Abstract
Glioblastoma is the most aggressive malignant primary brain tumor with a poor prognosis. Glioblastoma heterogeneous neuroimaging, pathologic, and molecular features provide opportunities for subclassification, prognostication, and the development of targeted therapies. Magnetic resonance imaging has the capability of quantifying specific phenotypic imaging features of these tumors. Additional insight into disease mechanism can be gained by exploring genetics foundations. Here, we use the gene expressions to evaluate the associations with various quantitative imaging phenomic features extracted from magnetic resonance imaging. We highlight a novel correlation by carrying out multi-stage genome-wide association tests at the gene-level through a non-parametric correlation framework that allows testing multiple hypotheses about the integrated relationship of imaging phenotype-genotype more efficiently and less expensive computationally. Our result showed several novel genes previously associated with glioblastoma and other types of cancers, as the LRRC46 (chromosome 17), EPGN (chromosome 4) and TUBA1C (chromosome 12), all associated with our radiographic tumor features.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Gliomas are the most common type of primary adult brain tumors that arise from glial cells. Gliomas have a very heterogeneous landscape, and they can be classified according to their grade into low-grade glioma, anaplastic glioma, and glioblastoma. The most common and aggressive type of glioma in adults is glioblastoma (GBM), which gives to the affected patient average survival time of only 10 to 18 months. The known molecular classification of GBM into classical, mesenchymal, neural and proneural subtypes is relatively accepted to be related to the expression of EGFR, NF1 and PDGFRA/IDH1 genes [1].
Imaging, specifically magnetic resonance imaging (MRI), can offer data towards promising biomarkers reflecting underlying tumor pathology and biological function. If imaging phenotypes of GBM obtained from routine clinical MRI studies can be associated with specific gene expression signatures, quantitative imaging phenotypes will serve as non-invasive surrogates for cancer genomic events and provide valuable information as to the diagnosis, prognosis, and optimal treatment.
Several radiogenomic studies have been carried out for many diseases [8,9,10,11,12,13,14,15,16]. For instance for schizophrenia pairs of SNP/Gene and MRI features have been mapped by using PLINK [8], and Parallel-ICA showed promising results [9]. Batmanghelich et al. [10] proposed a Bayesian framework to relate imaging and genetic data to phenotypes exploiting connection among these data modalities simultaneously in Alzheimer. Recently, correlations of connectomic features have been related to genes which are known to be related to Alzheimer progression [11]. In contrast to Alzheimer’s disease and schizophrenia, glioma lesions are generally not spread all over the brain, and local features from MRI can be used. An imaging-genomic analysis study [12], performed by using the tumor volume in T2-weighted FLuid-Attenuated Inversion Recovery (T2-FLAIR) images and large-scale genetic and micro-RNA expression probes demonstrated the potential for molecular subtyping and showed that the high median expression of POSTN gene results in a significant decrease in survival, and for that they used ANOVA and Tukey-Kramer test. Other studies [13, 14] showed correlations between image feature annotations and expression of genes with glioma molecular subtypes [1]. Specifically, Gutman et al. [13] found a significant association between contrast-enhanced tumor and these molecular subtypes [1], where proneural type expressed by PDGFRA/IDH1 gene showed low levels of contrast enhancement, and the classical type (i.e., primarily described by EGFR amplification) correlates with the increased percentage of contrast enhancement. The study used sher exact statistics.
Recent population-based studies have assessed the anatomical location of GBM in relation to distinct clinically-relevant molecular characteristics, and have identified the spatial distribution of the tumors being descriptive of their molecular status [14, 17,18,19,20,21,22]. Furthermore, the emerging research direction of radiomics has shown promise that texture analysis of the various tumor sub-regions in radiographic imaging can also be informative of the tumor’s molecular characterization [23,24,25]. Furthermore, using MRI features for GBM lesions, including texture and shape features, Haruka et al. proposed a classification imaging method and found three clusters of GBM patients [35]. In their method, they integrate copy number and gene expression data to estimate the molecular pathway activity and show that the three clusters reveal not only different molecular characteristics but also different survival probabilities.
