Abstract
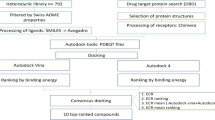
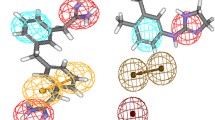
Needless to say that cure from malaria disease is a longstanding goal and helping the present worldwide effort to reduce the malarial disease burden by many folds. Chloroquine (CQ) and Artemisinin (ART) antimalarial drugs are among the most successful in treating malaria; however, CQ-resistant Plasmodium species are now at every corner of the malarial epidemic regions, while ART resistance is upcoming. Pharmacophore features derived from both drugs can be implemented to overcome the growing resistance as well as designing multi-targeted novel non-Chloroquine and non-Artemisinin scaffolds containing inhibitors. In the present study, two different pharmacophore modeling approaches are used to understand the pharmacophore feature distribution between the CQ-Sensitive and CQ-Resistance analogs. To exclude the possible resistance-causing features for CQ, a subtractive pharmacophore modeling protocol has been illustrated and further optimized to a CQ model which is used to screen novel antimalarial inhibitors. Along with CQ, Artemisinin analogs are also used for pharmacophore modeling and additive pharmacophore modeling has been implemented for novel antimalarials designing. We have discussed and used these models for searching novel inhibitors. The subtractive model is used to identify non-CQ scaffolds containing inhibitors while the additive model is used to design the non-ART & non-CQ scaffold containing inhibitors by chemical mining approach to overcome malarial resistance.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Abbreviations
- ART:
-
Artemisinin
- CQ:
-
Chloroquine
- CQR:
-
Chloroquine resistant
- CQS:
-
Chloroquine sensitive
- DHA:
-
dihydroartemisinin
- PC:
-
Physicochemical properties
- PH:
-
Pharmacophore
- QSAR:
-
Quantitative structure-activity relationship
- SBDD:
-
Structure-based drug design
References
Cui L, Mharakurwa S, Ndiaye D, Rathod PK, Rosenthal PJ (2015) Antimalarial drug resistance: literature review and activities and findings of the ICEMR network. Am J Trop Med Hyg 93(3 Suppl):57–68. https://doi.org/10.4269/ajtmh.15-0007
Klein EY (2013) Antimalarial drug resistance: a review of the biology and strategies to delay emergence and spread. Int J Antimicrob Agents 41(4):311–317. https://doi.org/10.1016/j.ijantimicag.2012.12.007
Schlitzer M (2007) Malaria chemotherapeutics part I: history of antimalarial drug development, currently used therapeutics, and drugs in clinical development. ChemMedChem 2(7):944–986. https://doi.org/10.1002/cmdc.200600240
Miotto O, Almagro-Garcia J, Manske M, Macinnis B, Campino S, Rockett KA, Amaratunga C, Lim P, Suon S, Sreng S, Anderson JM, Duong S, Nguon C, Chuor CM, Saunders D, Se Y, Lon C, Fukuda MM, Amenga-Etego L, Hodgson AV, Asoala V, Imwong M, Takala-Harrison S, Nosten F, Su XZ, Ringwald P, Ariey F, Dolecek C, Hien TT, Boni MF, Thai CQ, Amambua-Ngwa A, Conway DJ, Djimde AA, Doumbo OK, Zongo I, Ouedraogo JB, Alcock D, Drury E, Auburn S, Koch O, Sanders M, Hubbart C, Maslen G, Ruano-Rubio V, Jyothi D, Miles A, O'Brien J, Gamble C, Oyola SO, Rayner JC, Newbold CI, Berriman M, Spencer CC, McVean G, Day NP, White NJ, Bethell D, Dondorp AM, Plowe CV, Fairhurst RM, Kwiatkowski DP (2013) Multiple populations of artemisinin-resistant Plasmodium falciparum in Cambodia. Nat Genet 45(6):648–655. https://doi.org/10.1038/ng.2624
