Abstract
Atrial fibrillation, characterized by irregular atrial depolarization leading to an uncoordinated contraction of the atrial muscle, is the most common cardiac arrhythmia. The T-box transcription factor TBX5 regulates atrial rhythm and has been genetically associated with human atrial fibrillation by both familial and genome-wide association studies. Over the last 30 years, the pulmonary veins and peri-pulmonary region of the left atrium have been implicated as a physical locus that frequently initiates atrial fibrillation. Regional gene expression differences within the atria have been reported that may reflect distinct ontogeny of these regions and impact arrhythmia mechanisms. In this study, we investigated transcriptional differences between regions of the adult left atrium in the mouse. We performed and compared transcriptional profiling of the left atrial appendage and peri-pulmonary vein atrium of the adult mouse. We identified novel markers and transcriptional differences between the two regions, including of multiple known AF genes. Furthermore, we compared Tbx5-dependent genes between each region and identified Hcn4 as a novel dysregulated gene in the peri-pulmonary vein atrium. Regional and Tbx5-dependent gene expression differences between the left atrial appendage and peri-pulmonary vein atrium are considered in the context of the genetic basis of atrial fibrillation.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
- Atrial fibrillation
- Arrhythmia
- TBX5
- Cardiac conduction
- Pulmonary vein
- Atrium
- Gene regulatory network
- PITX2
- HCN4
- SHOX2
1 Introduction
1.1 Atrial Fibrillation and the Pulmonary Vein
Atrial fibrillation (AF) is the most common sustained cardiac arrhythmia, affecting more than 7 million Americans and 33 million people worldwide [1]. AF is characterized by an irregular pattern of atrial depolarization, resulting in rapid and disorganized atrial conduction and lack of effective atrial chamber contraction. The rhythm abnormality in AF manifests with circulatory deficits and systemic thromboembolism that greatly increase morbidity and mortality. Patients with AF have an increased risk of developing major complications including heart failure and stroke. In addition, the prevalence of AF is expected to rise significantly as the population ages. AF has become a major clinical and economic burden, owing to the limitations and side effects associated with current AF therapies. Although AF most often manifests in the context of pre-existing cardiac pathologies, idiopathic or lone AF forms indicated a heritable component [2]. Over the last decade, genome-wide association studies (GWAS) have identified over one hundred AF-associated loci [3, 4]. These advances in understanding the genetic basis of AF portend an understanding of the complex molecular pathways provoking AF.
The current paradigm of AF requires two arrhythmogenic components: (1) ectopic (triggered) atrial myocardial activity such as early or delayed afterdepolarizations and (2) a fibrillogenic substrate that propagates these triggers throughout the atrial chambers [5]. The most common source of triggered activity is the pulmonary veins, which connect the lungs to the left atria [6, 7]. It has been shown that the pulmonary vein (PV) and peri-pulmonary vein (PPV) myocardium can demonstrate automaticity which can trigger atrial depolarization [8]. A pacemaker current has been observed in the PV-left atrial junction in atrial fibrillation, and gene expression contributing to myocardial automaticity, such as the HCN4 channel protein, is observed in the pulmonary veins [9]. A common treatment for AF is to isolate the PV myocytes from the atrial myocardium by catheter ablation, severing the electrical connectivity between these two populations and preventing inappropriate PV depolarizations from initiating global atrial depolarization and atrial arrhythmias [6, 7].
An important advance in the understanding of the pathophysiology of AF has been the demonstration that the pulmonary vein plays an important role in the initiation of AF [6, 10]. Other initiating foci include the superior vena cava and the left atrium; however, the pulmonary vein remains the most common source of focal activity. This observation has stimulated consideration of the distinctions between the myocardium of the pulmonary/peri-pulmonary vein region and other regions of the atrium. However, the gene expression profile promoting focal activity around the PVs in humans has been difficult to elucidate, owing to the limitations of tissue samples from patients. The molecular identity of the PV myocardium, including specific ion channels and transcription factors expressed in this tissue, has only recently been described. Analysis of a whole-lung transcriptome data set uncovered a set of 24 transcripts, including sarcomeric structural proteins, genes regulating sarcomere assembly, ion transport proteins and hormone signaling in the pulmonary cardiomyocyte gene network [11]. Some of these genes have been linked to AF in humans, suggesting that perturbation of this transcriptional network might lead to altered calcium handling, altered myocardium contractile force and electrophysiological properties which may contribute to the initiation of atrial arrhythmias.
