Abstract
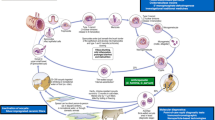
In humans in addition to diarrheal disease and sub-clinical infections the Cryptosporidium parasite can cause developmental delays and growth failure in malnourished infants. We hypothesized that genetic variants may be responsible for the differences in pathogen phenotypes. Resequencing thirty-two Bangladesh C. hominis isolates identified both polymorphic regions and evidence of frequent genetic recombination. This increased the genetic diversity of the parasites. Additional studies are now needed to identify the genetic changes predisposing the parasite to the different phenotypes.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Abrahamsen, M. S., Templeton, T. J., Enomoto, S., Abrahante, J. E., Zhu, G., Lancto, C. A., et al. (2004). Complete genome sequence of the apicomplexan, Cryptosporidium parvum. Science, 304, 441–445. https://doi.org/10.1126/science.1094786.
Acosta, A. M., Chavez, C. B., Flores, J. T., Olotegui, M. P., Pinedo, S. R., Trigoso, D. R., et al. (2014). The MAL-ED study: A multinational and multidisciplinary approach to understand the relationship between enteric pathogens, malnutrition, gut physiology, physical growth, cognitive development, and immune responses in infants and children up to 2 years of age in resource-poor environments. Clinical Infectious Diseases, 59, S193–S206. https://doi.org/10.1093/cid/ciu653.
Alam, G. M. J. (2009). Environmental pollution of Bangladesh—It’s effect and control. In International Conference on Mechanical Engineering 2009, Dhaka, pp. 1–6.
Amadi, B., Mwiya, M., Musuku, J., Watuka, A., Sianongo, S., Ayoub, A., et al. (2002). Effect of nitazoxanide on morbidity and mortality in Zambian children with cryptosporidiosis: A randomised controlled trial. Lancet, 360, 1375–1380. https://doi.org/10.1016/S0140-6736(02)11401-2.
Antony, H., & Parija, S. (2016). Antimalarial drug resistance: An overview. Tropical Parasitoloy, 6, 30. https://doi.org/10.4103/2229-5070.175081.
Bouzid, M., Hunter, P. R., Chalmers, R. M., & Tyler, K. M. (2013). Cryptosporidium pathogenicity and virulence. Clinical Microbiology Reviews, 26, 115–134. https://doi.org/10.1128/CMR.00076-12.
Cama, V. A., Bern, C., Roberts, J., Cabrera, L., Sterling, C. R., Ortega, Y., et al. (2008). Cryptosporidium species and subtypes and clinical manifestations in children, Peru. Emerging Infectious Diseases, 14, 1567–1574. https://doi.org/10.3201/eid1410.071273.
Cama, V. A., Ross, J. M., Crawford, S., Kawai, V., Chavez-Valdez, R., Vargas, D., et al. (2007). Differences in clinical manifestations among Cryptosporidium species and subtypes in HIV-infected persons. The Journal of Infectious Diseases, 196, 684–691. https://doi.org/10.1086/519842.
Checkley, W., White, A. C., Jaganath, D., Arrowood, M. J., Chalmers, R. M., Chen, X.-M., et al. (2015). A review of the global burden, novel diagnostics, therapeutics, and vaccine targets for Cryptosporidium. The Lancet Infectious Diseases, 15, 85–94. https://doi.org/10.1016/S1473-3099(14)70772-8.
Consortium T 1000 GP, 1000 Genomes Project Consortium, Auton, A., Brooks, L. D., Durbin, R. M., Garrison, E. P., Kang, H. M., Korbel, J. O., et al. (2015). A global reference for human genetic variation. Nature, 526, 68–74. https://doi.org/10.1038/nature15393.
Gilchrist, C. A., Cotton, J. A., Burkey, C., Arju, T., Gilmartin, A., Lin, Y., et al. (2018). Genetic diversity of Cryptosporidium hominis in a Bangladeshi community as revealed by whole-genome sequencing. Journal of Infectious Diseases, 218, 259–264. https://doi.org/10.1093/infdis/jiy121.
Guo, Y., Li, N., Lysén, C., Frace, M., Tang, K., Sammons, S., et al. (2015a). Isolation and enrichment of Cryptosporidium DNA and verification of DNA purity for whole-genome sequencing. Journal of Clinical Microbiology, 53, 641–647. https://doi.org/10.1128/JCM.02962-14.
