Abstract
Despite advances in screening, therapy, and surveillance that have improved patient survival rates, breast cancer is still the most commonly diagnosed cancer and the second leading cause of cancer mortality among women [1]. Breast cancer is a highly heterogeneous disease rooted in a genetic basis, influenced by extrinsic stimuli, and reflected in clinical behavior. The diversity of breast cancer hormone receptor status and the expression of surface molecules have guided therapy decisions for decades; however, subtype-specific treatment often yields diverse responses due to varying tumor evolution and malignant potential. Although the mechanisms behind breast cancer heterogeneity is not well understood, available evidence suggests that studying breast cancer metabolism has the potential to provide valuable insights into the causes of these variations as well as viable targets for intervention.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
- Breast cancer
- Estrogen receptor status
- Metabolic fingerprint
- Choline metabolism
- Estrogen metabolism
- Serine biosynthesis
- Glycolytic upregulation
- Intratumoral heterogeneity
- Metabolic adaptivity
-
Aberrant metabolic pathways present in breast cancer contribute to breast cancer heterogeneity.
-
Differences in glycolytic upregulation among breast cancer subtypes can be attributed to GLUT expression.
-
Choline metabolism in breast cancer is strongly associated with tumor grades.
-
Metabolic profiling of breast cancers can be used for clinical breast cancer diagnosis and prediction of recurrence or metastasis.
-
Breast cancer metabolism has heterogeneous and adaptive characteristics from a spatial and temporal basis.
-
Metabolic adaptability confers chemotherapy-resistant phenotypes and promotes tumor evolution.
1 Introduction
Despite advances in screening, therapy, and surveillance that have improved patient survival rates, breast cancer is still the most commonly diagnosed cancer and the second leading cause of cancer mortality among women [1]. Breast cancer is a highly heterogeneous disease rooted in a genetic basis, influenced by extrinsic stimuli, and reflected in clinical behavior. The diversity of breast cancer hormone receptor status and the expression of surface molecules have guided therapy decisions for decades; however, subtype-specific treatment often yields diverse responses due to varying tumor evolution and malignant potential. Although the mechanisms behind breast cancer heterogeneity is not well understood, available evidence suggests that studying breast cancer metabolism has the potential to provide valuable insights into the causes of these variations as well as viable targets for intervention.
2 Aberrant Metabolic Pathways Present in Breast Cancer Contribute to Breast Cancer Heterogeneity (Fig. 1)
In order to sustain tumorigenic proliferation, cancer cells exploit diverse metabolic pathways. The diversity of hormone receptors present within breast cancer cells is classified into different subtypes. Breast cancers with hormone-positive receptors such as estrogen receptors (ER) and progesterone receptors (PR) rely on their respective hormones for growth. Patients with HER2+ breast cancer have overexpression of human epidermal growth factor receptor 2 (HER2). Patients negative for all three receptors are considered to have triple-negative breast cancer (TNBC)—the most heterogeneous molecular profile. This diversity, in turn, reflects the different metabolic phenotypes of breast cancer. Some of these core metabolic aberrations have fundamental effects on breast cancer tumorigenicity and offer rationale behind the aggressiveness of specific subtypes. Tumor evolution results in the reprogramming of cell metabolism in order to adapt to support cell proliferation. Specific mutations in oncogenes and tumor-suppressor genes are hypothesized to cause metabolic reprogramming within different breast cancer subtypes. Although several mutations are commonly seen in breast cancers, they appear in various combinations that are reflective of the diverse metabolic behaviors of breast cancers. For example, mutations in BRAF, KRAS, and HRAS were found to be metabolic regulators of TNBC [2]. These genetic alterations are known to regulate glutamine metabolism, which renders cancer cells dependent on glutamine for proliferation and survival [3, 4]. The BRCA1 mutation is a good example of how genetic alterations lend to specific metabolic phenotypes that promote tumorigenesis. A study by Martinez-Outschoorn et al. showed that loss-of-function mutations in the BRCA1 tumor-suppressor gene resulted in the production of hydrogen peroxide and oxidative stress in epithelial breast cancer cells and stromal fibroblasts [5]. This loss of function also causes elevated expression of monocarboxylate transporter 4 (a functional marker of oxidative stress and glycolytic activity) to shuttle l-lactate out of cells.
Furthermore, the loss of caveolin-1 in cancer-associated fibroblasts is associated with elevated production of reactive oxygen species (ROS) and increased glycolysis in stromal cells, both of which play a fundamental role in tumorigenesis [6]. This encourages therapeutic targeting of cancer-associated fibroblasts that favor cancer progression [7]. Mutations in BRCA1 are marked by high rates of proliferation and substantial cellular inflammation. This study suggests that antioxidant agents present promising therapies for BRCA1-mutated breast cancer.