1.2 TBX5 and Pulmonary Vein Development
Holt-Oram syndrome is an autosomal dominant disorder caused by mutations in the T-box family transcription factor TBX5 [12,13,14], occurring in approximately 1 in 100,000 live births [15, 16]. The clinical diagnosis of Holt-Oram syndrome includes completely penetrant upper-limb malformations and variable congenital heart defects, most commonly septal and cardiac conduction defects [17]. Specifically, the conduction defects can manifest as long PR interval, AV and bundle branch block, bradycardia, sick sinus syndrome and atrial fibrillation with or without overt structural defects [18, 19]. The observation that conduction abnormalities can occur in the absence of structural heart defects was an early indication that TBX5 may play a direct role in controlling gene expression essential for normal cardiac conduction. Recently, GWAS have implicated TBX5 in altered cardiac conduction speed (PR and QRS intervals) and AF in structurally normal hearts. Although strict clinical diagnosis of Holt-Oram Syndrome does not involve defects of the pulmonary veins [14, 17], pulmonary vein abnormalities are often associated with Holt-Oram syndrome and TBX5 mutations [20,21,22]. Furthermore, homozygous mutation of Tbx5 in the mouse germline [23] results in complete morphological failure of the cardiopulmonary progenitors, which would normally generate portions of the atria, the atrial septum, pulmonary vein myocardium and smooth muscle cells and airways smooth muscle cells [24, 25]. These observations suggested that TBX5 may drive gene expression important for the development of the pulmonary vasculature in the embryo and for the suppression of atrial arrhythmias in the pulmonary veins or peri-pulmonary vein atrial myocardium in the adult.
Here, we describe regional differences between the transcriptome of the left atrial appendage and the peri-pulmonary vein of the left atrium from adult mice. RNA sequencing revealed previously known and novel regional differences, including several genes implicated in atrial fibrillation. We further compared Tbx5-depedent transcripts between the left atrial appendage and the peri-pulmonary vein of the left atrium from adult mice. Important distinctions were observed between the two regions including Hcn4, which demonstrated differential Tbx5-dependence. Our data suggests that underlying regional differences in the transcriptome of the atrial myocardium may enlighten susceptibility to atrial fibrillation.
2 Results
2.1 Comparison of Peri-Pulmonary Vein and Atrial Appendage Transcriptomes
We investigated transcriptional differences between regions of the left atrium in 7-week-old mice. The left atrium was microdissected into two parts: the left atrial appendage (LAA) and the peri-pulmonary vein (PPV) (Fig. 51.1a). Transcriptional profiling was performed by RNA sequencing (RNA-seq; n = 4 in each group) as previously described [26]. Principal component analysis (PCA) demonstrated that atrial region was responsible for the first principal component, accounting for 32% of the variation between samples (Fig. 51.1b). Differential expression testing between the PPV and LAA samples identified 747 significantly different genes (|log2FC| > 1; FDR < 0.01) with 582 expressed more strongly in the PPV samples and 165 expressed more strongly in the LAA samples (Fig. 51.1c, full table available at GEO accession GSE128870). Gene ontology (GO) terms and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways were examined using metascape (http://metascape.org) for both the LAA and PPV genes [27]. GO terms associated with the PPV indicated a preponderance of genes associated with immune cells. We observed GO terms that were unambiguously related to the immune system and only containing genes associated with the PPV, including phagocytosis (GO:0006909), inflammatory response (GO:0006954), humoral immune response (GO:0006959), lymphocyte differentiation (GO:0030098) and cytokine-cytokine receptor interaction (mmu04060). Although these 141 genes are of potential interest, they were separated from the remaining 441 PPV genes to focus on those genes previously verified as myocardially expressed or those with unknown roles or expression domains (Fig. 51.1c).
Transcriptional comparison of the left atrial appendage and peri-pulmonary vein atrium. (a) The left atrial appendage (LAA, blue) and the peri-pulmonary vein region of the left atrium (PPV, red) were microdissected from 7-week-old adult mice. (b) Principal component analysis comparing the LAA (blue) and PPV (red) transcriptomes. (c) Volcano plot showing log2(fold-change PPV/LAA). One hundred and sixty five were associated with LAA and 582 were associated with the PPV (|log2FC|>1 and FDR < 0.01). Of the PPV genes, 141 genes were associated with GO terms related to the immune system (red) and removed from downstream analysis. (d) Overlap of genes associated with atrial fibrillation and either the LAA (7 genes) or PPV (12 genes). P-values derived from Fisher’s exact test
Because of the relationship between the left atria, pulmonary vein and AF, we examined region-specific gene expression against a list of 235 AF associated genes [3, 28,29,30]. Of the 606 genes that demonstrated regionalized expression in our RNA-seq, 19 have been associated with AF (Fig. 51.1d). Specifically, Adrb1, Nkx2-5, Tmem182, Nppa, Nppb, Gja5 and Kcnh7 are more strongly expressed in the LAA while Ntrk2, Tubb4a, Cacna1d, Tbx3, Scn3b, Shox2, Kcne4, Hcn4, Adra1a, Adrb3, Tubb3 and Cacna2d2 have higher expression in the PPV.
