Abstract
Cancer stem cells (CSCs), also known as tumorinitiating cells (TICs), are a group of cells found within cancer cells. Like normal stem cells, CSCs can proliferate, engage in self-renewal, and are often implicated in the recurrence of tumors after therapy [1, 2]. The existence of CSCs in various types of cancer has been proven, such as in acute myeloid leukemia (AML) [3], breast [4], pancreatic [5], and lung cancers [6], to name a few. There are two theories regarding the origin of CSCs. First, CSCs may have arisen from normal stem/progenitor cells that experienced changes in their environment or genetic mutations. On the other hand, CSCs may also have originated from differentiated cells that underwent genetic and/or heterotypic modifications [7]. Either way, CSCs reprogram their metabolism in order to support tumorigenesis.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
- Cancer stem cell
- Metabolic plasticity
- Glucose metabolism
- Glutamine metabolism
- Mitochondrial metabolism
- Lipid metabolism
-
CSCs are different from stem cells and non-CSC tumor cells metabolically, but CSCs share some similarities with both stem cells and cancer cells in glucose, glutamine, oxidative phosphorylation (OXPHOS), and lipid metabolism.
-
The commonly shared metabolic pathways indicate that CSCs are a subtype of cancer cells that genetically evolved from normal stem/progenitor cells.
-
The commonly shared metabolic pathways between CSCs, stem cells, and non-CSC tumor cells indicate that these metabolic pathways are selected due to their ability to sustain the catabolic and anabolic needs of highly proliferating cells.
-
The fact that CSCs share metabolic pathways with stem cells and cancer cells poses a challenge for developing cancer metabolic drugs that do not target normal stem cells.
-
Further studies are needed to shed light on how to develop specific metabolic drugs targeting CSCs in specific cancer types.
1 Introduction
Cancer stem cells (CSCs), also known as tumor-initiating cells (TICs), are a group of cells found within cancer cells. Like normal stem cells, CSCs can proliferate, engage in self-renewal, and are often implicated in the recurrence of tumors after therapy [1, 2]. The existence of CSCs in various types of cancer has been proven, such as in acute myeloid leukemia (AML) [3], breast [4], pancreatic [5], and lung cancers [6], to name a few. There are two theories regarding the origin of CSCs. First, CSCs may have arisen from normal stem/progenitor cells that experienced changes in their environment or genetic mutations. On the other hand, CSCs may also have originated from differentiated cells that underwent genetic and/or heterotypic modifications [7]. Either way, CSCs reprogram their metabolism in order to support tumorigenesis.
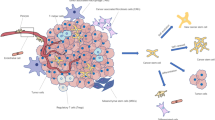
Metabolism plays a crucial role in cellular function and survival, and it is no different for CSCs. Metabolic rewiring is necessary for CSCs as it enables them to adapt to different environments and survive. For example, metabolic rewiring to oxidative phosphorylation (OXPHOS) from glycolysis makes CSCs more efficient in generating adenosine triphosphate (ATP) and more resistant to microenvironmental pressures such as lack of nutrients [8]. Understanding the metabolism of CSCs would enhance targeting CSCs, and in turn improve cancer therapy. In this chapter, we will be looking at four key metabolism of CSCs—glucose metabolism, glutamine metabolism, mitochondrial metabolism, and lipid metabolism—and how they affect the state of CSCs (Fig. 1).
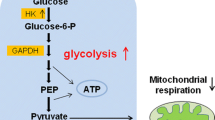
2 High Levels of Glycolytic Enzymes and Activities in CSCs (Fig. 2)
In Chap. 1, glucose metabolism and the Warburg effect have been described as key processes in cancer cells [9]. Not only are these processes observed in cancer cells, but they are also relevant to CSCs. Stemness features of CSCs, such as proliferation, are shown to be reliant on the Warburg effect [10].