2.1 Differences in Glycolytic Upregulation Among Breast Cancer Subtypes Can Be Attributed to Glucose Transporter (GLUT) Expression
First postulated by Otto Warburg in 1927 [8] and firmly established in the literature thereafter, a hallmark of cancer malignancy is an upregulation in aerobic glycolysis even in the presence of oxygen, known as the Warburg effect [9, 10]. Lactate dehydrogenase A, a key enzyme of the Warburg effect that catalyzes the conversion of pyruvate to lactate, has been a studied target in several cancers [11,12,13]. Breast cancer tumors are no exception to the Warburg effect; however, there are variations in glycolytic rates and metabolite-related protein expression among breast cancer subtypes that correlate with tumor aggressiveness. Previous in vitro studies have first observed that the glucose-dependent MCF-7 cell line is more sensitive to FX11, a lactate dehydrogenase A inhibitor, than the non-glucose-dependent MDA-MB-453 cell line [11]. Another in vitro study found that noninvasive breast cancer cell lines showed a significantly lower rate of glucose intake compared to more aggressive, metastatic cells [14]. Higher rates of glucose uptake are accompanied by altered gene expression and translation of metabolism-related proteins as well. Glucose transporter (GLUT) expression has been studied extensively in breast cancer. GLUTs are integral transmembrane proteins that facilitate glucose delivery across the plasma membrane. They serve as a rate-limiting step that controls the amount of glucose accessible to the cell [15]. Studies have shown that different isoforms of GLUTs have been detected and/or overexpressed in breast cancer cells. Different GLUT expression patterns are found to be associated with various pathological grades and tumor aggressiveness in patient-derived samples. Choi et al. discovered that GLUT1, one of the isoforms of the GLUT family, had the highest expression in the TNBC subtype and tumors with high histologic grade [16]. As a result of increased glucose uptake, the increased rate of glycolysis subjects the cell to intracellular lactic acidosis—leading to cell death. Interestingly, the same group showed that TNBC had the highest expression of carbonic anhydrase IX, an enzyme that prevents acidosis and provides TNBC with an acid-resistant phenotype [16], suggesting that aggressive breast cancer subtypes adopt metabolic phenotypes able to suppress apoptosis. GLUT1 overexpression has also been linked to invasiveness in breast cancer [17].
The link between metabolic reprogramming and protein expression offers an adaptive advantage that contributes to a level of aggression specific to certain subtypes of breast cancer like TNBC, making them characteristically resilient and harder to treat.
2.2 Choline Metabolism in Breast Cancer Is Strongly Associated with Tumor Grades
The deregulation of choline metabolism and elevated levels of choline-containing compounds are frequently observed in breast cancer progression [18,19,20,21]. Choline plays an important role in supplying methyl groups through its metabolism and is essential for cellular structure as a precursor of phospholipids. Choline metabolism in breast tissue is distributed between two central pathways: (1) the biosynthesis of phosphatidylcholine (PtdCho) known as the Kennedy pathway and (2) the oxidation to betaine, a methyl group donor in many methylation reactions. A study by Katz-Brull et al. revealed that breast cancer cells exhibited a higher choline transport rate compared to normal breast cells, and a majority of the choline was converted to phosphocholine (Pcho) through the Kennedy pathway while around approximately 25% was oxidized to betaine [19]. Although levels of phospholipid-related metabolites are enhanced in most breast cancers [22], significantly higher levels of Pcho were found to be associated with ER tumors and the more aggressive histologic grade 3 tumors [23]. Because of this, choline-containing compounds have often been seen as biomarkers for breast tumor malignancy. Oncogenic expression of choline kinase (CK), the enzyme responsible for the conversion of choline to Pcho, is responsible for elevated levels of Pcho in breast cancer cells [19]. Furthermore, CK also showed a strong association with high histologic grade and ER− subtypes [24]. For this reason, CK is an attractive antitumor target for subsequent studies. Whether or not choline metabolism represents an agent of disease progression or merely a marker for transformation has still not been defined. CK inhibitors blocking choline metabolism have shown promising antitumor results. A study by Rodríguez-González et al. discovered that blocking the enzyme had no effect on normal cells but disrupted phospholipid production in tumor cells—resulting in apoptosis due to the accumulation of cytotoxic ceramide, the simplest class of sphingolipids [25].
3 Different Roles of Estrogen in Estrogen Metabolism and ER Binding Promote Breast Cancer Tumorigenicity
Endogenous estrogens and their metabolism have been linked to breast carcinogenesis, especially in postmenopausal women [26]. 17b-Estradiol (E2), the main estrogen in breast tissue, acts as both a ligand for ER and a substrate in metabolism—roles which contribute to estrogen as a carcinogen. The mechanism of estrogen carcinogenesis is a combination of ER signaling and estrogen metabolism.
ERs, when activated, are responsible for the mediation of many downstream signaling pathways that function as transcription factors promoting cancer development [27]. In addition, ER signaling interacts with growth factor receptors and other signaling molecules to promote growth and anti-apoptotic signals [28]. ER activation has also been shown to promote downstream reprogramming in choline metabolism, an aberration in breast cancer [29].