Payne D (1987) Spread of chloroquine resistance in Plasmodium falciparum. Parasitol Today 3(8):241–246. https://doi.org/10.1016/0169-4758(87)90147-5
Hsu E (2006) Reflections on the 'discovery' of the antimalarial qinghao. Br J Clin Pharmacol 61(6):666–670. https://doi.org/10.1111/j.1365-2125.2006.02673.x
O'Neill PM, Barton VE, Ward SA (2010) The molecular mechanism of action of artemisinin – the debate continues. Molecules 15(3):1705–1721. https://doi.org/10.3390/molecules15031705
Phyo AP, Nkhoma S, Stepniewska K, Ashley EA, Nair S, McGready R, Ler Moo C, Al-Saai S, Dondorp AM, Lwin KM, Singhasivanon P, Day NP, White NJ, Anderson TJ, Nosten F (2012) Emergence of artemisinin-resistant malaria on the western border of Thailand: a longitudinal study. Lancet 379(9830):1960–1966. https://doi.org/10.1016/S0140-6736(12)60484-X
Hyde JE (2005) Drug-resistant malaria. Trends Parasitol 21(11):494–498. https://doi.org/10.1016/j.pt.2005.08.020
Ursos LM, Roepe PD (2002) Chloroquine resistance in the malarial parasite, Plasmodium falciparum. Med Res Rev 22(5):465–491. https://doi.org/10.1002/med.10016
Ashley EA, Phyo AP (2018) Drugs in development for malaria. Drugs 78(9):861–879. https://doi.org/10.1007/s40265-018-0911-9
Flannery EL, Chatterjee AK, Winzeler EA (2013) Antimalarial drug discovery – approaches and progress towards new medicines. Nat Rev Microbiol 11(12):849–862. https://doi.org/10.1038/nrmicro3138
Murithi JM, Owen ES, Istvan ES, Lee MCS, Ottilie S, Chibale K, Goldberg DE, Winzeler EA, Llinas M, Fidock DA, Vanaerschot M (2020) Combining stage specificity and metabolomic profiling to advance antimalarial drug discovery. Cell Chem Biol 27(2):158–171 e153. https://doi.org/10.1016/j.chembiol.2019.11.009
Okombo J, Chibale K (2017) Insights into integrated lead generation and target identification in malaria and tuberculosis drug discovery. Acc Chem Res 50(7):1606–1616. https://doi.org/10.1021/acs.accounts.6b00631
Gamo FJ, Sanz LM, Vidal J, de Cozar C, Alvarez E, Lavandera JL, Vanderwall DE, Green DV, Kumar V, Hasan S, Brown JR, Peishoff CE, Cardon LR, Garcia-Bustos JF (2010) Thousands of chemical starting points for antimalarial lead identification. Nature 465(7296):305–310. https://doi.org/10.1038/nature09107
Margineanu DG (2014) Systems biology, complexity, and the impact on antiepileptic drug discovery. Epilepsy Behav 38:131–142. https://doi.org/10.1016/j.yebeh.2013.08.029
Wright MH, Clough B, Rackham MD, Rangachari K, Brannigan JA, Grainger M, Moss DK, Bottrill AR, Heal WP, Broncel M, Serwa RA, Brady D, Mann DJ, Leatherbarrow RJ, Tewari R, Wilkinson AJ, Holder AA, Tate EW (2014) Validation of N-myristoyltransferase as an antimalarial drug target using an integrated chemical biology approach. Nat Chem 6(2):112–121. https://doi.org/10.1038/nchem.1830
Stein RM, Kang HJ, McCorvy JD, Glatfelter GC, Jones AJ, Che T, Slocum S, Huang XP, Savych O, Moroz YS, Stauch B, Johansson LC, Cherezov V, Kenakin T, Irwin JJ, Shoichet BK, Roth BL, Dubocovich ML (2020) Virtual discovery of melatonin receptor ligands to modulate circadian rhythms. Nature 579(7800):609–614. https://doi.org/10.1038/s41586-020-2027-0