CSCs across different types of cancers are found to express high levels of glycolytic genes. Song et al. found that CD133(+) liver CSCs exhibit increased expressions of glycolytic genes: glucose transporter 1 (GLUT1), hexokinase II (HK2), pyruvate dehydrogenase kinase 4 (PDK4), and phosphoglycerate mutase 1 (PGAM1). Furthermore, they observed a high extracellular acidification rate (ECAR) and decreased expressions of gluconeogenetic genes: glucose-6-phosphatase (G6Pase) and phosphoenolpyruvate carboxykinase (Pepck) [11]. These findings align with observations made in nasopharyngeal carcinoma (NPC) CSCs. In comparison to their parental cells, NPC CSCs express higher levels of GLUT1, HK2, and glucose-6-phosphate isomerase (GPI) [12]. The high levels of glycolytic genes found in CSCs suggest that CSCs depend on glucose metabolism as a source of energy.
Glucose and glutamine metabolism in cancer stem cells. GLUT1 glucose transporter 1, HK2 hexokinase II, LDH-A lactate dehydrogenase A, PDHC pyruvate dehydrogenase complex, PDK4 pyruvate dehydrogenase kinase 4, GOT glutamic-oxaloacetic transaminase, GLS1 glutaminase 1, ASCT2 alanine, serine, cysteine transporter 2
Studies have demonstrated the importance of glycolytic enzymes in CSCs, such as regulating the self-renewal and spheroid formation ability of CSCs. Shibuya et al. studied the role of GLUT1 in the CSCs of glioblastoma, pancreatic, and ovarian cancers. They found that the inhibition of GLUT1, through either siRNA-mediated knockdown or pharmacological inhibition by WZB117, a small-molecule inhibitor, resulted in a decrease in the uptake of glucose, a reduction in the expressions of stem cell markers (Sox2, Bmi1, Nanog), and an increase in the expressions of differentiation markers (GFAP in glioblastoma and B-actin in glioblastoma, pancreatic, and ovarian CSCs). Furthermore, these inhibitions impaired the spheroid-formation ability of CSCs, and inhibition by WZB117 did so without affecting their proliferative potential [13]. These findings suggest that GLUT1 plays a crucial role in maintaining the glucose metabolism and stemness characteristics of CSCs.
PDK4, one of the highly expressed genes in liver CSCs mentioned earlier, is an inhibitor of pyruvate dehydrogenase complex (PDHC). PDHC regulates the entrance of glucose-derived pyruvate into the mitochondria, which will subsequently form acetyl-CoA for the tricarboxylic acid (TCA) cycle. Song et al. found that low levels of liver-specific miR-122, which targets PDK4, are expressed in CD133+ cells. This allows for the upregulation of PDK4 and the subsequent inhibition of PDH, which in turn results in the failure of pyruvate to enter the mitochondrial matrix. Instead of going through subsequent oxidative phosphorylation (OXPHOS), lactate production in the cytosol occurs [11]. Therefore, this supports the idea that the Warburg effect is of relevance to CSCs. In fact, high levels of miR-122 impair the spheroid-formation ability of CSCs [11]. Hence, suppression of miR-122 in liver CSCs allows the induction of glucose metabolism and preference for fermentation instead of OXPHOS.
The conversion of pyruvate to lactate in the cytosol is regulated by lactate dehydrogenase A (LDH-A), which was found to be upregulated in breast CSCs compared to spheroid-derived adherent cells (SDACs) [14]. The importance of LDH-A can be concluded from the deleterious effect of LDH-A suppression on CSCs. The knockdown of LDH-A caused liver CD133+ CSCs to produce fewer transcriptional factors (Sox2, Nanog, Oct4) that give rise to stemness characteristics, and their spheroid-formation ability was impaired [11]. Furthermore, in non-small cell lung carcinoma (NSCLC), the suppression of LDH-A by shRNA resulted in a decrease of CSCs in the A549 cell line and a decrease in their ability to form tumorspheres [15]. These findings reinforce the idea that glucose metabolism, which results in lactate production, is central to the survival of CSCs and the maintenance of their stemness characteristics. LDH-A inhibitor such as FX-11 has been tested in several types of cancers [9, 16,17,18] and could be a potential strategy in targeting CSCs.