Guo, Y., Tang, K., Rowe, L. A., Li, N., Roellig, D. M., Knipe, K., et al. (2015b). Comparative genomic analysis reveals occurrence of genetic recombination in virulent Cryptosporidium hominis subtypes and telomeric gene duplications in Cryptosporidium parvum. BMC Genomics, 16, 320. https://doi.org/10.1186/s12864-015-1517-1.
Hadfield, S. J., Pachebat, J. A., Swain, M. T., Robinson, G., Cameron, S. J., Alexander, J., et al. (2015). Generation of whole genome sequences of new Cryptosporidium hominis and Cryptosporidium parvum isolates directly from stool samples. BMC Genomics, 16, 650. https://doi.org/10.1186/s12864-015-1805-9.
Haldar, K., Bhattacharjee, S., & Safeukui, I. (2018). Drug resistance in Plasmodium. Nature Reviews Microbiology, 16, 156–170. https://doi.org/10.1038/nrmicro.2017.161.
Ifeonu, O. O., Chibucos, M. C., Orvis, J., Su, Q., Elwin, K., Guo, F., et al. (2016). Annotated draft genome sequences of three species of Cryptosporidium: Cryptosporidium meleagridis isolate UKMEL1, C. baileyi isolate TAMU-09Q1 and C. hominis isolates TU502_2012 and UKH1. Pathogens and Disease, 74, ftw080. https://doi.org/10.1093/femspd/ftw080.
Insulander, M., Silverlås, C., Lebbad, M., Karlsson, L., Mattsson, J. G., & Svenungsson, B. (2013). Molecular epidemiology and clinical manifestations of human cryptosporidiosis in Sweden. Epidemiology and Infection, 141, 1009–1020. https://doi.org/10.1017/S0950268812001665.
Isaza, J. P., Galván, A. L., Polanco, V., Huang, B., Matveyev, A. V., Serrano, M. G., et al. (2015). Revisiting the reference genomes of human pathogenic Cryptosporidium species: Reannotation of C. parvum Iowa and a new C. hominis reference. Scientific Reports, 5, 16324. https://doi.org/10.1038/srep16324.
Islam, S., Begum, H. A., & Nili, N. Y. (2010). Bacteriological safety assessment of municipal tap water and quality of bottle water in Dhaka City: Health hazard analysis. Bangladesh Journal of Medical Microbiology, 4, 9–13. https://doi.org/10.3329/bjmm.v4i1.8462.
Jex, A. R., & Gasser, R. B. (2010). Genetic richness and diversity in Cryptosporidium hominis and C. parvum reveals major knowledge gaps and a need for the application of “next generation” technologies—Research review. Biotechnology Advances, 28, 17–26. https://doi.org/10.1016/j.biotechadv.2009.08.003.
King, P., Tyler, K. M., & Hunter, P. R. (2019). Anthroponotic transmission of Cryptosporidium parvum predominates in countries with poorer sanitation: A systematic review and meta-analysis. Parasites & Vectors, 12, 16. https://doi.org/10.1186/s13071-018-3263-0.
Korpe, P. S., Gilchrist, C. A., Burkey, C., Taniuchi, M., Ahmed, E., Madan, V., et al. (2018). Case-control study of Cryptosporidium transmission in Bangladeshi households. Clinical Infectious Diseases. https://doi.org/10.1093/cid/ciy593.
Korpe, P. S., Haque, R., Gilchrist, C. A., Valencia, C., Niu, F., Lu, M. M., et al. (2016). Natural history of cryptosporidiosis in a longitudinal study of slum-dwelling Bangladeshi children: Association with severe malnutrition. PLoS Neglected Tropical Diseases, 10, e0004564. https://doi.org/10.1371/journal.pntd.0004564.
Korpe, P. S., Liu, Y., Siddique, A., Kabir, M., Ralston, K., Ma, J. Z., et al. (2013). Breast milk parasite-specific antibodies and protection from amebiasis and cryptosporidiosis in Bangladeshi infants: A prospective cohort study. Clinical Infectious Diseases, 56, 988–992. https://doi.org/10.1093/cid/cis1044.
Kosek, M., Alcantara, C., Lima, A. A., & Guerrant, R. L. (2001). Cryptosporidiosis: An update. The Lancet Infectious Diseases, 1, 262–269. https://doi.org/10.1016/S1473-3099(01)00121-9.
Kotloff, K. L., Nataro, J. P., Blackwelder, W. C., Nasrin, D., Farag, T. H., Panchalingam, S., et al. (2013). Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): A prospective, case-control study. Lancet, 382, 209–222. https://doi.org/10.1016/S0140-6736(13)60844-2.