Romeo S, Dell'Agli M, Parapini S, Rizzi L, Galli G, Mondani M, Sparatore A, Taramelli D, Bosisio E (2004) Plasmepsin II inhibition and antiplasmodial activity of Primaquine-Statine ‘double-drugs’. Bioorg Med Chem Lett 14(11):2931–2934. https://doi.org/10.1016/j.bmcl.2004.03.030
Al-Bari MA (2015) Chloroquine analogues in drug discovery: new directions of uses, mechanisms of actions and toxic manifestations from malaria to multifarious diseases. J Antimicrob Chemother 70(6):1608–1621. https://doi.org/10.1093/jac/dkv018
Dai T, Jiang W, Guo Z, Xie Y, Dai R (2019) Comparison of in vitro/in vivo blood distribution and pharmacokinetics of artemisinin, artemether and dihydroartemisinin in rats. J Pharm Biomed Anal 162:140–148. https://doi.org/10.1016/j.jpba.2018.09.024
Hu YQ, Gao C, Zhang S, Xu L, Xu Z, Feng LS, Wu X, Zhao F (2017) Quinoline hybrids and their antiplasmodial and antimalarial activities. Eur J Med Chem 139:22–47. https://doi.org/10.1016/j.ejmech.2017.07.061
Lawrenson AS, Cooper DL, O'Neill PM, Berry NG (2018) Study of the antimalarial activity of 4-aminoquinoline compounds against chloroquine-sensitive and chloroquine-resistant parasite strains. J Mol Model 24(9):237. https://doi.org/10.1007/s00894-018-3755-z
Solomon VR, Haq W, Srivastava K, Puri SK, Katti SB (2007) Synthesis and antimalarial activity of side chain modified 4-aminoquinoline derivatives. J Med Chem 50(2):394–398. https://doi.org/10.1021/jm061002i
Reiling SJ, Rohrbach P (2019) Uptake of a fluorescently tagged chloroquine analogue is reduced in CQ-resistant compared to CQ-sensitive Plasmodium falciparum parasites. Malar J 18(1):342. https://doi.org/10.1186/s12936-019-2980-y
De D, Krogstad FM, Byers LD, Krogstad DJ (1998) Structure-activity relationships for antiplasmodial activity among 7-substituted 4-aminoquinolines. J Med Chem 41(25):4918–4926. https://doi.org/10.1021/jm980146x
Kaschula CH, Egan TJ, Hunter R, Basilico N, Parapini S, Taramelli D, Pasini E, Monti D (2002) Structure-activity relationships in 4-aminoquinoline antiplasmodials. The role of the group at the 7-position. J Med Chem 45(16):3531–3539. https://doi.org/10.1021/jm020858u
Sinha M, Dola VR, Agarwal P, Srivastava K, Haq W, Puri SK, Katti SB (2014) Antiplasmodial activity of new 4-aminoquinoline derivatives against chloroquine resistant strain. Bioorg Med Chem 22(14):3573–3586. https://doi.org/10.1016/j.bmc.2014.05.024
Sullivan Jr DJ, Gluzman IY, Russell DG, Goldberg DE (1996) On the molecular mechanism of chloroquine’s antimalarial action. Proc Natl Acad Sci U S A 93(21):11865–11870. https://doi.org/10.1073/pnas.93.21.11865
Ismail HM, Barton V, Phanchana M, Charoensutthivarakul S, Wong MH, Hemingway J, Biagini GA, O'Neill PM, Ward SA (2016) Artemisinin activity-based probes identify multiple molecular targets within the asexual stage of the malaria parasites Plasmodium falciparum 3D7. Proc Natl Acad Sci U S A 113(8):2080–2085. https://doi.org/10.1073/pnas.1600459113
Bridgford JL, Xie SC, Cobbold SA, Pasaje CFA, Herrmann S, Yang T, Gillett DL, Dick LR, Ralph SA, Dogovski C, Spillman NJ, Tilley L (2018) Artemisinin kills malaria parasites by damaging proteins and inhibiting the proteasome. Nat Commun 9(1):3801. https://doi.org/10.1038/s41467-018-06221-1