2.2 Tbx5-Dependent Transcriptome of Peri-Pulmonary Vein Atrium Identifies Genes Involved with Cardiac Conduction
We hypothesized that Tbx5-dependent gene expression may vary between different regions in the left atria. Previous work has demonstrated that Tbx5 regulates a gene regulatory network for atrial identity and its removal from the adult mouse results in atrial fibrillation [26]. Tbx5 is robustly expressed in both the LAA and PPV tissues with an average TPM of 79.7 ± 9.9 SEM and 97.1 ± 18.4 SEM, respectively putting Tbx5 within the top 8% of all genes expressed in either area of the left atrium (TPM > 1). To investigate differential Tbx5-dependent regional gene expression in the left atrium of the adult mouse, we performed Tbx5-dependent transcriptional profiling of the PPV tissue and compared it to previously described Tbx5-dependent transcriptional profiling of the left atrial appendage [26]. We performed RNA-seq on the PPV of an adult-specific conditional knockout of Tbx5 [26]. Control (R26 creERT2/creERT2; Tbx5+/+) and conditional mutant (R26creERT2/creERT2; Tbx5flox/flox) animals were treated with tamoxifen at 6-weeks old and tissue was harvested one week later prior to the onset or atrial arrhythmias (Fig. 51.1a) [26]. PCA analysis demonstrated clustering along PC1 and PC2 of samples (cumulative variation > 50%). PC1 demonstrated clustering associated with genotype (control versus conditional Tbx5 mutant), and PC2 demonstrated clustering associated with LA region (LAA versus PPV), indicating that genotype differences drove the majority of variance followed by regional atrial differences (Fig. 51.2a).
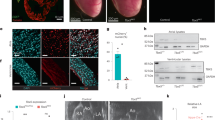
Tbx5 conditional knockout from the peri-pulmonary vein reveals Tbx5-dependence of multiple unique features. (a) Principal component analysis comparing the left atrial appendage (LAA, blue) and peri-pulmonary vein (PPV, red) transcriptomes from control (R26 creERT2/creERT2; Tbx5+/+, “Tbx5Ctrl”, circles) and Tbx5 conditional knockout (R26creERT2/creERT2; Tbx5flox/flox, “Tbx5CKO”, squares) adult mice. (b) Volcano plot comparing the PPV Tbx5 CKO and control samples. One hundred and forty nine genes were significantly up-regulated and 189 were significantly down-regulated (|log2FC|>1 and FDR< 0.01). (c) Summary GO terms representing all GO terms associated with up- and down-regulated genes (red and blue, respectively) with greater than five associated genes and −log10(P-value) greater than 5. (d) Full GO term list represented by “multicellular organismal signaling” with percent of genes identified (bar) and corresponding −log10(P-value) (dot). (e) Correlation of fold changes between the LAA and PPV samples. Colors and shapes represent whether the genes residuals were in the outermost one-thousandth quantiles (purple square, red circle, blue triangle, dark grey circle) or not (light grey circle). Furthermore, colors and shapes represent whether those outliers were significantly different in both datasets (purple square), the PPV only (red circle), the LAA only (blue triangle) or neither (dark grey circle)
We identified 338 significantly dysregulated genes (|log2FC| > 1; FDR < 0.01) comparing the PPV between Tbx5 conditional mutant and controls (Fig. 51.2b, GEO accession GSE128870). Of these, 149 were up-regulated and 189 were down-regulated in the Tbx5 mutants (Fig. 51.2b). Utilizing metascape [27], we identified seven representative GO terms associated with the up-regulated genes and two representative GO terms associated with the down-regulated genes (Fig. 51.2c). The down-regulated GO terms are reminiscent of the LAA previously reported [26]. In particular, examining the full set of GO terms under “multicellular organismal signaling,” we identified multiple GO terms associated with the regulation of cardiac conduction, including ion channels, action potential and transmembrane signaling (Fig. 51.2d).
We identified 1492 significantly dysregulated genes in the LAA conditional mutant and control, with 770 up-regulated and 722 down-regulated, similar to our prior analysis of this dataset [26]. Although the numbers of significantly dysregulated genes were different between the PPV and LAA, the dysregulation between the LAA and PPV experiments were significantly correlated (p-value < 2.2E-16, Fig. 51.2e), suggesting that the removal of Tbx5 had broadly similar effects in the LAA and PPV regions. In order to identify genes demonstrating distinct Tbx5-dependent effects between the atrial sub-regions, we correlated the Tbx5-dependent changes between the LAA and PPV samples. We identified the 28 genes with residuals in the outermost one-thousandth quantiles (Table 51.1). Amongst those genes, we identified 11 genes that were significantly different in both experiments with 10/11 demonstrating changes in the same direction, 4 genes that were significantly increased in only the LAA samples (Cemip, Col9a2, Itgb7, Lars2), 7 genes that were significantly different in only the PPV samples (increased: Gm42417, Lrtm1; decreased: Alox8, Cntn2, Hcn4, Scarna10, Vsnl1) and 6 genes that were not significantly different in either sample. Interestingly, among the genes that were in the outermost one-thousandth quantiles, we identified Hcn4, which encodes the potassium channel essential for the depolarizing “funny current” of the sinoatrial node. Hcn4 was significantly down-regulated only in the PPV and was also significantly more expressed in the PPV samples than the LAA (Fig. 51.1d).