The purpose of this paper is to identify significant associations between gene expressions, across the whole genome, and quantitative imaging phenomic features extracted from multi-modal MRI brain scans of patients diagnosed with de novo primary GBM. In line with the pre-mentioned studies, here we focus on evaluating the spatial location and texture features of GBM and investigate their associations with gene expressions.
2 Materials and Methods
2.1 Data
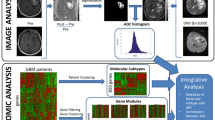
For the quantitative association analysis conducted here, we utilized a retrospective cohort of 135 de novo primary GBM patients from the TCGA-GBM collection [6], with available pre-operative multi-modal MRI scans in The Cancer Imaging Archive (TCIA) [7] and corresponding molecular characterization in The Cancer Genome Atlas (TGCA). The multi-modal MRI data we utilized comprise native (T1) and post-contrast T1-weighted (T1Gd), T2-weighted (T2), and T2-FLAIR modalities. The TCGA-GBM subset of 135 patients were identified by Bakas et al. [4] as brain scans without any surgically-imposed cavity, and their co-registered and skull-stripped imaging were provided in the TCIA Analysis Results together with expert manually annotated segmentation labels for the various histologically-distinct tumor sub-regions, i.e. enhancing tumor (ET), non-enhancing tumor (NET), peritumoral edematous/invaded tissue (ED) (Fig. 1) [4, 5]. The total sample size of GBM patients reduced to 88 after evaluating patients that had available imaging [6] and corresponding gene expressions. In total, we assessed expression energies for 17815 genes, 11 distinct descriptors of tumor spatial location (Fig. 2), and 517 radiomic/texture features (Fig. 2) for each patient’s brain tumor scan [2, 4, 5].
2.2 Quantitative Imaging Phenomic Features
Radiomic/Texture Features. We extracted an extensive panel of quantitative texture features, volumetrically (in 3D), for each tumor sub-region as provided by the expert annotations, across all available modalities. Specifically, the texture features we evaluated (i) capture global characteristics (i.e., variance, skewness, kurtosis) of each sub-region’s intensity distribution on each modality, and (ii) include features based on Gray-Level Co-occurrence Matrix (GLCM) [26] (Fig. 2), Gray-Level Run-Length Matrix (GLRLM) [27,28,29,30], Gray-Level Size Zone Matrix (GLSZM) [28,29,30], and Neighborhood Gray-Tone Difference Matrix (NGTDM) [31].
Spatial Distribution Patterns. Beyond texture features, we collected discrete spatial information about the anatomical location of each tumor on each brain scan (Fig. 2). To obtain these spatial distribution patterns we registered all brain tumor scans in a standardized healthy atlas space using an iterative Expectation-Maximization framework [3], while incorporating a biophysical tumor growth model (based on a reaction-diffusion-advection model [32,33,34]) to account for tumor mass effects in the brain parenchyma. We then retrieved the spatial distribution of each tumor according to the discretized anatomical locations of the (i) specific lobes (i.e., frontal, temporal, parietal, occipital), (ii) insula, (iii) basal ganglia, (iv) fornix, (v) cerebellum, and (vi) brain stem. In addition, we also included as distinct features the distances of (i) the tumor core (defined as the union of ET and NET), and (ii) the ED, from the ventricles.
To produce these quantitative features we have utilized GLISTRboost. Specifically, in the process to produce segmentations of the various tumor sub-regions, the generative part [37] of GLISTRboost, following an Expectation-Maximization framework registers a healthy population probabilistic atlas to glioma patients’ brain scans while incorporating a biophysical glioma growth model to account for mass effects. Then, after converting the predicted segmentation in the healthy atlas space, the percentage of the tumor core (i.e., enhancing and non-enhancing tumor) is calculated on each of the brain lobes in this healthy atlas.