As a substrate, the metabolism of estrogen through the 4-hydroxylation pathway produces specific catechol estrogens and estrogen quinones known to be carcinogenic. Estrogen is hydroxylated by cytochrome P450 enzymes and shuttled into three main pathways depending on the three different carbons hydroxylated: C2, C3, and C16. The catechol estrogens (2-OH E1, 2-OH E2, 4-OH E1, 4-OH E2) are either methylated by catechol-O-transferase (COMT), thereby reducing their mutational potential, or oxidized further to semiquinones or quinones. 4-OH catechol estrogen, when oxidized to a reactive estrogen quinone, leads to DNA damage by forming unstable DNA adducts between adenine and guanine nucleotides [30, 31]. Mutations caused by this mechanism have the potential to initiate breast cancer or increase cancer risk. In contrast, metabolites formed through the 2-OH pathway form stable DNA adducts and are anticarcinogenic—dubbing the 2-OH metabolites as “the good estrogen” in some cases [32]. Protective mechanisms such as estrogen quinone conjugation with glutathione via glutathione S-transferase P (GSTP) help lower the risk of cancerous mutations by detoxifying the estrogen quinones [30]. However, estrogenic imbalances lead to competition between the pathway forming the unstable DNA adducts and the detoxification of its cancer-promoting substrates [30]. Accordingly, hormone therapy for breast cancer has targeted ER+ subtypes with drugs such as tamoxifen, which acts as a competitive inhibitor that prevents estrogen from binding to the ER. Another important class of drugs inhibits aromatase, an important rate-limiting enzyme that converts androgens to estrogens, to lower estrogen levels in the body.
3.1 PHGDH Overexpression in Serine Biosynthesis Fuels TCA Anaplerosis
Serine biosynthesis is an essential pathway for breast cancer progression in specific subsets of breast tumors. Using RNAi-based loss-of-function screening, Possemato et al. identified phosphoglycerate dehydrogenase (PHGDH) in breast cancer with enhanced protein levels in 70% of aggressive ER− subtypes [33]. PHGDH catalyzes the committed-limiting step that oxidizes 3-phosphoglycerate (3PG) to 3-phosphohydroxypyruvate (3HP) substrates in the serine synthesis pathway. Enhanced PHGDH expression was associated with increased serine synthesis and glutamine uptake. Suppression of PHGDH expression led to a significant decrease in cell proliferation but did not affect intracellular serine levels; instead, researchers found a resulting drop in phosphoserine aminotransferase 1 (PSAT1)-dependent alpha-ketoglutarate (α-KG), an output of the serine pathway [33]. In cancer cells with overexpression of PHGDH, the serine synthesis pathway plays an important role in tricarboxylic acid (TCA) cycle anaplerosis—supplying α-KG to support cell proliferation [33]. In addition, suppression of PSAT1 and phosphoserine phosphatase (PSPH) enzymes downstream in the serine pathway inhibits cell proliferation in PHGDH-enhanced cell lines as well [33]. Subsequent studies have revealed that in addition to 3PG oxidation, PHGDH also catalyzes the reduction of α-KG to d-2-hydroxyglutarate (D-2HG) [34], an established oncometabolite [35, 36]. D-2HG acted as a competitive inhibitor of α-KG-dependent dioxygenases, resulting in aberrations in histone methylation and DNA hypermethylation [37]. High levels of D-2HG and N-acetyl-aspartate were found to accumulate preferentially in ER− and basal-like tumors, which may contribute to their aggressive phenotypes [36] in contrast to the mixed effects on glioblastoma [38] and other cancers. In vitro experiments revealed that accumulation of D-2HG is associated with increased cell proliferation and inhibited apoptosis [36]; however, the oncogenic effects of D-2HG on breast cancer still need to be defined. Because of its deregulated expression and oncogenic effects, PHGDH is considered a promising target for therapy in breast cancers that exhibit PHGDH overexpression. Although a preliminary PHGDH inhibitor has been recently developed [39], PHGDH-targeted therapy is still in its infancy.
Using a novel computational method, Jerby et al. contributed further evidence that the metabolic profiles of ER+ and ER− subtypes are vastly different [40]. The stoichiometric analysis revealed serine metabolism to be coupled with glutamine uptake [40]. ER+ tumors exhibit a stronger preference for glutamine biosynthesis and secretion than ER− tumors [40]. In addition, their model identified ER+ phenotypes as having more capacity to convert glucose to lactate than ER− tumors. Due to higher rates of serine metabolism, ER− subtypes are rationalized to preferably divert 3PG toward serine metabolism via PHGDH to exploit alternative pathways for glutaminolysis [40]. In addition, a high MYC overexpression [41, 42], and low thioredoxin-interacting protein expression, an inhibitor of glucose utilization, was found to be a characteristic gene signature of TNBC and no other subtypes [43].
4 The Clinical Applications of Metabolic Profiling
Metabolic profiling has garnered much research interest within the past decade [44]. Although the mechanisms behind breast cancer transformation have not been firmly established, changes in tumor evolution have been investigated through metabolic variation. The exploitation of these metabolic signatures has the potential to improve clinical results through diagnosis confirmation, early detection, and prediction of disease progression [45, 46] (Fig. 2).
4.1 Breast Cancer Diagnosis and Subtyping Using Metabolomics
Studies have used metabolic profiling for the general diagnosis of breast cancer—using different techniques to build prediction models that distinguish specific metabolic fingerprints of breast cancer hormone receptor status, histologic grade, and axillary lymphatic spread [47,48,49]. Jove et al. used a combination of random forest classification and multivariate statistics to identify combinations of metabolites that were used to distinguish breast cancer plasma samples from healthy control samples [47]. On the other hand, Huang et al. sought out a model more tolerant of breast cancer heterogeneity by following metabolic pathways rather than metabolite-based biomarkers for early diagnosis of breast cancer [48]. Other studies have used metabolic profiling to build models to distinguish breast cancer stages [49] and levels of malignancy [50].