Lequime, S., Fontaine, A., Ar Gouilh, M., Moltini-Conclois, I., & Lambrechts, L. (2016). Genetic drift, purifying selection and vector genotype shape dengue virus intra-host genetic diversity in mosquitoes. PLoS Genetics, 12, e1006111. https://doi.org/10.1371/journal.pgen.1006111.
Li, N., Xiao, L., Cama, V. A., Ortega, Y., Gilman, R. H., Guo, M., et al. (2013). Genetic recombination and Cryptosporidium hominis virulent subtype IbA10G2. Emerging Infectious Diseases, 19, 1573–1582. https://doi.org/10.3201/eid1910.121361.
Manjunatha, U. H., Chao, A. T., Leong, F. J., & Diagana, T. T. (2016). Cryptosporidiosis drug discovery: Opportunities and challenges. ACS Infectious Diseases, 2, 530–537. https://doi.org/10.1021/acsinfecdis.6b00094.
Manjunatha, U. H., Vinayak, S., Zambriski, J. A., Chao, A. T., Sy, T., Noble, C. G., et al. (2017). A Cryptosporidium PI(4)K inhibitor is a drug candidate for cryptosporidiosis. Nature, 546, 376–380. https://doi.org/10.1038/nature22337.
Mu, J., Awadalla, P., Duan, J., McGee, K. M., Joy, D. A., McVean, G. A. T., et al. (2005). Recombination hotspots and population structure in Plasmodium falciparum. PLoS Biology, 3, e335. https://doi.org/10.1371/journal.pbio.0030335.
Nader, J. L., Mathers, T. C., Ward, B. J., Pachebat, J. A., Swain, M. T., Robinson, G., et al. (2019). Evolutionary genomics of anthroponosis in Cryptosporidium. Nature Microbiology. https://doi.org/10.1038/s41564-019-0377-x.
Pal, S., Bhattacharya, S. K., Das, P., Chaudhuri, P., Dutta, P., De, S. P., et al. (1989). Occurrence and significance of Cryptosporidium infection in Calcutta. Transactions of the Royal Society of Tropical Medicine and Hygiene, 83, 520–521.
Pickering, A. J., Crider, Y., Sultana, S., Swarthout, J., Goddard, F. G., Anjerul Islam, S., et al. (2019). Effect of in-line drinking water chlorination at the point of collection on child diarrhoea in urban Bangladesh: A double-blind, cluster-randomised controlled trial. The Lancet Global Health, 7, e1247–e1256. https://doi.org/10.1016/S2214-109X(19)30315-8.
Platts-Mills, J. A., McCormick, B. J. J., Kosek, M., Pan, W. K., Checkley, W., Houpt, E. R., et al. (2014). Methods of analysis of enteropathogen infection in the MAL-ED cohort study. Clinical Infectious Diseases, 59, S233–S238. https://doi.org/10.1093/cid/ciu408.
Prüss-Ustün, A., Wolf, J., Bartram, J., Clasen, T., Cumming, O., Freeman, M. C., et al. (2019). Burden of disease from inadequate water, sanitation and hygiene for selected adverse health outcomes: An updated analysis with a focus on low- and middle-income countries. International Journal of Hygiene and Environmental Health, 222, 765–777. https://doi.org/10.1016/j.ijheh.2019.05.004.
Pumipuntu, N., & Piratae, S. (2018). Cryptosporidiosis: A zoonotic disease concern. Veterinary World, 11, 681–686. https://doi.org/10.14202/vetworld.2018.681-686.
Rafaluk, C., Gildenhard, M., Mitschke, A., Telschow, A., Schulenburg, H., & Joop, G. (2015). Rapid evolution of virulence leading to host extinction under host-parasite coevolution. BMC Evolutionary Biology, 15, 112. https://doi.org/10.1186/s12862-015-0407-0.
Rossignol, J., Kabil, S. M., El-Gohary, Y., & Younis, A. M. (2006). Effect of nitazoxanide in diarrhea and enteritis caused by Cryptosporidium species. Clinical Gastroenterology and Hepatology, 4, 320–324. https://doi.org/10.1016/j.cgh.2005.12.020.
Rossle, N. F., & Latif, B. (2013). Cryptosporidiosis as threatening health problem: A review. Asian Pacific Journal of Tropical Biomedicine, 3, 916–924. https://doi.org/10.1016/S2221-1691(13)60179-3.