3 Discussion
3.1 The Genetics of Atrial Fibrillation Implicates Developmental Transcription Factors
Most genetic loci implicated in AF encode ion channels that affect arrhythmia trigger or substrate. Initial transcriptional studies of AF have focused on a restricted set of candidates, associating remodeling with changes in ion channels and components of cellular signaling cascades [31, 32]. Early transcriptomic or proteomic investigation of AF has been limited to animal models and/or chronic AF [33,34,35,36]. For example, animal models based on disruption of specific ion channels do not exhibit spontaneous AF in the absence of concomitant cardiac pathophysiology [37]. However, a recent transcriptomic study using the left atrial appendage of patients with AF revealed that AF susceptibility was associated with decreased expression of transcription factors involved in inflammation, oxidation and the cellular stress response [38]. This study further suggested that remodeling of ion channel expression occurs later in the onset of the disease. Accumulating evidence suggests that transcription factors are important contributors to the pathogenesis of AF [39].
GWAS of AF have identified numerous transcription factor loci that play fundamental roles during cardiac development, including TBX5, NKX2-5, GATA4 and PITX2. The implication of transcription factor genes essential for cardiac development in in AF raises the fundamental question of whether AF is a function or a result of a developmental defect, for example, of the pulmonary veins, or a later manifestation of altered transcriptional control of cardiac rhythm genes in the adult. The first GWAS of AF identified a susceptibility locus on chromosome 4q25 adjacent to PITX2 [40]. There is evidence that Pitx2c affects adult gene expression in the atrium. PITX2 is expressed both in the left atrium and the pulmonary myocardial sleeve and that its levels are decreased in patients with AF [41, 42]. Atrial-specific deletion of Pitx2c leads to hallmarks of AF, which is also observed in Pitx2c heterozygous mice or adult-specific deletion of this gene [41, 43, 44]. Furthermore, microarray analysis of Pitx2c heterozygous mice revealed impaired gap and tight junction gene expression, consistent with the hypothesis that structural genes are remodeled later in the onset of AF [44].
We previously demonstrated that adult-specific deletion of Tbx5 causes spontaneous and sustained AF with atrial gene regulatory network dysfunction [26]. TBX5 drives the atrial expression of Pitx2, and TBX5 and PITX2 co-modulate the expression of cardiac rhythm effector genes, including Ryr2 and Atp2a2. These findings indicated that interactions between TBX5 and PITX2 provide tight control of an atrial rhythm gene regulatory network and that perturbation of this network triggered AF susceptibility. This example suggested that cardiac TFs implicated in AF by genetic association may co-regulate a gene regulatory network for atrial rhythm homeostasis in the adult atrium.
3.2 Transcription Factors and AF: A Pulmonary Vein Developmental Relationship?
The requirement of developmental TFs in the adult for normal cardiac rhythm control does not rule out the possibility that developmental defects caused by their deficiency in the embryo also contributes to AF susceptibility. During embryonic development, the cardiopulmonary progenitors give rise to parts of the atria, pulmonary vein and lungs [25]. Distinct functional characteristics between the PPV and LAA regions may reflect distinct developmental ontogeny. The myocardium of the pulmonary veins has a distinct developmental origin from that of the atrial free walls [45]. Subsequently, the myocardium surrounding the pulmonary veins is distinct from that of other regions in the atrium [46]. These developmental distinctions may set the stage for regional gene expression differences within the mature atrium of the adult that normally maintains atrial rhythm but which make the PPV region susceptible to trigger formation when faced with environmental or genetic insult.
Mutations in NKX2-5, PITX2C and TBX5 increase susceptibility to AF, and all of these transcription factors (TF) play critical roles during cardiovascular development [26, 47,48,49,50]. Each of these TFs are expressed in the pulmonary veins, suggesting that perturbations of the transcription program in the PV myocardium may underlie the pathogenesis of AF. The caudal myocardium, including the developing pulmonary vein myocardium, expresses Tbx5, Pitx2c, Nkx2-5 and Gja5 (Cx40) at high levels [51, 52] consistent with a fast-conducting atrial myocardial phenotype. In a Nkx2-5 haploinsufficient mouse model, Gja5 levels are decreased whereas Hcn4 levels are increased in the pulmonary vein myocardium. Nkx2-5 and Pitx2c are both required for the development and maintain identity of the pulmonary myocardium [45, 53]. The changes in gene expression observed in Nkx2-5 haploinsufficient mice could be reflective of a loss of fast conduction and acquisition of pacemaking conduction phenotype. This type of transition provided a conceivable model for how genetic changes could result in automaticity in the PPV region that resulted in arrhythmia initiation.