4.2 Metabolic Profiling as a Strategy for Prediction of Recurrence in Breast Cancer
Recurrence after initial therapy causes significant morbidity and mortality in breast cancer patients. Current methods for detecting recurrences such as medical imaging and serum tumor markers are not considered specific enough to be routinely recommended; therefore, there is still much room for improvement. A combination of nuclear magnetic resonance (NMR) and mass spectroscopy (MS) analysis and multivariate statistics on patient serum samples has been used to explore potential metabolic profiles sensitive to cancer recurrence [51]. Asiago et al. developed a prediction model built upon 11 biomarkers that correctly detected 55% of patients with breast cancer recurrence an average of 13 months prior to their clinical diagnosis using serum samples [51]. Although there is vast room for improvement on more specific and accurate models for early detection of recurrence, metabolic profiling of serum can be viewed as a promising noninvasive method for breast cancer surveillance.
Summary of the different levels of metabolic heterogeneity seen in breast cancer. Metabolic heterogeneity is demonstrated between molecular subtypes of breast cancer, between tumor core and periphery, in different stages of cancer progression, and in response to selective pressures from clinical treatment
4.3 Metabolic Fingerprinting in Breast Cancer Metastasis
Oakman et al. identified a preliminary metabolic fingerprint from patient serum samples that detected early and metastatic disease in breast cancer patients. In their study, higher levels of phenylalanine, glucose, proline, lysine, and N-acetyl cysteine and lower levels of lipids contributed to the metabolic profile of metastatic individuals [52]. Jobard et al. used similar serum NMR analysis to identify metabolic profiles between localized and metastatic breast cancer. They found eight statistically significant elevations of metabolite biomarkers in metastatic disease: histidine, acetoacetate, glycerol, pyruvate, N-acetyl glycoproteins, mannose, glutamate, and phenylalanine [53]. Although there are differences in biomarkers between the two studies, it is notable that the same trends of variation in glucose concentration and lowered lipid levels were seen between early and metastatic breast cancer [53]. Defining an accurate metabolic fingerprint specific across all metastatic breast cancers is a challenge due to the variability and high mutational load of metastatic disease. Under changing tumor microenvironments [54], metastatic breast cancer cells readily switch between glycolysis and oxidative phosphorylation [55]. The metabolic plasticity of metastatic breast cancer may contribute to the inconsistencies of biomarkers across different tumors. However, studies attempting to identify these metabolic patterns provide great insights into the general characteristics of advanced diseases.
4.4 Prediction of Response to Therapy Based on Metabolic Phenotypes
Metabolic fingerprinting has also been used to predict responses to therapy and drug resistance. Using a combination of NMR and liquid chromatography-mass spectrometry (LC-MS), Wei et al. were able to identify four altered metabolites (threonine, glutamine, isoleucine, and linolenic acid) as indicators of adjuvant chemotherapy response within breast cancer [56]. A prediction model derived from these metabolic markers was able to distinguish between complete, partial, and no tumor response to chemotherapy in a neoadjuvant setting using patient samples [56]. The model was able to correctly identify 80% of patients whose tumors did not show a complete pathologic response to chemotherapy [56]. Collectively, these studies highlight the potential impact of metabolic profiling on the integration of metabolomics into clinical practice. Further advancements in profiling could improve diagnosis and early detection or at least offer confirmation in the treatment of breast cancer quickly and at low cost. Although most of the prediction models and metabolic phenotypes presented in these studies are in their preliminary stages, improvements could make way for more individualized treatments specific to each patient.
5 Additional Perspectives on Breast Cancer Heterogeneity
5.1 Spatial Pathogenesis Observed in Breast Cancer Metabolism
Metabolic heterogeneity within a single tumor adds another layer of complexity when trying to understand the dynamic processes of breast cancer metabolism [57]. Several studies have identified metabolic distinctions between breast tumor periphery and center. A study by Xu et al. analyzed the mitochondrial redox states of breast cancer xenografts of varying aggressiveness. In general, the researchers found more oxidized metabolic states in central regions and more reduced states in peripheral regions of the tumors [58]. The tumors also exhibited higher glucose uptake and NADH levels in tumor peripheries compared to the centers [58]. The authors presumed this was due to higher substrate availability at the peripheries from the tumor microenvironment. Furthermore, higher degrees of metabolic heterogeneity were consistently observed in larger and higher staged tumors [58, 59]. When comparing metabolic profiles of clinical breast tumor samples, studies have observed higher levels of Pcho and phosphoethanolamine in the tumor core compared to tumor periphery [60]. Lactate and pyruvate were observed in higher levels in the tumor periphery compared to the tumor core [61]. Because the tumor periphery has direct interactions with the tumor microenvironment compared to the center, the differences in inputs translate into differences in metabolic phenotypes. In normal breast anatomy, the epithelia receive similar concentrations of oxygen, growth factors, and nutrients. The anatomic disorganization caused by breast cancer pathology alters the tumor microenvironment and intratumoral metabolism [57, 62]. It is unclear whether these observed differences are solely due to extrinsic inputs, genetic manifestations, or perhaps an interplay between both.
