Abstract
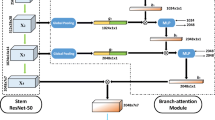
Accurate and automatic analysis of breast MRI plays a vital role in early diagnosis and successful treatment planning for breast cancer. Due to the heterogeneity nature, precise diagnosis of tumors remains a challenging task. In this paper, we propose to identify breast tumor in MRI by Cosine Margin Sigmoid Loss (CMSL) with deep learning (DL) and localize possible cancer lesion by COrrelation Attention Map (COAM) based on the learned features. The CMSL embeds tumor features onto a hyper-sphere and imposes a decision margin through cosine constraints. In this way, the DL model could learn more separable inter-class features and more compact intra-class features in the angular space. Furthermore, we utilize the correlations among feature vectors to generate attention maps that could accurately localize cancer candidates with only image-level labels. We build the largest breast cancer dataset involving 10,290 DCE-MRI scan volumes for developing and evaluating the proposed methods. The model driven by CMSL achieved a classification accuracy of 0.855 and AUC of 0.902 on the testing set, with sensitivity and specificity of 0.857 and 0.852, respectively, outperforming competitive methods overall. In addition, the proposed COAM accomplished more accurate localization of the cancer center compared with other state-of-the-art weakly supervised localization method.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
DeSantis, C.E., et al.: Breast cancer statistics, racial disparity in mortality by state. CA Cancer J. Clin. 67(6), 439–448 (2017)
Kuhl, C., et al.: Prospective multicenter cohort study to refine management recommendations for women at elevated familial risk of breast cancer: the EVA trial. J. Clin. Oncol. 28(9), 1450–1457 (2010)
Zheng, H., Gu, Y., Qin, Y., Huang, X., Yang, J., Yang, G.-Z.: Small lesion classification in dynamic contrast enhancement MRI for breast cancer early detection. In: Frangi, A.F., Schnabel, J.A., Davatzikos, C., Alberola-López, C., Fichtinger, G. (eds.) MICCAI 2018. LNCS, vol. 11071, pp. 876–884. Springer, Cham (2018). https://doi.org/10.1007/978-3-030-00934-2_97
Amit, G., et al.: Classification of breast MRI lesions using small-size training sets: comparison of deep learning approaches. In: Medical Imaging 2017: Computer-Aided Diagnosis, vol. 10134. International Society for Optics and Photonics (2017)
Amit, G., et al.: Hybrid mass detection in breast MRI combining unsupervised saliency analysis and deep learning. In: Descoteaux, M., Maier-Hein, L., Franz, A., Jannin, P., Collins, D.L., Duchesne, S. (eds.) MICCAI 2017. LNCS, vol. 10435, pp. 594–602. Springer, Cham (2017). https://doi.org/10.1007/978-3-319-66179-7_68
Maicas, G., Bradley, A.P., Nascimento, J.C., Reid, I., Carneiro, G.: Training medical image analysis systems like radiologists. In: Frangi, A.F., Schnabel, J.A., Davatzikos, C., Alberola-López, C., Fichtinger, G. (eds.) MICCAI 2018. LNCS, vol. 11070, pp. 546–554. Springer, Cham (2018). https://doi.org/10.1007/978-3-030-00928-1_62
Wang, H., et al.: CosFace: large margin cosine loss for deep face recognition. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (2018)
Zhou, B., et al.: Learning deep features for discriminative localization. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (2016)
Gatys, L., Ecker, A.S., Bethge, M.: Texture synthesis using convolutional neural networks. In: Advances in Neural Information Processing Systems, pp. 262–270 (2015)
Fu, J., et al.: Dual attention network for scene segmentation. arXiv preprint arXiv:1809.02983 (2018)
Frangi, A.F., Niessen, W.J., Vincken, K.L., Viergever, M.A.: Multiscale vessel enhancement filtering. In: Wells, W.M., Colchester, A., Delp, S. (eds.) MICCAI 1998. LNCS, vol. 1496, pp. 130–137. Springer, Heidelberg (1998). https://doi.org/10.1007/BFb0056195
He, K., et al.: Deep residual learning for image recognition. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 770–778 (2016)
Wu, J., et al.: Deep multiple instance learning for image classification and auto-annotation. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (2015)
Zhu, W., Lou, Q., Vang, Y.S., Xie, X.: Deep multi-instance networks with sparse label assignment for whole mammogram classification. In: Descoteaux, M., Maier-Hein, L., Franz, A., Jannin, P., Collins, D.L., Duchesne, S. (eds.) MICCAI 2017. LNCS, vol. 10435, pp. 603–611. Springer, Cham (2017). https://doi.org/10.1007/978-3-319-66179-7_69
Liu, J., et al.: Integrate domain knowledge in training CNN for ultrasonography breast cancer diagnosis. In: Frangi, A.F., Schnabel, J.A., Davatzikos, C., Alberola-López, C., Fichtinger, G. (eds.) MICCAI 2018. LNCS, vol. 11071, pp. 868–875. Springer, Cham (2018). https://doi.org/10.1007/978-3-030-00934-2_96
Acknowledgement
This work was supported by Research Grants Council of Hong Kong Special Administrative Region under Project No. CUHK14225616 and Hong Kong Innovation and Technology Fund under Project No. ITS/426/17FP.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Luo, L. et al. (2019). Deep Angular Embedding and Feature Correlation Attention for Breast MRI Cancer Analysis. In: Shen, D., et al. Medical Image Computing and Computer Assisted Intervention – MICCAI 2019. MICCAI 2019. Lecture Notes in Computer Science(), vol 11767. Springer, Cham. https://doi.org/10.1007/978-3-030-32251-9_55
Download citation
DOI: https://doi.org/10.1007/978-3-030-32251-9_55
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-32250-2
Online ISBN: 978-3-030-32251-9
eBook Packages: Computer ScienceComputer Science (R0)