Absence of the lungs and pulmonary vasculature in Tbx5 knockout mice suggests that it may play an essential developmental role specifically within the pulmonary veins as well [24]. We observed that a decrement in Tbx5 in the PPV region decreased Hcn4 expression, which is not overtly consistent with this transformation model. On the other hand, we observed that many genes with important roles in myocardial automaticity, including Tbx3, Shox2, Cacna1d, Cacna2d2 and Hcn4 were more highly expressed in the PPV than the LAA region, and remained more highly expressed in the context of adult-specific Tbx5 conditional deletion. Furthermore, other Tbx5-dependent genes in the PPV region are classical fast conduction genes, including Scn5a, Gja5 and Kcnk3. Reduction in the expression of these fast conduction genes in the context of retained high expression of automaticity genes such as Hcn4 could diminish fast conduction physiology and allow the emergence of myocardial automaticity, providing a nidus for the initiation of atrial arrhythmias including AF.
4 Methods
4.1 Animal Experiments and Ethics Statement
All murine experiments were performed under University of Chicago Institutional Animal Care and Use Committee (IACUC) protocol number 71737. Mice harboring the ROSA26 tamoxifen-inducible cre recombinase transgene, Gt(ROSA)26Sor tm1(cre/ERT2)Tyj (R26 creERT2), were bred with mice harboring the Tbx5 tm1Jse allele (Tbx5 flox) to generate Tbx5 conditional mutants, R26 creERT2/creERT2; Tbx5flox/flox, and controls, R26 creERT2/creERT2; Tbx5+/+ [23, 54]. Tamoxifen was administered by intraperitoneal injection for three consecutive days at 6 weeks of age and tissue samples were harvested 1 week after treatment as previously reported [26].
4.2 Transcriptional Profiling and Analysis
Transcriptional profiling was performed on the peri-pulmonary vein region of the left atrium (Fig. 51.1a). Total RNA was extracted from four controls and six conditional mutants. Following rRNA removal (Ribo-Zero rRNA Removal Kit, Illumina), libraries were prepared using the TruSeq RNA Sample prep kit v2 (Illumina). Samples were pooled in equimolar ratios and sequenced on the Illumina HiSeq4000 platform. Library preparation and sequencing were performed by the Genomics Core Facility at the University of Chicago.
The peri-pulmonary vein samples were compared with the previously published corresponding left atrial appendage samples [26]. Both the peri-pulmonary vein and left atrial appendage samples were analyzed in parallel as described below.
Between 20 and 22 million reads were generated for each replicate (14–19 million reads for left atrial appendage) and aligned to the GRCm38/mm10 build of the Mus musculus genome (retrieved from UCSC May 23, 2012) using TopHat v2.1.1 [55, 56]. Reads were filtered to remove poorly aligned (MAPQ<10), duplicated and unmapped reads using BamTools v2.4.1 [57]. Post-alignment, post-filtered reads were assigned to transcripts using StringTie v1.3.3 [58, 59].
Downstream analysis was performed using R v3.4.0 [60]. Differential expression testing was performed using edgeR v3.18.1 [61] and limma v3.32.10 [62]. Low level genes were removed within each condition using median log-transformed counts per gene per million mapped reads (cpm) cutoff of 1. A generalized linear model (GLM) framework was used to test for differential expressed genes between Tbx5 fl/fl;R26 cre-ERT2 conditional mutants and Tbx5 +/+;R26 cre-ERT2 controls for each tissue and between controls of the left atrial appendage and peri-pulmonary vein tissues. Data was plotted using the ggplot2 v2.3.0 [63] and scales v0.5.0 [64] packages. GO term analysis was performed using metascape (http://metascape.org) [27]. The raw and processed data as well as the full outputs were deposited in GEO at accession number GSE128870.
References
Nishida K, Datino T, Macle L, Nattel S. Atrial fibrillation ablation: translating basic mechanistic insights to the patient. J Am Coll Cardiol. 2014;64(8):823–31.
Tucker NR, Ellinor PT. Emerging directions in the genetics of atrial fibrillation. Circ Res. 2014;114(9):1469–82.
Fatkin D, Santiago CF, Huttner IG, Lubitz SA, Ellinor PT. Genetics of Atrial Fibrillation: State of the Art in 2017. Heart Lung Circ. 2017;26(9):894–901.
Nielsen JB, et al. Genome-wide Study of Atrial Fibrillation Identifies Seven Risk Loci and Highlights Biological Pathways and Regulatory Elements Involved in Cardiac Development. Am J Hum Genet. 2018;102(1):103–15.
Andrade J, Khairy P, Dobrev D, Nattel S. The clinical profile and pathophysiology of atrial fibrillation: relationships among clinical features, epidemiology, and mechanisms. Circ Res. 2014;114(9):1453–68.
Haissaguerre M, et al. Spontaneous initiation of atrial fibrillation by ectopic beats originating in the pulmonary veins. N Engl J Med. 1998;339(10):659–66.
Jais P, et al. A focal source of atrial fibrillation treated by discrete radiofrequency ablation. Circulation. 1997;95(3):572–6.