Talevi A (2015) Multi-target pharmacology: possibilities and limitations of the “skeleton key approach” from a medicinal chemist perspective. Front Pharmacol 6:205. https://doi.org/10.3389/fphar.2015.00205
Gray DA, Wenzel M (2020) Multitarget approaches against multiresistant superbugs. ACS Infect Dis 6(6):1346–1365. https://doi.org/10.1021/acsinfecdis.0c00001
Kumar P, Ghosh I (2021) Molecular multi-target approach on COVID-19 for designing novel chemicals. In: Roy K (ed) Methods in pharmacology and toxicology. Springer, New York, pp 1–24. https://doi.org/10.1007/7653_2020_52
Kumar P, Kaalia R, Srinivasan A, Ghosh I (2018) Multiple target-based pharmacophore design from active site structures. SAR QSAR Environ Res 29(1):1–19. https://doi.org/10.1080/1062936X.2017.1401555
Bhattacharjee AK, Kyle DE, Vennerstrom JL (2001) Structural analysis of chloroquine resistance reversal by imipramine analogs. Antimicrob Agents Chemother 45(9):2655–2657. https://doi.org/10.1128/aac.45.9.2655-2657.2001
Bagchi MC, Mills D, Basak SC (2007) Quantitative structure-activity relationship (QSAR) studies of quinolone antibacterials against M. fortuitum and M. smegmatis using theoretical molecular descriptors. J Mol Model 13(1):111–120. https://doi.org/10.1007/s00894-006-0133-z
Hawkins DM, Basak SC, Shi X (2001) QSAR with few compounds and many features. J Chem Inf Comput Sci 41(3):663–670. https://doi.org/10.1021/ci0001177
Masand VH, Toropov AA, Toropova AP, Mahajan DT (2014) QSAR models for anti-malarial activity of 4-aminoquinolines. Curr Comput Aided Drug Des 10(1):75–82. https://doi.org/10.2174/1573409910666140303114621
Mahmud AW, Shallangwa GA, Uzairu A (2020) QSAR and molecular docking studies of 1,3-dioxoisoindoline-4-aminoquinolines as potent antiplasmodium hybrid compounds. Heliyon 6(3):e03449. https://doi.org/10.1016/j.heliyon.2020.e03449
Bhattacharjee AK, Kyle DE, Vennerstrom JL, Milhous WK (2002) A 3D QSAR pharmacophore model and quantum chemical structure – activity analysis of chloroquine(CQ)-resistance reversal. J Chem Inf Comput Sci 42(5):1212–1220. https://doi.org/10.1021/ci0200265
Hocart SJ, Liu H, Deng H, De D, Krogstad FM, Krogstad DJ (2011) 4-Aminoquinolines active against chloroquine-resistant Plasmodium falciparum: basis of antiparasite activity and quantitative structure-activity relationship analyses. Antimicrob Agents Chemother 55(5):2233–2244. https://doi.org/10.1128/AAC.00675-10
Sahu NK, Sharma MC, Mourya V, Kohli DV (2014) QSAR studies of some side chain modified 7-chloro-4-aminoquinolines as antimalarial agents. Arab J Chem 7(5):701–707. https://doi.org/10.1016/j.arabjc.2010.12.005
Shibi IG, Aswathy L, Jisha RS, Masand VH, Divyachandran A, Gajbhiye JM (2015) Molecular docking and QSAR analyses for understanding the antimalarial activity of some 7-substituted-4-aminoquinoline derivatives. Eur J Pharm Sci 77:9–23. https://doi.org/10.1016/j.ejps.2015.05.025
Singh B, Vishwakarma RA, Bharate SB (2013) QSAR and pharmacophore modeling of natural and synthetic antimalarial prodiginines. Curr Comput Aided Drug Des 9(3):350–359. https://doi.org/10.2174/15734099113099990020
