Abstract
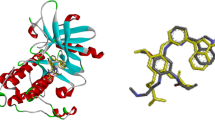
Leukemia inhibitory factor (LIF) stimulates the terminal differentiation of the cells of myelogenous leukemia by interacting with gp130 (dimeric) and LIF receptor (LIFR). Janus protein-tyrosine kinases (JAK) and STAT3 get triggered by phosphorylation. This leads to target gene expression and thus inhibition of the growth of leukemic cells occurs. Wet-laboratory experimental studies were involved so far, but the residue-based molecular-level indulgement and structural changes in the protein complexes were not explored hitherto. This study therefore involves X-ray crystal structures of the three proteins. Through molecular docking techniques, the best cluster-sized complex was selected and optimized. Electrostatic surface potential for gp130 protein, net solvent accessibility and an ascent in the ΔG value from −2101.57 kcal/mol to −2124.28 kcal/mol showed spontaneous and firmer interaction after optimization. Conformational fluctuations in gp130 had a shift for increased β-sheet conformation to stabilize the complex. All evaluations were statistically significant. Protein interaction residues and binding patterns revealed that ionic interactions were the predominant ones with Asp and Arg playing chief role. After optimization, even ionic interactions increased to 11. Other interactions including hydrogen bonding ones were also seen. Thus, it might instigate the clinical and pharmaceutical research for drug discovery and to study the mutational impacts upon interaction.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Blanchard F et al (2000) Stimulation of leukemia inhibitory factor receptor degradation by extracellular signal-regulated kinase. J Biol Chem 275:28793–28801
Mathieu ME et al (2012) LIF-dependent signaling: new pieces in the lego. Stem Cell Rev Rep 8(1):1–15
Sun X, Bartos A, Whitsett JA, Dey SK (2013) Uterine deletion of Gp130 or Stat3 shows implantation failure with increased estrogenic responses. Mol Endocrinol 27(9):1492–1501
Metz S et al (2008) Novel inhibitors for murine and human leukemia inhibitory factor based on fused soluble receptors. J Biol Chem 283:5985–5995
Berman MH et al (2000) The protein data bank. Nucleic Acids Res 28:235–242
Boulanger MJ, Bankovich AJ, Kortemme T, Baker D, Garcia KC (2003) Convergent mechanisms for recognition of divergent cytokines by the shared signaling receptor gp130. Mol Cell 12:577–589
Skiniotis G, Lupardus PJ, Martick M, Walz T, Garcia KC (2008) Structural organization of a full-length gp130/LIF-R cytokine receptor transmembrane complex. Mol Cell 31:737–748
Comeau SR et al (2004) ClusPro: an automated docking and discrimination method for the prediction of protein complexes. Bioinformatics 20:45–50
Pierce BG et al (2014) ZDOCK server: interactive docking prediction of protein-protein complexes and symmetric multimers. Bioinformatics 30(12):1771–1773
Tovchigrechko A, Vakser IA (2006) GRAMM-X public web server for protein-protein docking. Nucleic Acids Res 34:W310–W314
Xu D, Zhang Y (2011) Improving the physical realism and structural accuracy of protein models by a two- step atomic level energy minimization. Biophys J 101:2525–2534
Abrahama MJ et al (2015) GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2:19–25
Tina KG, Bhadra R, Srinivasan N (2007) PIC: protein interactions calculator. Nucleic Acids Res 35:W473–W476
DeLano WL (2002) The PyMOL molecular graphics system. DeLano Scientific, San Carlos
Yuedong Y, Yaoqi Z (2008) Specific interactions for ab initio folding of protein terminal regions with secondary structures. Proteins 72:793–803
Gerstein M (1992) A resolution sensitive procedure for comparing protein surfaces and its application to the comparison of antigen-combining sites. Acta Crystallogr A 48:271–276
Paul DT, Ken DA (1993) Local and nonlocal interactions in globular proteins and mechanisms of alcohol denaturation. Protein Sci 2:2050–2065
Toniolo C, Benedetti E (1991) The polypeptide 310-helix. Trends Biochem Sci 16:350–353
Banerjee A, Ray S (2015) Molecular computing and structural biology for interactions in ERα and bZIP proteins from homo sapiens: an insight into the signal transduction in breast cancer metastasis. Adv Intell Syst Comput 404:43–55. https://doi.org/10.1007/978-81-322-2695-6_5
Banerjee A, Ray S (2016) Molecular modeling, mutational analysis and conformational switching in IL27: an in silico structural insight towards AIDS research. Gene 576(1):72–78
Acknowledgement
High gratefulness is rendered to the Department of Biochemistry and Biophysics, University of Kalyani, for the support. Authors are also grateful to the Amity Institute of Biotechnology, Amity University, Kolkata, for the cooperation and support as well.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Banerjee, A., Dasgupta, R., Ray, S. (2020). Molecular and Protein Interaction Studies for Inhibiting Growth of Human Leukemic Cells: An In Silico Structural Approach to Instigate Drug Discovery. In: Sadhukhan, P., Premi, S. (eds) Biotechnological Applications in Human Health. Springer, Singapore. https://doi.org/10.1007/978-981-15-3453-9_9
Download citation
DOI: https://doi.org/10.1007/978-981-15-3453-9_9
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-3452-2
Online ISBN: 978-981-15-3453-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)