Chen YJ, Chen SA, Chang MS, Lin CI. Arrhythmogenic activity of cardiac muscle in pulmonary veins of the dog: implication for the genesis of atrial fibrillation. Cardiovasc Res. 2000;48(2):265–73.
Nguyen BL, Fishbein MC, Chen LS, Chen PS, Masroor S. Histopathological substrate for chronic atrial fibrillation in humans. Heart Rhythm. 2009;6(4):454–60.
Chen SA, et al. Initiation of atrial fibrillation by ectopic beats originating from the pulmonary veins: electrophysiological characteristics, pharmacological responses, and effects of radiofrequency ablation. Circulation. 1999;100(18):1879–86.
Boutilier JK, et al. Gene expression networks in the murine pulmonary myocardium provide insight into the pathobiology of atrial fibrillation. G3 (Bethesda). 2017;7(9):2999–3017.
Basson CT, et al. Mutations in human TBX5 [corrected] cause limb and cardiac malformation in Holt-Oram syndrome. Nat Genet. 1997;15(1):30–5.
Li QY, et al. Holt-Oram syndrome is caused by mutations in TBX5, a member of the Brachyury (T) gene family. Nat Genet. 1997;15(1):21–9.
McDermott DA, et al. TBX5 genetic testing validates strict clinical criteria for Holt-Oram syndrome. Pediatr Res. 2005;58(5):981–6.
Elek C, Vitez M, Czeizel E. Holt-Oram syndrome. Orv Hetil. 1991;132(2):73–4.. 77-78
Barisic I, et al. Holt Oram syndrome: a registry-based study in Europe. Orphanet J Rare Dis. 2014;9:156.
Holt M, Oram S. Familial heart disease with skeletal malformations. Br Heart J. 1960;22:236–42.
Basson CT, et al. The clinical and genetic spectrum of the Holt-Oram syndrome (heart-hand syndrome). N Engl J Med. 1994;330(13):885–91.
Newbury-Ecob RA, Leanage R, Raeburn JA, Young ID. Holt-Oram syndrome: a clinical genetic study. J Med Genet. 1996;33(4):300–7.
Huang T, et al. Causes of clinical diversity in human TBX5 mutations. Cold Spring Harb Symp Quant Biol. 2002;67:115–20.
Bohm J, et al. Functional analysis of the novel TBX5 c.1333delC mutation resulting in an extended TBX5 protein. BMC Med Genet. 2008;9:88.
Boogerd CJ, et al. Functional analysis of novel TBX5 T-box mutations associated with Holt-Oram syndrome. Cardiovasc Res. 2010;88(1):130–9.
Bruneau BG, et al. A murine model of Holt-Oram syndrome defines roles of the T-box transcription factor Tbx5 in cardiogenesis and disease. Cell. 2001;106(6):709–21.
Steimle JD, et al. Evolutionarily conserved Tbx5-Wnt2/2b pathway orchestrates cardiopulmonary development. Proc Natl Acad Sci U S A. 2018;115(45):E10615–24.
Peng T, et al. Coordination of heart and lung co-development by a multipotent cardiopulmonary progenitor. Nature. 2013;500(7464):589–92.
Nadadur RD, et al. Pitx2 modulates a Tbx5-dependent gene regulatory network to maintain atrial rhythm. In: Sci Transl Med 8(354):354ra115; 2016.
Tripathi S, et al. Meta- and orthogonal integration of influenza “OMICs” data defines a role for UBR4 in virus budding. Cell Host Microbe. 2015;18(6):723–35.
Bapat A, Anderson CD, Ellinor PT, Lubitz SA. Genomic basis of atrial fibrillation. Heart. 2018;104(3):201–6.
van Setten J, et al. PR interval genome-wide association meta-analysis identifies 50 loci associated with atrial and atrioventricular electrical activity. Nat Commun. 2018;9(1):2904.
Kramer A, Green J, Pollard J Jr, Tugendreich S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics. 2014;30(4):523–30.
Goette A, et al. Increased expression of extracellular signal-regulated kinase and angiotensin-converting enzyme in human atria during atrial fibrillation. J Am Coll Cardiol. 2000;35(6):1669–77.
Brundel BJ, et al. Alterations in potassium channel gene expression in atria of patients with persistent and paroxysmal atrial fibrillation: differential regulation of protein and mRNA levels for K+ channels. J Am Coll Cardiol. 2001;37(3):926–32.
Barth AS, et al. Reprogramming of the human atrial transcriptome in permanent atrial fibrillation: expression of a ventricular-like genomic signature. Circ Res. 2005;96(9):1022–9.
Lamirault G, et al. Gene expression profile associated with chronic atrial fibrillation and underlying valvular heart disease in man. J Mol Cell Cardiol. 2006;40(1):173–84.
Cardin S, et al. Contrasting gene expression profiles in two canine models of atrial fibrillation. Circ Res. 2007;100(3):425–33.
De Souza AI, et al. Proteomic and metabolomic analysis of atrial profibrillatory remodelling in congestive heart failure. J Mol Cell Cardiol. 2010;49(5):851–63.