Gupta AK, Chakroborty S, Srivastava K, Puri SK, Saxena AK (2010) Pharmacophore modeling of substituted 1,2,4-trioxanes for quantitative prediction of their antimalarial activity. J Chem Inf Model 50(8):1510–1520. https://doi.org/10.1021/ci100180e
Kumar P, Shandilya A, Jayaram B, Ghosh I (2017) Integrative method for finding antimalarials using in silico approach. In: Kholmurodov KT (ed) Computer design for new drugs and materials: molecular dynamics of nanoscale phenomena. Nova Science Publishers
Gaulton A, Hersey A, Nowotka M, Bento AP, Chambers J, Mendez D, Mutowo P, Atkinson F, Bellis LJ, Cibrian-Uhalte E, Davies M, Dedman N, Karlsson A, Magarinos MP, Overington JP, Papadatos G, Smit I, Leach AR (2017) The ChEMBL database in 2017. Nucleic Acids Res 45(D1):D945–D954. https://doi.org/10.1093/nar/gkw1074
Ekoue-Kovi K, Yearick K, Iwaniuk DP, Natarajan JK, Alumasa J, de Dios AC, Roepe PD, Wolf C (2009) Synthesis and antimalarial activity of new 4-amino-7-chloroquinolyl amides, sulfonamides, ureas and thioureas. Bioorg Med Chem 17(1):270–283. https://doi.org/10.1016/j.bmc.2008.11.009
Iwaniuk DP, Whetmore ED, Rosa N, Ekoue-Kovi K, Alumasa J, de Dios AC, Roepe PD, Wolf C (2009) Synthesis and antimalarial activity of new chloroquine analogues carrying a multifunctional linear side chain. Bioorg Med Chem 17(18):6560–6566. https://doi.org/10.1016/j.bmc.2009.08.003
Natarajan JK, Alumasa JN, Yearick K, Ekoue-Kovi KA, Casabianca LB, de Dios AC, Wolf C, Roepe PD (2008) 4-N-, 4-S-, and 4-O-chloroquine analogues: influence of side chain length and quinolyl nitrogen pKa on activity vs chloroquine resistant malaria. J Med Chem 51(12):3466–3479. https://doi.org/10.1021/jm701478a
Yearick K, Ekoue-Kovi K, Iwaniuk DP, Natarajan JK, Alumasa J, de Dios AC, Roepe PD, Wolf C (2008) Overcoming drug resistance to heme-targeted antimalarials by systematic side chain variation of 7-chloro-4-aminoquinolines. J Med Chem 51(7):1995–1998. https://doi.org/10.1021/jm800106u
BIOVIA DS (2016) BIOVIA discovery studio. Dassault Systèmes, San Diego
Dixon SL, Smondyrev AM, Knoll EH, Rao SN, Shaw DE, Friesner RA (2006) PHASE: a new engine for pharmacophore perception, 3D QSAR model development, and 3D database screening: 1. Methodology and preliminary results. J Comput Aided Mol Des 20(10–11):647–671. https://doi.org/10.1007/s10822-006-9087-6
Dixon SL, Smondyrev AM, Rao SN (2006) PHASE: a novel approach to pharmacophore modeling and 3D database searching. Chem Biol Drug Des 67(5):370–372. https://doi.org/10.1111/j.1747-0285.2006.00384.x
2018-3 SdR (2018) Phase. Schrödinger, LLC, NewYork
Roy K, Mitra I, Kar S, Ojha PK, Das RN, Kabir H (2012) Comparative studies on some metrics for external validation of QSPR models. J Chem Inf Model 52(2):396–408. https://doi.org/10.1021/ci200520g
Basak SC (2019) Importance of proper statistical practices in the use of chemodescriptors and biodescriptors in the twenty-first century. Future Med Chem 11(21):2755–2758. https://doi.org/10.4155/fmc-2019-0250
Hawkins DM, Basak SC, Mills D (2003) Assessing model fit by cross-validation. J Chem Inf Comput Sci 43(2):579–586. https://doi.org/10.1021/ci025626i