5.2 Temporal Pathogenesis Observed in Breast Cancer Metabolism: Metabolic Differences Between Early Stage and Advanced Stage
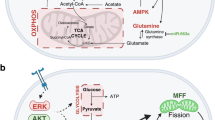
Temporal pathogenesis refers to tumor progression over time, starting from a single cancer cell to the formation of a primary tumor and then metastatic spread. The epithelial-to-mesenchymal transition (EMT) is one of the prerequisites of early metastasis. It describes the transition in which epithelial cells lose their polarity and cell adhesions to become mesenchymal cells with migratory properties. Cancer cells detach from their extracellular matrix (ECM) when they decide to metastasize. It has been shown in vivo that mammary epithelial cells with lost ECM attachment are unable to survive due to ATP deficiency from glucose deprivation [63]. ECM detachment is also accompanied by increased ROS [63, 64]. Overexpression of the HER2 oncogene rescued these cells by restoring glucose uptake and reducing ROS through the oxidative pentose phosphate pathway (PPP) [63]. Nicotinamide adenine dinucleotide phosphate (NADPH), a product of the PPP, serves as a reducing agent able to combat oxidative stress. The study by Schafer et al. also showed that the treatment of antioxidants alone was able to rescue matrix-detached cells—identifying oxidative resistance as an important property needed for metastatic migration [63]. Once cancer cells detach from the ECM, they will need to survive the journey in the oxidizing bloodstream. Many cells will undergo apoptosis in this environment, but the cells that acquire oxidative resistance have the adaptive advantage to metastasize. Although aerobic glycolysis is the most well-known hallmark of cancer metabolism [10], research has also identified the importance of oxidative phosphorylation in cancer progression as well [65]. Increased mitochondrial biogenesis and respiration have been observed in cancer cell metastases through the modulation of peroxisome proliferation-activated receptor gamma coactivator-1a (PGC-1a)—a regulator of mitochondrial biogenesis and energy metabolism [66]. This is corroborated by the correlation between PGC-1a expression and formation of distant metastasis from patient breast tumors and breast cancer cell lines [66]. These changes in metabolic phenotype seen in migrating breast cancer cells are examples that highlight the importance of metabolic plasticity for cancer progression.
5.3 Metabolic Heterogeneity Influences Effective Breast Cancer Drug Treatment
In silico modeling of tumor progression by Robertson-Tessi et al. proposed that early stages of tumor growth have a stratified composition. Tumor centers have higher glycolytic activity and are, therefore, more aggressive compared to the periphery [67]. It is argued that cancer treatments should decrease or slow selective pressures in cells through the maintenance of less aggressive cancer cells within a tumor rather than aiming for eradication [67, 68]. This concept is demonstrated in the antiangiogenic treatment of breast cancer. Antiangiogenic therapies aim to starve cancer cells of oxygen and nutrients by inhibiting tumor vascularization, creating pockets of intratumoral hypoxia. Cutting off nutrient supply in this way may be effective in stopping cancer cell growth, but it may also select for cells that are able to alter metabolism to adapt to hypoxic conditions [54], resulting in a drug-resistant phenotype. Although antiangiogenic therapies in breast cancer patients have been able to lengthen progression-free survival, data has shown that it does little to improve overall patient survival [69]. Aggressive relapse and enhanced metastasis in treated patients are not uncommon either [70]. Conley et al. were able to show that breast cancer xenografts, when treated with antiangiogenic drugs, developed hypoxia-driven cancer stem cell stimulation, which promoted tumorigenesis—opposite to the intended effect [70]. It would seem that the goal of chemotherapy is to halt tumorigenesis and shrink existing tumor populations as quickly and effectively as possible by delivering the drug at the highest dosage allowed. This objective, however, is a double-edged sword: if treatment is too aggressive, it puts selective pressure on the cells to enhance drug-resistant phenotypes that, in turn, escalate cancer progression. The adaptive nature of cancer metabolism is a significant obstacle for creating effective drug therapies. An effective treatment aims to find the delicate balance of delivering maximum cytotoxic effects while avoiding selective resistance.
6 Conclusion
Metabolomics serves as an essential utility in breast cancer research by offering a perspective that represents the net interactions between the tumor, the host, and the environment and within the tumor itself. The metabolic nuances across different breast cancer subtypes and treatment timelines can be taken advantage of when thinking about potential prognostic markers, prediction models, and mechanisms involved with breast cancer. Metabolic heterogeneity in breast cancer can be seen within a single tumor and in the different stages of the tumor’s progression (Fig. 3). Understanding these dynamic processes and applying them to drug discovery and clinical practice have the potential to improve the lives of not only breast cancer patients but also all cancer patients.
Change history
22 October 2021
After initial publication of the book, various errors were identified that needed correction. All corrections listed below have been updated within the current version.