3 Effects of Deregulation of Glutamine Metabolism on CSCs (Fig. 2)
Glutamine metabolism is also a source of energy for cancer cells that is an alternative to glucose metabolism [19, 20]. For CSCs, a lack of glucose-derived pyruvate for the TCA cycle would mean that reliance on other metabolites to drive the TCA cycle is needed. Compared to non-CSCs, metabolites from oxidative glutamine metabolism contribute to the TCA cycle in CSCs [10]. Additionally, both glucose deprivation and treatment using the glycolytic inhibitor 2-deoxy-glucose (2-DG) resulted in an increase in the uptake of glutamine into CSCs but not into non-CSCs, further suggesting the metabolic plasticity of CSCs [10]. Colorectal CSCs of the SW620 cell line were shown to be resistant to the effects of inhibition of mitochondrial metabolism by metformin because they compensated by deriving energy from glutamine metabolism [21]. These findings suggest the importance of glutamine metabolism in CSC survival. Targeting of glutamine metabolism has been extensively studied by many researchers [22,23,24,25,26,27,28,29]. These strategies could have the potential of eliminating CSCs.
Glutaminolysis starts with the intake of glutamine into the cell via transporters such as alanine, serine, and cysteine transporter 2 (ASCT2). Kim et al. found that the CSCs of colorectal cancer, both HT29 and SW260 cell lines, have a higher amount of ASCT2 than non-CSCs, which means that a higher glutamine uptake occurs in CSCs. After ASCT2 knockdown, the number of CD133+/CD44+ CSCs drastically decreased. In metformin-resistant cell line SW260, the siRNA-mediated knockdown of glutaminase 1 (GLS1) and ASCT2 decreased the number of CSCs, with a more significant decrease observed during ASCT2 knockdown only [21].
Another study on pancreatic CSCs has demonstrated the importance of glutamine metabolism to CSCs. When there is a lack of glutamine, pancreatic CSCs have a decreased ability to form both primary and secondary spheroids. Furthermore, the lack of glutamine also resulted in reduced expression of stemness-related genes and accumulation of intracellular reactive oxygen species (ROS). Oxaloacetate (OAA), an intermediate of the TCA cycle, was found to alleviate the high ROS levels. Furthermore, the knockdown of cytosolic glutamic-oxaloacetic transaminase 1 (GOT1), which converts aspartate to OAA, together with the administration of low-dose ionizing radiation (IR) resulted in impairment of spheroid-formation ability, and the spheroids formed experienced apoptosis within 10 days of their formation [30]. These findings revealed the important role that OAA plays in ensuring a balanced ROS level in CSCs. Therefore, the noncanonical pathway of glutamine metabolism helps CSCs in their ability to resist ROS accumulation and its effects.
4 Mitochondrial Metabolism (Fig. 3)
The term “mitostemness” describes the crucial role that mitochondria play in the ability of CSCs to self-renew and resist differentiating and losing their stemness properties [31]. The ability of CSCs to switch across different types of metabolism for better survival in different environmental conditions has resulted in contradictory findings regarding their metabolism. Some studies have shown that CSCs depend on glycolysis [32, 33], while others have described CSCs to be mostly dependent on OXPHOS [33,34,35].
In addition to their energetics contributions, mitochondrial metabolic pathways also epigenetically regulate CSC stemness. Mitochondrial metabolism is important because the TCA cycle and OXPHOS are involved in the production of metabolites such as acetyl-CoA, alpha-ketoglutarate, NAD+, and S-adenosyl methionine, which are substrates for chromatin-state modifications, whether acetylation or methylation [36]. In turn, these epigenetic modifications have been found to alter the stemness of CSCs [37].