2.3 Data Analysis
Initially, we combined the two types of data (imaging - genetics) using the patient ID as a primary column. As a first stage, we used the gene expressions and the spatial distribution patterns to perform a non-parametric test of association. To assess the associations, we computed the Spearman correlation coefficient (\(r_s\)) between the gene expressions, individually, as a with each of the spatial distribution patterns described in Sect. 2.2. We then assessed the significant of the correlation coefficient by calculating the p-values as described below.
For each quantitative feature and each gene, We obtained the p-value associated with Spearman correlation coefficient test statistic. That is, the p-value of the correlation between a single gene expression with a single feature of the tumor’s location in the brain. The Spearman correlation coefficient model for a given feature (y) and given gene expression (x) is;
Where \(d_i\) is the difference between the ranks of \(x_i\) and \(y_i\), and N is 88; representing the number of GBM patients [38]. \(r_s\) can take any real value between \(+1\) and \(-1\); \(+1\) represents a strong positive association, \(-1\) means a perfect negative association and 0 indicates no association between the ranks of x and y. Our hypothesis of interest is:
-
\(H_0\): There is no association between the gene expression and the tumor’s feature under study
$$\begin{aligned} \text {vs} \end{aligned}$$ -
\(H_a\): There is an association between the gene expression and the tumor’s feature under study, alternatively:
-
\(H_0\): \(r_s\) = 0 vs \(H_0\): \(r_s \ne 0\)
To determine the significance of \(r_s\), one can use the t test statistic defined as
\(t_c\) follows approximately the Student’s t distribution with a \(N - 2\) degrees of freedom under the null hypothesis [38]. At a certain significance level, the calculated value of \(t_c\) can be compared to the table value obtained from the Student’s t distribution (as described previously). The significance of \(r_s\) can also be determined using the p-value, which is simply the integration, or the area under the curve from \(t_c\) to infinity.
Briefly, in this first stage, the association test was initially conducted to six features of the tumor location (Sect. 2.2). More specifically, for each gene, we computed six p-values, then considered only the minimum p-value at each gene (see Fig. 3 for the analysis workflow). The latter is referred to as meta-analysis in Fig. 3 (step(c)). All results reported in Sect. 3 use the summary statistics of the meta-analysis. Moreover, out of the all the association results, we excluded all the genes with p-values greater than or equal 0.05. Here we meant to exclude the genes that have very low (and not significant) association with the spatial pattern, which we believe is an important phenotype. This step is referred to as (d) in Fig. 3. In the second stage, we proceeded with all the genes with p-value less than 0.05, excluding the least significant genes, and we carried the same analysis as in the first stage but using the radiomic features (Sect. 2.2. Table 1 shows the thresholds at both 5% and 10% significance level), along with the number of genes used and remained in each stage.
It is worth mentioning that, out of the total number of genes, we were able to annotate 15009 genes and assign them to their defined physical locations in the DNA. We carried on the first stage of the analysis using those genes (Table 1).
3 Results
The incidence of tumors specific for region is summarized in Table 2. The Manhattan plot for the p-values obtained from the meta-analysis is illustrated in Fig. 4. The plot shows two horizontal lines which associate with the thresholds of 5% significance level (top line), and 10% significance level (bottom line), after correcting for multiple comparisons. The x-axis is the physical position of genes in the DNA, and the y-axis is the negative log10 of the p-values. Figure 4 also shows the qq-plot of all the genes used in the association analysis. Likewise, each dot corresponds to a p-value of a single gene and \(- log10\) of the p-value is used instead. The qq-plot reported with each Manhattan plot, and it compares the observed distribution of p-values (y-axis) to the expected distribution (x-axis), for each gene tested, where the diagonal line is the null distribution.
Table 3 shows (only) the highest ten p-values and the corresponding genes of the first stage of the analysis. In this stage, non of the p-values was less than \(3.3e^{-6}\) or \(6.7e^{-6}\) (see Table 1); therefore, no gene was significantly associated with any of the features. Table 3 reports the gene symbol, its start and end position, the associated p-value and feature, and the chromosome.
