves their reliability. It is important to emphasize that there is no ideal model and it is highly recommended to choose a common data model most likely to become part of the initiatives already in place in your institution, having an active voice in the process, and helping the community to decide the future direction of the model.
2.2.2 Data Mapping
With access to the EHR source database acquired and a data model determined, mapping the data will be main activity of both data architects and clinicians (Step 2, Fig. 4.1). This step is the longest in the process, so obtaining documentation from the source database will prove helpful and can shorten the time needed. Data architects will need to dive into the specifics of the source database, and work on the Extracting, Transform and Load (ETL) process, and fitting the data in the chosen data model. The clinicians’ role in this stage is to help the data architects in finding the information in the source database, by browsing through the EHR system and identifying where data is stored. The clinicians will also need to validate the data whenever new information is added to the ETL, verifying if the information being stored corresponds with their actual clinical meaning, making each iteration of the data mapping more reliable and consistent. If the clinicians do not work in the source hospital, their expertise will be used to validate the iterations based on whether the given value for each variable is reasonable for its type.
2.2.3 Homologation
If the validation steps during the iterations of data mapping were performed well, the next step, homologation (Step 3, Fig. 4.1), will be short and only require small adjustments to the mapping. Homologation consists of checking to make sure all the mapped data are correct, and have not been corrupted during the ETL process, as a result of improper deletion or modification of information, inclusion of irrelevant and confounding data, and inaccurate verification of correct clinical meaning. During this process, some of the clinicians’ current patients are randomly chosen and information from their latest stay is verified by comparing the information in their medical record to the mapped data. If real time access to the EHR source database was not obtained, this process can be more time consuming as the information from the randomly chosen patients needs to be adapted to the current conditions. If the clinicians on the database development team do not have access to the EHR system, they must homologate the records using their expert knowledge, as they did when validating the data mapping. It is very important that the database development team be thorough during the homologation process, as every piece of mapped information must be checked in order to guarantee the consistency of the data.
2.2.4 Data Pipeline and Quality
After completing the homologation process, the prototype of the database is essentially completed. The next step in the process is to establish the automatic input of the mapped data into a repository, by using a pipeline that assesses data quality (Step 4, Fig. 4.1). The pipeline is made up of sequentially executed computer tasks, scheduled and ordered according to desired intervals and availability of the source database, i.e. real-time or daily, in order to maintain the consistency of the data and the relationships between them. The last and most important task before the final incorporation of the data into the repository must be checking data quality, for example looking for values that differ significantly from current historical data, thereby preventing the inclusion of possibly corrupted data in studies utilizing the database.
2.2.5 De-identification
With the completion of the data pipeline, a usable database is in place. In order to have a clinical database that can be used by other researchers and applications, most institutions and governments require further development to comply with privacy policies and regulations. Additional steps, commonly referred to as de-identification, need to be included in the pipeline (Step 5, Fig. 4.1), in order to produce a database which complies with these requirements. For structured data, i.e. columns of a database, these methods rely on categorizing information, and then deleting or cryptographing the ones flagged as protected. For unstructured data, such as clinicians’ notes, various methods of natural language processing are used, from simple regular expressions, that are pattern matching sequences, to sophisticated neural networks, ultimately trying to identify all protected information throughout the free text for deletion or cryptography (Neamatullah et al. 2008; Dernoncourt et al. 2017). These methods have been included in software and services (Amazon Comprehend Medical 2018) to assist healthcare institutions to comply with patient privacy policies.
2.2.6 Feedback and Correction to Continually Improve and Maintain the Database
After completing the first version of the database, the process is not over. It is essential to understand that constructing a database is a continuous process, relying on continual user feedback to improve and maintain the integrity of the information. The users will have important insights on what can be improved in future versions. The data architect who will be responsible for the maintenance of the database must continually monitor the source database and the pipeline for any possible data corruption. Additional data quality assessments with expert knowledge from clinicians are recommended, who can provide ongoing input regarding whether the data is being properly populated in the database. This can help detect problems in the EHR system, source database or inform directives on how clinicians input information in the EHR system.
References
Aboelsoud, M., Siddique, O., Morales, A., Seol, Y., & Al-Qadi, M. (2018). Early biliary drainage is associated with favourable outcomes in critically-ill patients with acute cholangitis. Przeglad Gastroenterologiczny, 13(1), 16–21.
Amazon Comprehend Medical. Retrieved from December 2018, from https://aws.amazon.com/comprehend/medical/.
Bailly, S., Meyfroidt, G., & Timsit, J. F. (2018). What’s new ICU in 2050: Big data and machine learning. Intensive Care Medicine, 44, 1524–1527.
Block, J. P., Bailey, L. C., Gillman, M. W., Lunsford, D., Boone-Heinonen, J., Cleveland, L. P., et al. (2018). PCORnet antibiotics and childhood growth study: Process for cohort creation and cohort description. Academic Pediatric, 18(5), 569–576.
Collins, F. S., Hudson, K. L., Briggs, J. P., & Lauer, M. S. (2014). PCORnet: Turning a dream into reality. Journal of the American Medical Informatics Association, 21(4), 576–577.
Computing NCfB. (2018). i2b2 (Informatics for Integrating Biology and the Bedside). Retrieved October 2018, from https://www.i2b2.org.