4.1 OXPHOS
OXPHOS, which involves the mitochondrial respiratory chain, is the last step of aerobic glucose metabolism where energy is generated as ATP, which is used for cell survival and growth. In a study done on pancreatic CSCs, it was found that in comparison to normal adherent cells, CSCs have a higher number of mitochondria and form more components that are required for OXPHOS. Furthermore, compared to non-CSCs (CD133− differentiated cells), CD133+ CSCs have a higher mitochondrial oxygen consumption rate (OCR) [35], suggesting the reliance of CSCs on OXPHOS for their growth and survival. In fact, the inhibition of OXPHOS resulted in CSCs experiencing an energy crisis. However, there is a subset of CSCs that are resistant to inhibition of mitochondrial metabolism by metformin, and they are found to have enhanced glycolytic capacity [35], suggesting the metabolic plasticity of CSCs.
Studies have shown that the inhibition of OXPHOS can have detrimental effects on CSCs. Salinomycin, a K+ ionophore, which changes the mitochondrial membrane potential of cells, inhibits OXPHOS [38] and decreases the number of CSCs in breast [39], gastric [40], and pancreatic cancers [41]. The change in mitochondrial membrane potential caused by salinomycin causes mitochondrial imbalance. In turn, ATP depletion activates AMP-activated protein kinase (AMPK) and results in apoptosis of CSCs [42]. A balanced and well-regulated mitochondrion is, therefore, important to CSCs.
The mitochondrial respiratory chain is comprised of four enzymatic complexes for the transfer of electrons to O2 and generate ATP, which is used for cell survival. Only complex II is completely encoded by the nuclear genome, while for complexes I, III, and IV, mitochondrial DNA (mtDNA) plays a role in encoding them [43]. The inhibition of these complexes is detrimental to the survival and growth of CSCs and, therefore, they serve as possible targets for eliminating CSCs. The inhibition of complex I in pancreatic cancer by metformin resulted in a cell cycle arrest in non-CSCs, while in CSCs, it led to apoptosis [34]. This observation aligns with the finding that metformin results in the selective killing of CD44+/CD24− CSCs of MCF10-A and MCF-7 cell lines, and works well with doxorubicin in reducing both CSCs and non-CSCs [44]. Another study showed that the inhibition of complexes I and II by pyrvinium pamoate caused dysregulated mitochondrial metabolism in the fat pad (FP) CSCs, but this specific mechanism was not toxic to CSCs [45]. Treatment using atovaquone, a complex III inhibitor, decreased proliferation of CD44+/CD24− MCF7 breast CSCs, and induced apoptosis in dose-independent conditions, but had no effect on mitochondrial respiration in normal fibroblasts [46]. Furthermore, complex V inhibitor bedaquiline, which causes ATP depletion, was found to target MCF7 breast CSCs preferentially and did not affect the viability of cancer cells and normal human fibroblasts [47]. These studies indicated the importance of OXPHOS across different CSCs as the inhibition of the complexes involved in OXPHOS negatively impacted the survival and growth of CSCs. However, it is important to keep in mind the different responses to mitochondrial respiratory chain complexes’ inhibition seen in different CSCs. For instance, complex I inhibition by metformin in pancreatic CSCs led to their apoptosis, but no cytotoxic effect was brought about in FP-CSCs when pyrvinium pamoate inhibited complexes I and II. Further studies regarding the differences in the mechanism and regulation of OXPHOS across different CSCs are required. Nevertheless, it is certain that OXPHOS is necessary and important to the survival of CSCs and its enzymatic complexes serve as potential targets in the elimination of CSCs.
4.2 Resistance of CSCs Against ROS
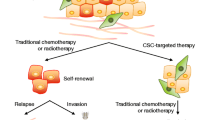
ROS are reactive products generated from mitochondrial metabolism. Mitochondria are the major source of intracellular ROS, such as free radicals, superoxide, and peroxides [42, 48]. Excess ROS causes apoptosis via oxidative stress [49]. Radiotherapy and most of the chemotherapies increase ROS levels and thereby affect cancer cell viability and growth [50, 51]. Despite such therapy, tumor recurrence does occur due to the failure to eliminate CSCs during such treatment [52, 53]. One of the mechanisms of CSC resistance is through increased Bcl-2 expression and radioresistance.