Riley G, Syeda F, Kirchhof P, Fabritz L. An introduction to murine models of atrial fibrillation. Front Physiol. 2012;3:296.
Deshmukh A, et al. Left atrial transcriptional changes associated with atrial fibrillation susceptibility and persistence. Circ Arrhythm Electrophysiol. 2015;8(1):32–41.
Mahida S. Transcription factors and atrial fibrillation. Cardiovasc Res. 2014;101(2):194–202.
Gudbjartsson DF, et al. Variants conferring risk of atrial fibrillation on chromosome 4q25. Nature. 2007;448(7151):353–7.
Chinchilla A, et al. PITX2 insufficiency leads to atrial electrical and structural remodeling linked to arrhythmogenesis. Circ Cardiovasc Genet. 2011;4(3):269–79.
Kirchhof P, et al. PITX2c is expressed in the adult left atrium, and reducing Pitx2c expression promotes atrial fibrillation inducibility and complex changes in gene expression. Circ Cardiovasc Genet. 2011;4(2):123–33.
Wang J, et al. Pitx2 prevents susceptibility to atrial arrhythmias by inhibiting left-sided pacemaker specification. Proc Natl Acad Sci U S A. 2010;107(21):9753–8.
Tao Y, et al. Pitx2, an atrial fibrillation predisposition gene, directly regulates ion transport and intercalated disc genes. Circ Cardiovasc Genet. 2014;7(1):23–32.
Mommersteeg MT, et al. Pitx2c and Nkx2-5 are required for the formation and identity of the pulmonary myocardium. Circ Res. 2007;101(9):902–9.
Soufan AT, et al. Reconstruction of the patterns of gene expression in the developing mouse heart reveals an architectural arrangement that facilitates the understanding of atrial malformations and arrhythmias. Circ Res. 2004;95(12):1207–15.
Huang RT, Xue S, Xu YJ, Zhou M, Yang YQ. A novel NKX2.5 loss-of-function mutation responsible for familial atrial fibrillation. Int J Mol Med. 2013;31(5):1119–26.
Xie WH, et al. Prevalence and spectrum of Nkx2.5 mutations associated with idiopathic atrial fibrillation. Clinics (Sao Paulo). 2013;68(6):777–84.
Qiu XB, et al. PITX2C loss-of-function mutations responsible for idiopathic atrial fibrillation. Clinics (Sao Paulo). 2014;69(1):15–22.
Wang J, et al. Pitx2-microRNA pathway that delimits sinoatrial node development and inhibits predisposition to atrial fibrillation. Proc Natl Acad Sci U S A. 2014;111(25):9181–6.
Christoffels VM, et al. Formation of the venous pole of the heart from an Nkx2-5-negative precursor population requires Tbx18. Circ Res. 2006;98(12):1555–63.
Xie L, et al. Tbx5-hedgehog molecular networks are essential in the second heart field for atrial septation. Dev Cell. 2012;23(2):280–91.
Ye W, et al. A common Shox2-Nkx2-5 antagonistic mechanism primes the pacemaker cell fate in the pulmonary vein myocardium and sinoatrial node. Development. 2015;142(14):2521–32.
Ventura A, et al. Restoration of p53 function leads to tumour regression in vivo. Nature. 2007;445(7128):661–5.
Kim D, et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013;14(4):R36.
Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 2009;25(9):1105–11.
Barnett DW, Garrison EK, Quinlan AR, Stromberg MP, Marth GT. BamTools: a C++ API and toolkit for analyzing and managing BAM files. Bioinformatics. 2011;27(12):1691–2.
Pertea M, Kim D, Pertea GM, Leek JT, Salzberg SL. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat Protoc. 2016;11(9):1650–67.
Pertea M, et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol. 2015;33(3):290–5.
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing. 2017.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26(1):139–40.
Ritchie ME, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43(7):e47.
Wickham H. ggplot2: Elegant graphics for data analysis. New York: Springer; 2009.
Wickham H. Scales: Scale Functions for Visualization. R package version 0.5.0. 2017.
Nowak KJ, et al. Mutations in the skeletal muscle alpha-actin gene in patients with actin myopathy and nemaline myopathy. Nat Genet. 1999;23(2):208–12.
Perera D, et al. Bub1 maintains centromeric cohesion by activation of the spindle checkpoint. Dev Cell. 2007;13(4):566–79.
Christodoulou A, Lederer CW, Surrey T, Vernos I, Santama N. Motor protein KIFC5A interacts with Nubp1 and Nubp2, and is implicated in the regulation of centrosome duplication. J Cell Sci. 2006;119(Pt 10):2035–47.
Chan GK, Jablonski SA, Starr DA, Goldberg ML, Yen TJ. Human Zw10 and ROD are mitotic checkpoint proteins that bind to kinetochores. Nat Cell Biol. 2000;2(12):944–7.