Deliberato, R. O., Ko, S., Komorowski, M., de La Hoz Armengol, M. A., Frushicheva, M.P., & Raffa, J., et al. (2018, March). Severity of illness may misclassify critically ill obese patients. Crit Care, 46(3), 394–400.
Dernoncourt, F., Lee, J. Y., Uzuner, O., & Szolovitz, P. (2017). De-identification of patients notes with recurrent neural networks. Journal of the American Medical Informatics Association, 24(3), 596–606.
Desautels, T., Calvert, J., Hoffman, J., Jay, M., Kerem, Y., Shieh, L., et al. (2016). Prediction of sepsis in the intensive care unit with minimal electronic health record data: A machine learning approach. JMIR Med Inform., 4(3), e28.
Desautels, T., Das, R., Calvert, J., Trivedi, M., Summers, C., Wales, D. J., et al. (2017). Prediction of early unplanned intensive care unit readmission in a UK tertiary care hospital: A cross-sectional machine learning approach. British Medical Journal Open, 7(9), e017199.
Farhan, W., Wang, Z., Huang, Y., Wang, S., Wang, F., & Jiang, X. (2016). A predictive model for medical events based on contextual embedding of temporal sequences. JMIR Medical Informatics, 4(4), e39.
Feng, M., McSparron, J. I., Kien, D. T., Stone, D. J., Roberts, D. H., Schwartzstein, R. M., et al. (2018). Transthoracic echocardiography and mortality in sepsis: Analysis of the MIMIC-III database. Intensive Care Medicine, 44(6), 884–892.
Fleurence, R. L., Curtis, L. H., Califf, R. M., Platt, R., Selby, J. V., & Brown, J. S. (2014). Launching PCORnet, a national patient-centered clinical research network. Journal of the American Medical Informatics Association, 21(4), 578–582.
Ghassemi, M., Marshall, J., Singh, N., Stone, D. J., & Celi, L. A. (2014). Leveraging a critical care database: Selective serotonin reuptake inhibitor use prior to ICU admission is associated with increased hospital mortality. Chest, 145(4), 745–752.
Johnson, A. E., Pollard, T. J., Shen, L., Lehman, L. W., Feng, M., Ghassemi, M., et al. (2016). MIMIC-III, a freely accessible critical care database. Scientific Data, 3, 160035.
Johnson, A. E. W., Aboab, J., Raffa, J., Pollard, T. J., Deliberato, R. O., Celi, L. A., et al. (2018). A comparative analysis of sepsis identification methods in a electronic database. Critical Care, 46(4), 494–499.
Komorowski, M., Celi, L. A., Badawi, O., Gordon, A. C., Faisal, A. A. (2018, October 22). The artificial intelligence clinician learns optimal treatment strategies for sepsis in intensive care. Nat Med. [Epub ahead of print].
List of databases which have been converted to OMOP CDM. (2018). Retrieved October 2018, from http://www.ohdsi.org/web/wiki/doku.php?id=resources:2018_data_network.
Neamatullah, I., Douglas, M. M., Lehman, L. W., Reisner, A., Villarroel, M., Long, W. J., et al. (2008). Automated de-identification of free text medical records. BMC Medical Informatics and Decision Making, 24(8), 32.
Observational Health Data Sciences and Informatics (OHDSI) OMOP Common Data Model V5.0. Retrieved October 2018, from https://www.ohdsi.org.
Pollard, T. J., Johnson, A. E. W., Raffa, J. D., Celi, L. A., Mark, R. G., & Badawi, O. (2018). The eICU collaborative research database, a freely available multi-center database for critical care research. Scientific Data, 5, 180178.
Sanchez-Pinto, L. N., Luo, Y., Churpek, M. M. (2018, May 9). Big data and data science in critical care. Chest pii: S0012-3692(18)30725-6 [Epub ahead of print].
Sandfort,V., Johnson, A. E. W., Kunz, L. M., Vargas, J. D., Rosing, D. R. (2018). Prolonged elevated heart rate and 90-day survival in acutely Ill patients: Data from the MIMIC-III database. Journal of Intensive Care Medicine, 885066618756828.
Serpa Neto, A, Deliberato, R. O., Johnson, A. E. W., Bos, L. D., Amorim, P., Pereira, S. M., et al. (2018, October 5). Mechanical power of ventilation is associated with mortality in critically ill patients: an analysis of patients in two observational cohorts. Intensive Care Medicine [Epub ahead of print].
Waudby-Smith, I. E. R., Tran, N., Dubin, J. A., & Lee, J. (2018). Sentiment in nursing notes as an indicator of out-of-hospital mortality in intensive care patients. PLoS ONE, 13(6), e0198687.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2020 The Author(s)
About this chapter
Cite this chapter
Bulgarelli, L., Núñez-Reiz, A., Deliberato, R.O. (2020). Building Electronic Health Record Databases for Research. In: Celi, L., Majumder, M., Ordóñez, P., Osorio, J., Paik, K., Somai, M. (eds) Leveraging Data Science for Global Health. Springer, Cham. https://doi.org/10.1007/978-3-030-47994-7_4
Download citation
DOI: https://doi.org/10.1007/978-3-030-47994-7_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-47993-0
Online ISBN: 978-3-030-47994-7
eBook Packages: Computer ScienceComputer Science (R0)