Studies have shown the resistance of CSCs against the effects of ROS in CD44+ CSCs of prostate, breast, and colorectal cancers [54,55,56]. An isoform of CD44, CD44v, was found to protect CSCs of gastric cancer from the detrimental effects of ROS accumulation. CD44v stabilizes the xCT subunit of cystine-glutamate exchange transporter, allowing for the uptake of cystine needed for glutathione (GSH) synthesis. The upregulation of GSH synthesis makes CSCs more antioxidative [57]. Other than containing CD44, CSCs increase the expression of antioxidant enzyme genes that neutralize ROS and the multifunctional protein apurinic/apyrimidinic endonuclease/redox effector factor (Ape1/Ref-1). In turn, they neutralize the intracellular ROS and reduce DNA damage via DNA repair [58]. Therefore, through their antioxidative capability, CSCs are resistant to the effects of ROS produced during therapy and can, therefore, cause recurrence of tumors [58, 59]. The defense mechanism against ROS of CSCs needs to be targeted for improved cancer treatment.
5 Lipid Metabolism (Fig. 4)
Lipid anabolic metabolism consists of both ex novo incorporation of lipids and de novo syntheses of lipids [60, 61]. De novo lipogenesis consists of three major pathways: fatty acid synthesis (FAS), mevalonate biosynthesis, and cholesterol biosynthesis [62]. CSCs exhibit both ex novo and de novo lipid anabolic pathways. Interestingly, despite the activation of anabolic pathways, CSCs also activate catabolic lipid pathways, such as fatty acid oxidation (FAO). Classically, FAS and FAO are antagonistic pathways due to the role of FAS intermediate malonyl CoA, which suppresses carnitine palmitoyltransferase 1 (CPT1), inactivating FAO in normal cellular metabolism [63]. However, it seems that in CSCs, both pathways are activated to allow them to adapt best to their environmental pressures and cellular demands for proliferation.
Lipogenesis via uptake from the environment has been reported in CSCs [64]. Given the complexity of certain lipids such as phospholipids and cholesterol, which rely on the stoichiometric balancing of fatty acid with glycerol in various combinations, it makes sense that CSCs have been found to incorporate these complex lipids. Mass spectrometry studies reveal that CSCs have abundant unsaturated lipids, cholesterol, and triglycerides (TGs) which are important for their stemness [61]. The combination of external lipid uptake and de novo lipogenesis seems to be the most efficient route for fulfilling the increasing demands of membrane production for the proliferating CSCs.
De novo lipogenesis, however, still plays a major role despite CSC’s ex novo sources of lipids. The FAS pathway, being the initial step of de novo lipogenesis, is activated in CSCs, as it is in some cancer cells, such as those driven by MYC [62, 65, 66]. The key enzymes in the FAS pathway are ATP citrate lyase (ACLY), acetyl carboxylase A (ACC1), fatty acid synthase (FASN), and stearoyl-CoA desaturase (SCD). ACLY is connected directly to the TCA cycle via citrate synthase (CS). Thus, increased TCA mitochondrial metabolism in CSCs would also drive the flux into FAS via CS into ACLY and the rest of the FAS pathway [62]. Though de novo lipogenesis may appear to be a secondary or passenger effect of increased glucose and glutamine metabolism in CSCs, inhibition of virtually any enzyme of the FAS pathway in CSCs leads to apoptosis. ACLY knockdown reduced the proliferation of lung cancer [67] and breast cancer CSCs [68]. ACC1 inhibition suppressed the stemness of CSCs [69]. Inhibition of FASN reduced the stemness of glioma stem cells [70]. SCD inhibition has been correlated with the suppression of various CSCs [71].
Like the FAS pathway, the cholesterol synthesis pathways via mevalonate are regulated by sterol regulatory element-binding proteins (SREBPs). SREBP2 actively transcribes HMG-CoA synthase (HMGCS), HMG-CoA reductase (HMGCR), mevalonate kinase (MVK), and phosphomevalonate kinase (PMVK) for the production of pyrophosphate-containing intermediates, which leads to the production of cholesterol. The mevalonate and cholesterol synthesis pathways are connected to the acetyl-CoA pool from ACLY in FAS via acetoacetyl CoA transferase (ACAT), which feeds into HMGCS. During epithelial-mesenchymal transition (EMT), the mevalonate and cholesterol pathways are activated, such that inhibition of any of the aforementioned enzymes disrupts CSCs’ stemness and the overall EMT [72].