Abbreviations
- 3HP:
-
3-Phosphohydroxypyruvate
- 3PG:
-
3-Phosphoglycerate
- CK:
-
Choline kinase
- COMT:
-
Catechol-O-methyltransferase
- D-2HG:
-
d-2-Hydroxyglutarate
- E2:
-
17b-Estradiol
- ECM:
-
Extracellular matrix
- EMT:
-
Epithelial-to-mesenchymal transition
- ER:
-
Estrogen receptor
- GLUT:
-
Glucose transporter
- GSTP:
-
Glutathione S-transferase P
- HER2:
-
Human epidermal growth factor receptor 2
- LCMS:
-
Liquid chromatography mass spectrometry
- MS:
-
Mass spectrometry
- NADPH:
-
Nicotinamide adenine dinucleotide phosphate
- NMR:
-
Nuclear magnetic resonance
- PCho:
-
Phosphocholine
- PGC-1a:
-
Peroxisome proliferation-activated receptor gamma coactivator-1a
- PHGDH:
-
Phosphoglycerate dehydrogenase
- PPP:
-
Pentose phosphate pathway
- PR:
-
Progesterone receptor
- PSAT1:
-
Phosphoserine aminotransferase 1
- PSPH:
-
Phosphoserine phosphatase
- PtdCho:
-
Phosphatidylcholine
- ROS:
-
Reactive oxygen species
- TCA:
-
Tricarboxylic acid
- TNBC:
-
Triple-negative breast cancer
- α-KG:
-
Alpha-ketoglutarate
References
Gutierrez, T., et al. (2013). IL-21 promotes the production of anti-DNA IgG but is dispensable for kidney damage in lyn(-/-) mice. European Journal of Immunology, 43(2), 382–393.
Hu, X., et al. (2009). Genetic alterations and oncogenic pathways associated with breast cancer subtypes. Molecular Cancer Research, 7(4), 511–522.
Li, T., Copeland, C., & Le, A. (2021). Glutamine metabolism in cancer. Advances in Experimental Medicine and Biology, 1311, https://doi.org/10.1007/978-3-030-65768-0_2.
Elgogary, A., et al. (2016). Combination therapy with BPTES nanoparticles and metformin targets the metabolic heterogeneity of pancreatic cancer. Proceedings of the National Academy of Sciences of the United States of America, 113(36), E5328–E5336.
Martinez-Outschoorn, U. E., et al. (2012). BRCA1 mutations drive oxidative stress and glycolysis in the tumor microenvironment: Implications for breast cancer prevention with antioxidant therapies. Cell Cycle, 11(23), 4402–4413.
Sazeides, C., & Le, A. (2021). Metabolic relationship between cancer-associated fibroblasts and cancer cells. Advances in Experimental Medicine and Biology, 1311, https://doi.org/10.1007/978-3-030-65768-0_14.
Jung, J. G., & Le, A. (2021). Targeting metabolic cross talk between cancer cells and cancer-associated fibroblasts. Advances in Experimental Medicine and Biology, 1311, https://doi.org/10.1007/978-3-030-65768-0_15.
Warburg, O., Wind, F., & Negelein, E. (1927). The metabolism of tumors in the body. The Journal of General Physiology, 8(6), 519–530.
Vander Heiden, M. G., Cantley, L. C., & Thompson, C. B. (2009). Understanding the Warburg effect: The metabolic requirements of cell proliferation. Science, 324(5930), 1029–1033.
Bose, S., Zhang, C., & Le, A. (2021). Glucose metabolism in cancer: The Warburg effect and beyond. Advances in Experimental Medicine and Biology, 1311, https://doi.org/10.1007/978-3-030-65768-0_1.
Le, A., et al. (2010). Inhibition of lactate dehydrogenase A induces oxidative stress and inhibits tumor progression. Proceedings of the National Academy of Sciences of the United States of America, 107(5), 2037–2042.
Rajeshkumar, N. V., et al. (2015). Therapeutic targeting of the Warburg effect in pancreatic cancer relies on an absence of p53 function. Cancer Research, 75(16), 3355–3364.
Dutta, P., et al. (2013). Evaluation of LDH-A and glutaminase inhibition in vivo by hyperpolarized 13C-pyruvate magnetic resonance spectroscopy of tumors. Cancer Research, 73(14), 4190–4195.
Gatenby, R. A., & Gillies, R. J. (2004). Why do cancers have high aerobic glycolysis? Nature Reviews. Cancer, 4(11), 891–899.
Waki, A., et al. (1998). The importance of glucose transport activity as the rate-limiting step of 2-deoxyglucose uptake in tumor cells in vitro. Nuclear Medicine and Biology, 25(7), 593–597.
Choi, J., Jung, W. H., & Koo, J. S. (2013). Metabolism-related proteins are differentially expressed according to the molecular subtype of invasive breast cancer defined by surrogate immunohistochemistry. Pathobiology, 80(1), 41–52.
Grover-McKay, M., et al. (1998). Role for glucose transporter 1 protein in human breast cancer. Pathology Oncology Research, 4(2), 115–120.
Lloyd, S. M., Arnold, J., & Sreekumar, A. (2015). Metabolomic profiling of hormone-dependent cancers: A bird’s eye view. Trends in Endocrinology and Metabolism, 26(9), 477–485.
Katz-Brull, R., et al. (2002). Metabolic markers of breast cancer: Enhanced choline metabolism and reduced choline-ether-phospholipid synthesis. Cancer Research, 62(7), 1966–1970.