We then pruned the genes used in the previous stage to a smaller set, by removing the genes that have p-values less than 0.05. With the 5401 genes remaining, we took over the second stage and repeated the same analysis with the texture characteristics of the tumor. The Manhattan and qq-plot for the texture features are shown in Fig. 5, and Table 4 shows the top 10 significant genes. Total of significant genes in this stage is 37 (at 5% significance level).
4 Discussion
GBM is a fatal malignant disease that so far is incurable. The identification of genetic risk factors that affect the tumor characteristics improves our understanding of the underlying biological processes for GBM, and contribute to therapeutic discovery. In this study, we proposed a framework that allows quantifying the non-parametric correlations to test associations between gene expressions and different quantitative imaging phenomic characteristics of GBM. Our result has shown a high genetic enrichment through the Manhattan and qq-plots, especially for the texture features (Fig. 5).
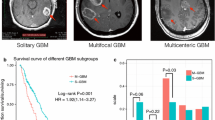
Our results highlighted several genes that significantly associated with the tumor texture features, including LRRC46, USP38, EPGN, TUBA1C, ZNF284, IPO8, MMP7, TLL2, TRIM55 and UBAP1, as the top ten significant genes (Table 4). However, there are, in total, 37 genes are significantly associated with the texture features (Fig. 5). EPGN expression associates significantly (\(r_s=0.501\), p-value\(\,=6.542e{-}07\)) with GLSZM LGZE in the T1Gd modality (Table 4). EPGN previously reported to be one of the top ten upregulated genes after EBLN1 silencing in oligodendroglia cells [39]. Moreover, the emergence of EPGN was marked in another study by Duhem-Tonnelle et al. in EGF ligands expression profile, between glioblastoma cell lines and biopsies [40]. Located at chromosome 4, USP38 (\(r_s=-0.511\), p-value \(=3.648e{-}07\)) [41]. Moreover, as it is illustrated in the Manhattan plot of the spatial features of the tumor (Fig. 4 and Table 3), no gene shows significant association with any of the location features. In addition to the latter, the number of GBM lesions in the cerebellum in clinical settings are quite rare [36], as also shown in our summary Table 2. Our study can give some insight into this rare type of GBM lesion. Nevertheless, the investigation excluding the patients having those lesions have to be repeated as a future work.
5 Conclusion
As the understanding of gliomagenesis grows, several medical imaging biomarkers and genetic variations can be identified, and new hypotheses can be formed. The hereby proposed genome-wide association framework aims at identifying differentially expressed genes that significantly correlate with various aspects of GBM. The identification of such genes may contribute to the development of targeted therapies that focus on the resistance mechanisms of individual patients.
Through the systematic testing of associations and shrinking of the number of genes at every stage, this pipeline facilitates the evaluation of various hypotheses and reduces the computational complexity. In future work, we plan to extend the study by integrating more quantitative imaging phenomic tumor characteristics, inclusive of morphological, intensity, and volumetric descriptors, as well as parameters derived by biophysical tumor growth modeling.