In addition to the induction of the anabolic lipid metabolic pathways mentioned above (ex novo incorporation of lipids and the three de novo lipogenesis pathways: FAS, mevalonate, and cholesterol), CSCs also upregulate the catabolic lipid metabolic pathway: FAO [73]. FAO takes place in mitochondria following the uptake of fatty acids via CPT1. CPT1 can be inhibited by malonyl CoA, which is a product of ACC1 of the FAS pathway, ensuring the inhibition of running a futile cycle between FAS and FAO. However, due to the highly proliferative state of FAO and the ex vivo uptake of lipids, FAO is also upregulated in CSCs. This FAO pathway may be necessary for CSCs to replenish NADH and FADH2, which will help facilitate the electron transport chain (ETC) in OXPHOS [63]. Indeed, inhibition of FAO has been reported to lower the number of CSCs [74].
6 Conclusion
From our brief overview, it is clear that CSCs have diverse metabolic profiles that are quite specific to each cancer type and tissue type, which is a break from convention with normal cell metabolism. For example, the activation of OXPHOS alongside the Warburg effect presents a complexity that demands further studies. The shunting of glucose-derived pyruvate to lactate instead of the TCA cycle by the Warburg effect should preclude the flow of glucose carbons to OXPHOS unless somehow the glutaminolysis carbons can compensate for all the TCA cycle to push forward to OXPHOS independent of glucose [20]. Regardless, CSCs seem to exhibit both the Warburg effect and OXPHOS simultaneously. Another example is the activation of mitochondrial metabolism alongside lipogenesis, which also challenges conventional wisdom concerning cellular metabolism. Typically, the activation of lipogenesis would block fatty acid oxidation in the mitochondria due to the inhibition of CPT1 by malonyl-CoA. However, CSCs seem to have no problem activating both pathways.
The metabolic assessment of CSCs using metabolomics technologies [75] reflects variation within the CSC population. Single-cell metabolic analyses would enable us to identify such diversity in the CSC population. Moreover, there is tissue and microenvironmental specificity to CSC metabolic profiles. For example, breast CSCs that are surrounded by lipids may “prefer” to activate fatty acid oxidation instead of inducing de novo lipogenesis. This may not be the case for lymphomas that travel within the blood vessels, which may prefer to use glucose as a primary energy source instead of lipids. Furthermore, CSCs seem to run futile cycles metabolically (i.e., activating lipogenesis and fatty acid oxidation and OXPHOS simultaneously), because these processes are not absolutely efficient. Thus, CSCs activate both pathways to make sure that all residual metabolites are processed efficiently catabolically and anabolically to support their high demand for energy and biomass.
Despite the differences in CSC metabolic profiles, CSCs do share some similarities with both stem cells and cancer cells in glucose, glutamine, OXPHOS, and lipid metabolism. This makes it difficult to distinguish CSCs from stem cells and non-CSC tumor cells metabolically, and conventional CSC markers are still needed (i.e., CD44 and CD133). However, the fact that there are commonly activated metabolic pathways between CSCs, stem cells, and non-CSC tumor cells is consistent with three ideas pertaining to the nature of CSCs. First, the commonly shared metabolic pathways indicate that CSCs are indeed a subtype of cancer cells that genetically evolved from normal stem/progenitor cells. Second, the commonly shared metabolic pathways between the three indicate that these metabolic pathways are selected due to their ability to sustain the catabolic and anabolic needs of highly proliferating cells. Third, the fact that CSCs share metabolic pathways with stem cells and cancer cells presents a challenge for developing cancer metabolic drugs that do not target normal stem cells.
Overall, this is a fascinating field in metabolism, where researchers need to further delineate metabolic similarities and differences within the CSC population itself as well as with other stem cell and cancer cell types. These intra- and inter-differences will shed light on how to best develop specific metabolic drugs targeting CSCs in specific cancer types that would not be toxic to normal stem cells.