Sikora, P., Andersson, S., Winiecka-Krusnell, J., Hallström, B., Alsmark, C., Troell, K., et al. (2017). Genomic variation in IbA10G2 and other patient-derived Cryptosporidium hominis subtypes. Journal of Clinical Microbiology, 55, 844–858. https://doi.org/10.1128/JCM.01798-16.
Sirajul Islam, M., Brooks, A., Kabir, M. S., Jahid, I. K., Shafiqul Islam, M., Goswami, D., et al. (2007). Faecal contamination of drinking water sources of Dhaka city during the 2004 flood in Bangladesh and use of disinfectants for water treatment. Journal of Applied Microbiology, 103, 80–87. https://doi.org/10.1111/j.1365-2672.2006.03234.x.
Sow, S. O., Muhsen, K., Nasrin, D., Blackwelder, W. C., Wu, Y., Farag, T. H., et al. (2016). The burden of Cryptosporidium diarrheal disease among children <24 months of age in moderate/high mortality regions of Sub-Saharan Africa and South Asia, utilizing data from the global enteric multicenter study (GEMS). PLoS Neglected Tropical Diseases, 10, e0004729. https://doi.org/10.1371/journal.pntd.0004729.
Steiner, K. L., Ahmed, S., Gilchrist, C. A., Burkey, C., Cook, H., Ma, J. Z., et al. (2018). Species of cryptosporidia causing subclinical infection associated with growth faltering in rural and urban Bangladesh: A birth cohort study. Clinical Infectious Diseases, 67, 1347–1355. https://doi.org/10.1093/cid/ciy310.
U.S. Department of Health and Human Services. (2003). New drug for parasitic infections in children. FDA Consumer, 37, 4.
Van den Bergh, B., Swings, T., Fauvart, M., & Michiels, J. (2018). Experimental design, population dynamics, and diversity in microbial experimental evolution. Microbiology and Molecular Biology Reviews, 82. https://doi.org/10.1128/MMBR.00008-18.
Wang, N., Akey, J. M., Zhang, K., Chakraborty, R., & Jin, L. (2002). Distribution of recombination crossovers and the origin of haplotype blocks: The interplay of population history, recombination, and mutation. American Journal of Human Genetics, 71, 1227–1234. https://doi.org/10.1086/344398.
Widmer, G., & Lee, Y. (2010). Comparison of single- and multilocus genetic diversity in the protozoan parasites Cryptosporidium parvum and C. hominis. Applied and Environment Microbiology, 76, 6639–6644. https://doi.org/10.1128/AEM.01268-10.
Xu, P., Widmer, G., Wang, Y., Ozaki, L. S., Alves, J. M., Serrano, M. G., et al. (2004). The genome of Cryptosporidium hominis. Nature, 431, 1107–1112. https://doi.org/10.1038/nature02977.
Working Group on Waterborne Cryptosporidiosis. (1995). Cryptosporidium and Water: A Public Health Handbook. Georgia: Atlanta.
Acknowledgements
We are grateful for the willing participation of the parents and children at the icddr,b study sites and wish to thank the field workers, nurses, laboratory staff of the Parasitology Laboratory of icddr,b without whom we could not have completed this research.
Conflict of Interest Statement
The author has no conflict of interest to disclose.
Funding
The work mentioned in this chapter was supported by the National Institute of Allergy and Infectious Diseases (NIAID) R21 AI142656 and R01 AI-043596, the Bill and Melinda Gates Foundation Grant (OPP1100514). Sequencing was at the Wellcome Trust Sanger Institute which is supported by the Wellcome Trust. Work at icddr,b is supported by the core donors (Government of the People’s Republic of Bangladesh, GAC, Sida, UKAid). The funders had no participation in the writing of this chapter or the decision to publish.
Ethical Considerations
The work reported here as reviewed by the Ethical and Research Review Committees of the International Centre for Diarrhoeal Disease Research, Bangladesh and by the Institutional Review Board of the University of Virginia. Informed written consent was obtained from the parents or guardians for the participation of their child in the study.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this paper
Cite this paper
Gilchrist, C.A. (2020). Cryptosporidium Infection in Bangladesh Children. In: Guillen, N. (eds) Eukaryome Impact on Human Intestine Homeostasis and Mucosal Immunology. Springer, Cham. https://doi.org/10.1007/978-3-030-44826-4_7
Download citation
DOI: https://doi.org/10.1007/978-3-030-44826-4_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-44825-7
Online ISBN: 978-3-030-44826-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)