Abstract
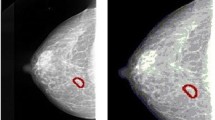
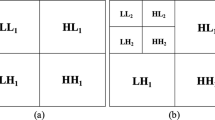
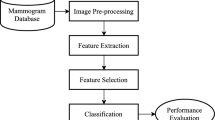
Automatic detection and classification of lesions in mammography remains one of the most important and challenging problems in the development of computer-aided diagnosis systems. Several machine learning approaches have been proposed for supporting the detection and classification of mammographic findings, and are used as computational tools during different diagnosis process by the radiologists. However, the effectiveness of these approaches depends on the accuracy of the feature representation and classification techniques. In this paper, a radiomic strategy based on texture features is explored for identifying abnormalities in mammographies. For doing that, a complete study of five feature extraction approaches, ten selection methods, and five classification models was carried out for identifying findings contained in regions of interest extracted from mammography. The proposed strategy starts with a region extraction process. Some square regions of interest (ROI) were manually extracted from the Mammographic Image Analysis Society (miniMIAS) database. Then, each ROI was decomposed into different resolution levels by using a Wavelet transform approach, and a set of radiomic features based on texture information was computed. Finally, feature selection algorithms and machine learning models were applied to decide whether the ROI undergoing analysis contains or not a mammographic abnormality. The obtained results showed that radiomic texture descriptors extracted from wavelet detail coefficients improved the performance obtained by radiomic features extracted from the original image.
G.M. Díaz—This work was supported by Colciencias (RFC 740-2017) and the Instituto Tecnológico Metropolitano.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Beura, S., Majhi, B., Dash, R.: Mammogram classification using two dimensional discrete wavelet transform and gray-level co-occurrence matrix for detection of breast cancer. Neurocomputing 154, 1–14 (2015). https://doi.org/10.1016/j.neucom.2014.12.032
Erickson, B.J., Korfiatis, P., Akkus, Z., Kline, T.L.: Machine learning for medical imaging. Radiographics 37(2), 505–515 (2017)
Gillies, R.J., Kinahan, P.E., Hricak, H.: Radiomics: images are more than pictures, they are data. Radiology 278(2), 563–577 (2015)
Görgel, P., Sertbas, A., Ucan, O.N.: Mammographical mass detection and classification using local seed region growing-spherical wavelet transform (LSRG-SWT) hybrid scheme. Comput. Biol. Med. 43(6), 765–774 (2013). https://doi.org/10.1016/j.compbiomed.2013.03.008
Haralick, R., Shanmugan, K., Dinstein, I.: Textural features for image classification (1973). https://doi.org/10.1109/TSMC.1973.4309314
Jona, J.B.: A hybrid swarm optimization approach for feature set reduction in digital mammograms. WSEAS Trans. Inf. Sci. Appl. 9(11), 340–349 (2012)
Lambin, P., et al.: Radiomics: extracting more information from medical images using advanced feature analysis. Eur. J. Cancer 48(4), 441–446 (2012)
Lee, A.Y., et al.: Inter-reader variability in the use of bi-rads descriptors for suspicious findings on diagnostic mammography: a multi-institution study of 10 academic radiologists. Acad. Radiol. 24(1), 60–66 (2017)
Li, J., et al.: Feature selection: a data perspective (2016). https://doi.org/10.1145/3136625
Narváez, F., Díaz, G., Poveda, C., Romero, E.: An automatic BI-RADS description of mammographic masses by fusing multiresolution features. Expert. Syst. Appl. 74, 82–95 (2017)
Ojala, T., Pietikainen, M., Harwood, D.: Performance evaluation of texture measures with classification based on Kullback discrimination of distributions. In: Proceedings of 12th International Conference on Pattern Recognition, vol. 1, pp. 582–585 (1994). https://doi.org/10.1109/ICPR.1994.576366
Parmar, C., Grossmann, P., Bussink, J., Lambin, P., Aerts, H.J.: Machine learning methods for quantitative radiomic biomarkers. Sci. Rep. 5, 1–11 (2015). https://doi.org/10.1038/srep13087
Pedregosa, F., et al.: Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011)
Stewart, B.W., Wild, C.P.: World cancer report 2014 (2014)
Subashini, T.S., Ramalingam, V., Palanivel, S.: Automated assessment of breast tissue density in digital mammograms. Comput. Vis. Image Underst. 114(1), 33–43 (2010). https://doi.org/10.1016/j.cviu.2009.09.009
Suckling, J., et al.: The mammographic image analysis society digital mammogram database. In: Experta Medica, International Congress Series, vol. 1069, pp. 375–378, January 1994
Wang, D., Shi, L., Heng, P.A.: Automatic detection of breast cancers in mammograms using structured support vector machines. Neurocomputing 72(13–15), 3296–3302 (2009). https://doi.org/10.1016/j.neucom.2009.02.015
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Rincón, J.S., Castro-Ospina, A.E., Narváez, F.R., Díaz, G.M. (2019). Machine Learning Methods for Classifying Mammographic Regions Using the Wavelet Transform and Radiomic Texture Features. In: Botto-Tobar, M., Pizarro, G., Zúñiga-Prieto, M., D’Armas, M., Zúñiga Sánchez, M. (eds) Technology Trends. CITT 2018. Communications in Computer and Information Science, vol 895. Springer, Cham. https://doi.org/10.1007/978-3-030-05532-5_47
Download citation
DOI: https://doi.org/10.1007/978-3-030-05532-5_47
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-05531-8
Online ISBN: 978-3-030-05532-5
eBook Packages: Computer ScienceComputer Science (R0)