References
Verhaak, R.G.W., et al.: Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17(1), 98–110 (2010)
Davatzikos, C., et al.: Cancer imaging phenomics toolkit: quantitative imaging analytics for precision diagnostics and predictive modeling of clinical outcome. J. Med. Imaging 5(1), 011018 (2018)
Bakas, S., et al.: GLISTRboost: combining multimodal MRI segmentation, registration, and biophysical tumor growth modeling with gradient boosting machines for glioma segmentation. In: Crimi, A., Menze, B., Maier, O., Reyes, M., Handels, H. (eds.) BrainLes 2015. LNCS, vol. 9556, pp. 144–155. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-30858-6_13
Bakas, S., et al.: Advancing the cancer genome atlas glioma MRI collections with expert segmentation labels and radiomic features. Nat. Sci. Data 4, 170117 (2017)
Bakas, S., et al.: Segmentation labels and radiomic features for the pre-operative scans of the TCGA-GBM collection. The Cancer Imaging Archive (2017). https://doi.org/10.7937/K9/TCIA.2017.KLXWJJ1Q
Scarpace, L., et al.: Radiology data from the cancer genome atlas glioblastoma multiforme [TCGA-GBM] collection. Cancer Imaging Arch. 11, 4 (2016)
Clark, K., et al.: The Cancer Imaging Archive (TCIA): maintaining and operating a public information repository. J. Digit. Imaging 26(6), 1045–1057 (2013)
Stein, J.L., et al.: Voxelwise genome-wide association study (vGWAS). Neuroimage 53(3), 1160–1174 (2010)
Liu, J., et al.: Combining fMRI and SNP data to investigate connections between brain function and genetics using parallel ICA. Hum. Brain Mapp. 30(1), 241–255 (2009)
Batmanghelich, N.K., Dalca, A.V., Sabuncu, M.R., Golland, P.: Joint modeling of imaging and genetics. In: Gee, J.C., Joshi, S., Pohl, K.M., Wells, W.M., Zöllei, L. (eds.) IPMI 2013. LNCS, vol. 7917, pp. 766–777. Springer, Heidelberg (2013). https://doi.org/10.1007/978-3-642-38868-2_64
Elsheikh, S., et al.: Relating connectivity changes in brain networks to genetic information in Alzheimer patients. In: 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018). IEEE (2018)
Zinn, P.O., et al.: Radiogenomic mapping of edema/cellular invasion MRI-phenotypes in glioblastoma multiforme. PLoS ONE 6(10), e25451 (2011)
Gutman, D.A., et al.: MR imaging predictors of molecular profile and survival: multi-institutional study of the TCGA glioblastoma data set. Radiology 267(2), 560–569 (2013)
Macyszyn, L., et al.: Imaging patterns predict patient survival and molecular subtype in glioblastoma via machine learning techniques. Neuro-oncology 18(3), 417–425 (2015)
Binder, Z., et al.: Epidermal growth factor receptor extracellular domain mutations in glioblastoma present opportunities for clinical imaging and therapeutic development. Cancer Cell 34, 163–177 (2018)
Bakas, S., et al.: In vivo detection of EGFRvIII in glioblastoma via perfusion magnetic resonance imaging signature consistent with deep peritumoral in ltration: the \(\varphi \)-index. Clin. Cancer Res. 23, 4724–4734 (2017)
Cancer Genome Atlas Research Network: Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N. Engl. J. Med. 372(26), 2481–2498 (2015)
Ellingson, B.M., et al.: Probabilistic radiographic atlas of glioblastoma phenotypes. Am. J. Neuroradiol. 34(3), 533–540 (2012)
Ellingson, B.M.: Radiogenomics and imaging phenotypes in glioblastoma: novel observations and correlation with molecular characteristics. Curr. Neurol. Neurosci. Rep. 15(1), 506 (2015)
Steed, T.C., et al.: Differential localization of glioblastoma subtype: implications on glioblastoma pathogenesis. Oncotarget 7(18), 24899 (2016)
Bilello, M., et al.: Population-based MRI atlases of spatial distribution are specific to patient and tumor characteristics in glioblastoma. NeuroImage: Clin. 12, 34–40 (2016)
Akbari, H., et al.: In vivo evaluation of EGFRvIII mutation in primary glioblastoma patients via complex multiparametric MRI signature. Neuro-Oncology 20(8), 1068–1079 (2018)
Aerts, H.J.W.L.: The potential of radiomic-based phenotyping in precision medicine: a review. JAMA Oncol. 2(12), 1636–1642 (2016)