Change history
22 October 2021
After initial publication of the book, various errors were identified that needed correction. All corrections listed below have been updated within the current version.
Abbreviations
- 2-DG:
-
2-Deoxy-d-glucose
- ACC1:
-
Acetyl carboxylase A
- ACLY:
-
ATP citrate lyase
- AML:
-
Acute myeloid leukemia
- AMPK:
-
AMP-activated protein kinase
- Ape1/Ref-1:
-
Apurinic/apyrimidinic endonuclease/redox effector factor
- ASCT2:
-
Alanine, serine, cysteine transporter 2
- ATP:
-
Adenosine triphosphate
- CPT1:
-
Carnitine palmitoyltransferase 1
- CS:
-
Citrate synthase
- CSCs:
-
Cancer stem cells
- ECAR:
-
Extracellular acidification rate
- EMT:
-
Epithelial-mesenchymal transition
- FAO:
-
Fatty acid oxidation
- FAS:
-
Fatty acid synthesis
- FASN:
-
Fatty acid synthase
- FP:
-
Fat pad
- G6Pase:
-
Glucose-6-phosphatase
- GLS1:
-
Glutaminase 1
- GLUT1:
-
Glucose transporter 1
- GOT1:
-
Glutamic-oxaloacetic transaminase 1
- GPI:
-
Glucose-6-phosphate isomerase
- HK2:
-
Hexokinase II
- HMGCR:
-
HMG-CoA reductase
- HMGCS:
-
HMG-CoA synthase
- IR:
-
Ionizing radiation
- LDH-A:
-
Lactate dehydrogenase A
- mtDNA:
-
Mitochondrial deoxyribonucleic acid
- MVK:
-
Mevalonate kinase
- NSCLC:
-
Non-small cell lung carcinoma
- OAA:
-
Oxaloacetate
- OCR:
-
Oxygen consumption rate
- OXPHOS:
-
Oxidative phosphorylation
- PDHC:
-
Pyruvate dehydrogenase complex
- PDK4:
-
Pyruvate dehydrogenase kinase 4
- Pepck:
-
Phosphoenolpyruvate carboxykinase
- PGAM1:
-
Phosphoglycerate mutase 1
- PMVK:
-
Phosphomevalonate kinase
- PPPs:
-
Pyrophosphates
- ROS:
-
Reactive oxygen species
- SCD:
-
Stearoyl CoA desaturase
- SREBPs:
-
Sterol regulatory element-binding proteins
- TCA:
-
Tricarboxylic acid
- TGs:
-
Triglycerides
- TICs:
-
Tumor-initiating cells
References
Dick, J. E. (2008). Stem cell concepts renew cancer research. Blood, 112(13), 4793–4807.
Reya, T., et al. (2001). Stem cells, cancer, and cancer stem cells. Nature, 414(6859), 105–111.
Bonnet, D., & Dick, J. E. (1997). Human acute myeloid leukemia is organized as a hierarchy that originates from a primitive hematopoietic cell. Nature Medicine, 3(7), 730–737.
Crabtree, J. S., & Miele, L. (2018). Breast cancer stem cells. Biomedicine, 6, 3.
Hermann, P. C., et al. (2007). Distinct populations of cancer stem cells determine tumor growth and metastatic activity in human pancreatic cancer. Cell Stem Cell, 1(3), 313–323.
Ho, M. M., et al. (2007). Side population in human lung cancer cell lines and tumors is enriched with stem-like cancer cells. Cancer Research, 67(10), 4827–4833.
Yu, Z., et al. (2012). Cancer stem cells. The International Journal of Biochemistry & Cell Biology, 44(12), 2144–2151.
Sancho, P., Barneda, D., & Heeschen, C. (2016). Hallmarks of cancer stem cell metabolism. British Journal of Cancer, 114(12), 1305–1312.