Eliyahu, G., Kreizman, T., & Degani, H. (2007). Phosphocholine as a biomarker of breast cancer: Molecular and biochemical studies. International Journal of Cancer, 120(8), 1721–1730.
Aboagye, E. O., & Bhujwalla, Z. M. (1999). Malignant transformation alters membrane choline phospholipid metabolism of human mammary epithelial cells. Cancer Research, 59(1), 80–84.
Park, J. K., et al. (2021). The heterogeneity of lipid metabolism in cancer. Advances in Experimental Medicine and Biology, 1311, https://doi.org/10.1007/978-3-030-65768-0_3.
Hilvo, M., et al. (2011). Novel theranostic opportunities offered by characterization of altered membrane lipid metabolism in breast cancer progression. Cancer Research, 71(9), 3236–3245.
Ramirez de Molina, A., et al. (2002). Increased choline kinase activity in human breast carcinomas: Clinical evidence for a potential novel antitumor strategy. Oncogene, 21(27), 4317–4322.
Rodriguez-Gonzalez, A., et al. (2004). Choline kinase inhibition induces the increase in ceramides resulting in a highly specific and selective cytotoxic antitumoral strategy as a potential mechanism of action. Oncogene, 23(50), 8247–8259.
Fuhrman, B. J., et al. (2012). Estrogen metabolism and risk of breast cancer in postmenopausal women. Journal of the National Cancer Institute, 104(4), 326–339.
Cicatiello, L., et al. (2010). Estrogen receptor alpha controls a gene network in luminal-like breast cancer cells comprising multiple transcription factors and microRNAs. The American Journal of Pathology, 176(5), 2113–2130.
Acconcia, F., & Kumar, R. (2006). Signaling regulation of genomic and nongenomic functions of estrogen receptors. Cancer Letters, 238(1), 1–14.
Jia, M., et al. (2016). Estrogen receptor alpha promotes breast cancer by reprogramming choline metabolism. Cancer Research, 76(19), 5634–5646.
Devanesan, P., et al. (2001). Catechol estrogen conjugates and DNA adducts in the kidney of male Syrian golden hamsters treated with 4-hydroxyestradiol: Potential biomarkers for estrogen-initiated cancer. Carcinogenesis, 22(3), 489–497.
Cavalieri, E., et al. (2000). Estrogens as endogenous genotoxic agents—DNA adducts and mutations. Journal of the National Cancer Institute. Monographs, 27, 75–93.
Bradlow, H. L., et al. (1996). 2-Hydroxyestrone: The ‘good’ estrogen. The Journal of Endocrinology, 150(Suppl), S259–S265.
Possemato, R., et al. (2011). Functional genomics reveal that the serine synthesis pathway is essential in breast cancer. Nature, 476(7360), 346–350.
Fan, J., et al. (2015). Human phosphoglycerate dehydrogenase produces the oncometabolite D-2-hydroxyglutarate. ACS Chemical Biology, 10(2), 510–516.
Rakheja, D., et al. (2013). The emerging role of d-2-hydroxyglutarate as an oncometabolite in hematolymphoid and central nervous system neoplasms. Frontiers in Oncology, 3, 169.
Terunuma, A., et al. (2014). MYC-driven accumulation of 2-hydroxyglutarate is associated with breast cancer prognosis. The Journal of Clinical Investigation, 124(1), 398–412.
Yue, W., et al. (2003). Genotoxic metabolites of estradiol in breast: Potential mechanism of estradiol induced carcinogenesis. The Journal of Steroid Biochemistry and Molecular Biology, 86(3–5), 477–486.
Quinones, A., & Le, A. (2021). The multifaceted glioblastoma: From genomic alterations to metabolic adaptations. Advances in Experimental Medicine and Biology, 1311, https://doi.org/10.1007/978-3-030-65768-0_4.
Mullarky, E., et al. (2016). Identification of a small molecule inhibitor of 3-phosphoglycerate dehydrogenase to target serine biosynthesis in cancers. Proceedings of the National Academy of Sciences of the United States of America, 113(7), 1778–1783.
Jerby, L., et al. (2012). Metabolic associations of reduced proliferation and oxidative stress in advanced breast cancer. Cancer Research, 72(22), 5712–5720.
Dang, C. V., Le, A., & Gao, P. (2009). MYC-induced cancer cell energy metabolism and therapeutic opportunities. Clinical Cancer Research, 15(21), 6479–6483.
Le, A., & Dang, C. V. (2013). Studying Myc’s role in metabolism regulation. Methods in Molecular Biology, 1012, 213–219.
Shen, L., et al. (2015). Metabolic reprogramming in triple-negative breast cancer through Myc suppression of TXNIP. Proceedings of the National Academy of Sciences of the United States of America, 112(17), 5425–5430.
Hoang, G., Udupa, S., & Le, A. (2019). Application of metabolomics technologies toward cancer prognosis and therapy. International Review of Cell and Molecular Biology, 347, 191–223.
Dang, C. V., et al. (2011). Therapeutic targeting of cancer cell metabolism. Journal of Molecular Medicine (Berlin), 89(3), 205–212.
Hirschey, M. D., et al. (2015). Dysregulated metabolism contributes to oncogenesis. Seminars in Cancer Biology, 35(Suppl), S129–S150.