Young SS, Ge N (2004) Design of diversity and focused combinatorial libraries in drug discovery. Curr Opin Drug Discov Devel 7(3):318–324
Geysen HM, Schoenen F, Wagner D, Wagner R (2003) Combinatorial compound libraries for drug discovery: an ongoing challenge. Nat Rev Drug Discov 2(3):222–230. https://doi.org/10.1038/nrd1035
Gilson MK, Liu T, Baitaluk M, Nicola G, Hwang L, Chong J (2016) BindingDB in 2015: a public database for medicinal chemistry, computational chemistry and systems pharmacology. Nucleic Acids Res 44(D1):D1045–D1053. https://doi.org/10.1093/nar/gkv1072
Sterling T, Irwin JJ (2015) ZINC 15 – ligand discovery for everyone. J Chem Inf Model 55(11):2324–2337. https://doi.org/10.1021/acs.jcim.5b00559
Cereto-Massague A, Guasch L, Valls C, Mulero M, Pujadas G, Garcia-Vallve S (2012) DecoyFinder: an easy-to-use python GUI application for building target-specific decoy sets. Bioinformatics 28(12):1661–1662. https://doi.org/10.1093/bioinformatics/bts249
Sakkiah S, Lee KW (2012) Pharmacophore-based virtual screening and density functional theory approach to identifying novel butyrylcholinesterase inhibitors. Acta Pharmacol Sin 33(7):964–978. https://doi.org/10.1038/aps.2012.21
Empereur-Mot C, Guillemain H, Latouche A, Zagury JF, Viallon V, Montes M (2015) Predictiveness curves in virtual screening. J Cheminform 7:52. https://doi.org/10.1186/s13321-015-0100-8
Koes DR, Camacho CJ (2011) Pharmer: efficient and exact pharmacophore search. J Chem Inf Model 51(6):1307–1314. https://doi.org/10.1021/ci200097m
Landrum G (2012) RDKit: Open-source cheminformatics. https://www.rdkit.org
John S, Thangapandian S, Arooj M, Hong JC, Kim KD, Lee KW (2011) Development, evaluation and application of 3D QSAR pharmacophore model in the discovery of potential human renin inhibitors. BMC Bioinformatics 12(Suppl 14):S4. https://doi.org/10.1186/1471-2105-12-S14-S4
Acknowledgments
PK would like to acknowledge the Indian Council of medical Research (ICMR) for ICMR-SRF fellowship, and IG wishes to thank the Department of Biotechnology CCPM projects and the DST purse, Government of India, for support. We are also thankful to Dassault Systems and Schrodinger for providing the Discovery Studio and Phase software, without which a part of this work would have not been possible.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Ethics declarations
Funding
There is no funding available presently for writing the article, but SRF scholarship from ICMR, DST PURSE & DBT project funding supported the research work earlier.
Ethical Approval
This manuscript is a review of previously published accounts in the Thesis, as such no animal or human studies were performed.
Informed Consent
No patients were studied in this chapter.
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Kumar, P., Ghosh, I. (2021). Probing with Pharmacophore Modeling the Chloroquine Resistance and Designing Novel Antimalarials. In: Saxena, A.K. (eds) Biophysical and Computational Tools in Drug Discovery. Topics in Medicinal Chemistry, vol 37. Springer, Cham. https://doi.org/10.1007/7355_2021_131
Download citation
DOI: https://doi.org/10.1007/7355_2021_131
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-85280-1
Online ISBN: 978-3-030-85281-8
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)