Lambin, P., et al.: Radiomics: extracting more information from medical images using advanced feature analysis. Eur. J. Cancer 48(4), 441–446 (2012)
Aerts, H.J.W.L., et al.: Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat. Commun. 5, 4006 (2014)
Haralick, R.M., et al.: Textural features for image classification. IEEE Trans. Syst. Man Cybern. 3, 610–621 (1973)
Galloway, M.M.: Texture analysis using grey level run lengths. Comput. Graph. Image Process. 4, 172–179 (1975)
Chu, A., et al.: Use of gray value distribution of run lengths for texture analysis. Pattern Recogn. Lett. 11, 415–419 (1990)
Dasarathy, B.V., Holder, E.B.: Image characterizations based on joint gray level-run length distributions. Pattern Recogn. Lett. 12, 497–502 (1991)
Tang, X.: Texture information in run-length matrices. IEEE Trans. Image Process. 7, 1602–1609 (1998)
Amadasun, M., King, R.: Textural features corresponding to textural properties. IEEE Trans. Syst. Man Cybern. 19, 1264–1274 (1989)
Hogea, C., et al.: An image-driven parameter estimation problem for a reaction-diffusion glioma growth model with mass effects. J. Math. Biol. 56, 793–825 (2008)
Hogea, C., et al.: A robust framework for soft tissue simulations with application to modeling brain tumor mass effect in 3D MR images. Phys. Med. Biol. 52, 6893–6908 (2007)
Hogea, C., Davatzikos, C., Biros, G.: Modeling glioma growth and mass effect in 3D MR images of the brain. In: Ayache, N., Ourselin, S., Maeder, A. (eds.) MICCAI 2007. LNCS, vol. 4791, pp. 642–650. Springer, Heidelberg (2007). https://doi.org/10.1007/978-3-540-75757-3_78
Itakura, H., et al.: Magnetic resonance image features identify glioblastoma phenotypic subtypes with distinct molecular pathway activities. Sci. Transl. Med. 7(303), 303ra138 (2015)
Drabycz, S., et al.: An analysis of image texture, tumor location, and MGMT promoter methylation in glioblastoma using magnetic resonance imaging. Neuroimage 49(2), 1398–1405 (2010)
Gooya, A., et al.: GLISTR: glioma image segmentation and registration. IEEE Trans. Med. Imaging 31(10), 1941–1954 (2012)
Kendall, M.G.: The advanced theory of statistics. In: The Advanced Theory of Statistics, 2nd edn (1946)
He, P., et al.: Knock-down of endogenous bornavirus-like nucleoprotein 1 inhibits cell growth and induces apoptosis in human oligodendroglia cells. Int. J. Mol. Sci. 17(4), 435 (2016)
Duhem-Tonnelle, V., et al.: Differential distribution of erbB receptors in human glioblastoma multiforme: expression of erbB3 in CD133-positive putative cancer stem cells. J. Neuropathol. Exp. Neurol. 69(6), 606–622 (2010)
Carminati, P.O., et al.: Alterations in gene expression profiles correlated with cisplatin cytotoxicity in the glioma U343 cell line. Genet. Mol. Biol. 33(1), 159–168 (2010)
Acknowledgement
Research reported in this publication was partly supported by the National Institutes of Health (NIH) under award numbers NIH/NINDS:R01NS042645, NIH/NCI:U24CA189523, NIH/NCATS:UL1TR001878, the ITMAT of the University of Pennsylvania as well as by the Swedish International Development Cooperation Agency (SIDA) through the Organization for Women in Science for the Developing World (OWSD). Computations were performed using facilities provided by the University of Cape Town’s ICTS High Performance Computing team: hpc.uct.ac.za.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Elsheikh, S.S.M., Bakas, S., Mulder, N.J., Chimusa, E.R., Davatzikos, C., Crimi, A. (2019). Multi-stage Association Analysis of Glioblastoma Gene Expressions with Texture and Spatial Patterns. In: Crimi, A., Bakas, S., Kuijf, H., Keyvan, F., Reyes, M., van Walsum, T. (eds) Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries. BrainLes 2018. Lecture Notes in Computer Science(), vol 11383. Springer, Cham. https://doi.org/10.1007/978-3-030-11723-8_24
Download citation
DOI: https://doi.org/10.1007/978-3-030-11723-8_24
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-11722-1
Online ISBN: 978-3-030-11723-8
eBook Packages: Computer ScienceComputer Science (R0)