Jove, M., et al. (2017). A plasma metabolomic signature discloses human breast cancer. Oncotarget, 8(12), 19522–19533.
Huang, S., et al. (2016). Novel personalized pathway-based metabolomics models reveal key metabolic pathways for breast cancer diagnosis. Genome Medicine, 8(1), 34.
Giskeodegard, G. F., et al. (2010). Multivariate modeling and prediction of breast cancer prognostic factors using MR metabolomics. Journal of Proteome Research, 9(2), 972–979.
Mountford, C. E., et al. (2001). Diagnosis and prognosis of breast cancer by magnetic resonance spectroscopy of fine-needle aspirates analysed using a statistical classification strategy. The British Journal of Surgery, 88(9), 1234–1240.
Asiago, V. M., et al. (2010). Early detection of recurrent breast cancer using metabolite profiling. Cancer Research, 70(21), 8309–8318.
Oakman, C., et al. (2011). Identification of a serum-detectable metabolomic fingerprint potentially correlated with the presence of micrometastatic disease in early breast cancer patients at varying risks of disease relapse by traditional prognostic methods. Annals of Oncology, 22(6), 1295–1301.
Jobard, E., et al. (2014). A serum nuclear magnetic resonance-based metabolomic signature of advanced metastatic human breast cancer. Cancer Letters, 343(1), 33–41.
Antonio, M. J., Zhang, C., & Le, A. (2021). Different tumor microenvironments lead to different metabolic phenotypes. Advances in Experimental Medicine and Biology, 1311, https://doi.org/10.1007/978-3-030-65768-0_10.
Simoes, R. V., et al. (2015). Metabolic plasticity of metastatic breast cancer cells: Adaptation to changes in the microenvironment. Neoplasia, 17(8), 671–684.
Wei, S., et al. (2013). Metabolomics approach for predicting response to neoadjuvant chemotherapy for breast cancer. Molecular Oncology, 7(3), 297–307.
Nabi, K., & Le, A. (2021). The intratumoral heterogeneity of cancer metabolism. Advances in Experimental Medicine and Biology, 1311, https://doi.org/10.1007/978-3-030-65768-0_11.
Xu, H. N., et al. (2013). Characterizing the metabolic heterogeneity in human breast cancer xenografts by 3D high resolution fluorescence imaging. Springerplus, 2(1), 73.
Son, S. H., et al. (2014). Prognostic implication of intratumoral metabolic heterogeneity in invasive ductal carcinoma of the breast. BMC Cancer, 14, 585.
Park, V. Y., et al. (2016). Intratumoral agreement of high-resolution magic angle spinning magnetic resonance spectroscopic profiles in the metabolic characterization of breast cancer. Medicine (Baltimore), 95(15), e3398.
Gallagher, F. A., et al. (2020). Imaging breast cancer using hyperpolarized carbon-13 MRI. Proceedings of the National Academy of Sciences of the United States of America, 117(4), 2092–2098.
Marusyk, A., Janiszewska, M., & Polyak, K. (2020). Intratumor heterogeneity: The Rosetta Stone of therapy resistance. Cancer Cell, 37(4), 471–484.
Schafer, Z. T., et al. (2009). Antioxidant and oncogene rescue of metabolic defects caused by loss of matrix attachment. Nature, 461(7260), 109–113.
Lee, Y. J., et al. (1998). Glucose deprivation-induced cytotoxicity and alterations in mitogen-activated protein kinase activation are mediated by oxidative stress in multidrug-resistant human breast carcinoma cells. The Journal of Biological Chemistry, 273(9), 5294–5299.
Ahn, C. S., & Metallo, C. M. (2015). Mitochondria as biosynthetic factories for cancer proliferation. Cancer & Metabolism, 3(1), 1.
LeBleu, V. S., et al. (2014). PGC-1alpha mediates mitochondrial biogenesis and oxidative phosphorylation in cancer cells to promote metastasis. Nature Cell Biology, 16(10), 992–1003. 1–15.
Robertson-Tessi, M., et al. (2015). Impact of metabolic heterogeneity on tumor growth, invasion, and treatment outcomes. Cancer Research, 75(8), 1567–1579.
Kerbel, R. S. (2009). Issues regarding improving the impact of antiangiogenic drugs for the treatment of breast cancer. Breast, 18(Suppl 3), S41–S47.
Ma, S., et al. (2018). The role of tumor microenvironment in resistance to anti-angiogenic therapy. F1000Res, 7, 326.
Conley, S. J., et al. (2012). Antiangiogenic agents increase breast cancer stem cells via the generation of tumor hypoxia. Proceedings of the National Academy of Sciences of the United States of America, 109(8), 2784–2789.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2021 The Author(s)
About this chapter
Cite this chapter
Tan, J., Le, A. (2021). The Heterogeneity of Breast Cancer Metabolism. In: Le, A. (eds) The Heterogeneity of Cancer Metabolism. Advances in Experimental Medicine and Biology, vol 1311. Springer, Cham. https://doi.org/10.1007/978-3-030-65768-0_6
Download citation
DOI: https://doi.org/10.1007/978-3-030-65768-0_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-65767-3
Online ISBN: 978-3-030-65768-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)