Barton PJ, et al. The myosin alkali light chains of mouse ventricular and slow skeletal muscle are indistinguishable and are encoded by the same gene. J Biol Chem. 1985;260(14):8578–84.
Moldovan GL, et al. Inhibition of homologous recombination by the PCNA-interacting protein PARI. Mol Cell. 2012;45(1):75–86.
Lu X, et al. The cytokine-driven regulation of secretoglobins in normal human upper airway and their expression, particularly that of uteroglobin-related protein 1, in chronic rhinosinusitis. Respir Res. 2011;12:28.
Kitamura T, Takagi S, Naganuma T, Kihara A. Mouse aldehyde dehydrogenase ALDH3B2 is localized to lipid droplets via two C-terminal tryptophan residues and lipid modification. Biochem J. 2015;465(1):79–87.
Tomsig JL, Creutz CE. Copines: a ubiquitous family of Ca(2+)-dependent phospholipid-binding proteins. Cell Mol Life Sci. 2002;59(9):1467–77.
Fagerberg L, et al. Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol Cell Proteomics. 2014;13(2):397–406.
Yue F, et al. A comparative encyclopedia of DNA elements in the mouse genome. Nature. 2014;515(7527):355–64.
Yoshida H, et al. KIAA1199, a deafness gene of unknown function, is a new hyaluronan binding protein involved in hyaluronan depolymerization. Proc Natl Acad Sci U S A. 2013;110(14):5612–7.
Eyre D. Collagen of articular cartilage. Arthritis Res. 2002;4(1):30–5.
Erle DJ, Ruegg C, Sheppard D, Pytela R. Complete amino acid sequence of an integrin beta subunit (beta 7) identified in leukocytes. J Biol Chem. 1991;266(17):11009–16.
Pang YL, Poruri K, Martinis SA. tRNA synthetase: tRNA aminoacylation and beyond. Wiley Interdiscip Rev RNA. 2014;5(4):461–80.
Smith CL, et al. Mouse Genome Database (MGD)-2018: knowledgebase for the laboratory mouse. Nucleic Acids Res. 2018;46(D1):D836–42.
Lopez Jimenez N, et al. Examination of FGFRL1 as a candidate gene for diaphragmatic defects at chromosome 4p16.3 shows that Fgfrl1 null mice have reduced expression of Tpm3, sarcomere genes and Lrtm1 in the diaphragm. Hum Genet. 2010;127(3):325–36.
Jisaka M, Kim RB, Boeglin WE, Nanney LB, Brash AR. Molecular cloning and functional expression of a phorbol ester-inducible 8S-lipoxygenase from mouse skin. J Biol Chem. 1997;272(39):24410–6.
Lu Z, et al. Molecular Architecture of Contactin-associated Protein-like 2 (CNTNAP2) and Its Interaction with Contactin 2 (CNTN2). J Biol Chem. 2016;291(46):24133–47.
Stevens DR, et al. Hyperpolarization-activated channels HCN1 and HCN4 mediate responses to sour stimuli. Nature. 2001;413(6856):631–5.
Darzacq X, et al. Cajal body-specific small nuclear RNAs: a novel class of 2′-O-methylation and pseudouridylation guide RNAs. EMBO J. 2002;21(11):2746–56.
Burgoyne RD. Neuronal calcium sensor proteins: generating diversity in neuronal Ca2+ signalling. Nat Rev Neurosci. 2007;8(3):182–93.
DeChiara TM, Efstratiadis A, Robertson EJ. A growth-deficiency phenotype in heterozygous mice carrying an insulin-like growth factor II gene disrupted by targeting. Nature. 1990;345(6270):78–80.
Schluter OM, Schmitz F, Jahn R, Rosenmund C, Sudhof TC. A complete genetic analysis of neuronal Rab3 function. J Neurosci. 2004;24(29):6629–37.
Steinberg SJ, Wang SJ, McGuinness MC, Watkins PA. Human liver-specific very-long-chain acyl-coenzyme A synthetase: cDNA cloning and characterization of a second enzymatically active protein. Mol Genet Metab. 1999;68(1):32–42.
Huang W, et al. The long non-coding RNA SNHG3 functions as a competing endogenous RNA to promote malignant development of colorectal cancer. Oncol Rep. 2017;38(3):1402–10.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2020 The Author(s)
About this paper
Cite this paper
Steimle, J.D., Laforest, B., Nadadur, R.D., Broman, M.T., Moskowitz, I.P. (2020). Regional and TBX5-Dependent Gene Expression in the Atria: Implications for Pulmonary Vein Development and Atrial Fibrillation. In: Nakanishi, T., Baldwin, H., Fineman, J., Yamagishi, H. (eds) Molecular Mechanism of Congenital Heart Disease and Pulmonary Hypertension. Springer, Singapore. https://doi.org/10.1007/978-981-15-1185-1_51
Download citation
DOI: https://doi.org/10.1007/978-981-15-1185-1_51
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-1184-4
Online ISBN: 978-981-15-1185-1
eBook Packages: MedicineMedicine (R0)