Bose, S., Zhang, C., & Le, A. (2021). Glucose metabolism in cancer: The Warburg effect and beyond. Advances in Experimental Medicine and Biology, 1311, https://doi.org/10.1007/978-3-030-65768-0_1
Aguilar, E., et al. (2016). Metabolic reprogramming and dependencies associated with epithelial cancer stem cells independent of the epithelial-mesenchymal transition program. Stem Cells, 34(5), 1163–1176.
Song, K., et al. (2015). Active glycolytic metabolism in CD133(+) hepatocellular cancer stem cells: Regulation by MIR-122. Oncotarget, 6(38), 40822–40835.
Shen, Y. A., et al. (2015). Metabolic reprogramming orchestrates cancer stem cell properties in nasopharyngeal carcinoma. Cell Cycle, 14(1), 86–98.
Shibuya, K., et al. (2015). Targeting the facilitative glucose transporter GLUT1 inhibits the self-renewal and tumor-initiating capacity of cancer stem cells. Oncotarget, 6(2), 651–661.
Ciavardelli, D., et al. (2014). Breast cancer stem cells rely on fermentative glycolysis and are sensitive to 2-deoxyglucose treatment. Cell Death & Disease, 5, e1336.
Xie, H., et al. (2014). Targeting lactate dehydrogenase--a inhibits tumorigenesis and tumor progression in mouse models of lung cancer and impacts tumor-initiating cells. Cell Metabolism, 19(5), 795–809.
Le, A., et al. (2010). Inhibition of lactate dehydrogenase A induces oxidative stress and inhibits tumor progression. Proceedings of the National Academy of Sciences of the United States of America, 107(5), 2037–2042.
Rajeshkumar, N. V., et al. (2015). Therapeutic targeting of the Warburg effect in pancreatic cancer relies on an absence of p53 function. Cancer Research, 75(16), 3355–3364.
Dutta, P., et al. (2013). Evaluation of LDH-A and glutaminase inhibition in vivo by hyperpolarized 13C-pyruvate magnetic resonance spectroscopy of tumors. Cancer Research, 73(14), 4190–4195.
Li, T., Copeland, C., & Le, A. (2021). Glutamine metabolism in cancer. Advances in Experimental Medicine and Biology, 1311, https://doi.org/10.1007/978-3-030-65768-0_2
Le, A., et al. (2012). Glucose-independent glutamine metabolism via TCA cycling for proliferation and survival in B cells. Cell Metabolism, 15(1), 110–121.
Kim, J. H., et al. (2018). Effects of metformin on colorectal cancer stem cells depend on alterations in glutamine metabolism. Scientific Reports, 8(1), 409.
Zimmermann, S. C., et al. (2016). Allosteric glutaminase inhibitors based on a 1,4-di(5-amino-1,3,4-thiadiazol-2-yl)butane scaffold. ACS Medicinal Chemistry Letters, 7(5), 520–524.
Rais, R., et al. (2016). Discovery of 6-diazo-5-oxo-l-norleucine (DON) prodrugs with enhanced CSF delivery in monkeys: a potential treatment for glioblastoma. Journal of Medicinal Chemistry, 59(18), 8621–8633.
Xiang, Y., et al. (2015). Targeted inhibition of tumor-specific glutaminase diminishes cell-autonomous tumorigenesis. The Journal of Clinical Investigation, 125(6), 2293–2306.
Dang, C. V., et al. (2011). Therapeutic targeting of cancer cell metabolism. Journal of Molecular Medicine (Berlin), 89(3), 205–212.
Hirschey, M. D., et al. (2015). Dysregulated metabolism contributes to oncogenesis. Seminars in Cancer Biology, 35(Suppl), S129–S150.
Elgogary, A., et al. (2016). Combination therapy with BPTES nanoparticles and metformin targets the metabolic heterogeneity of pancreatic cancer. Proceedings of the National Academy of Sciences of the United States of America, 113(36), E5328–E5336.
Udupa, S., et al. (2019). Upregulation of the glutaminase II pathway contributes to glutamate production upon glutaminase 1 inhibition in pancreatic cancer. Proteomics, 19(21–22), e1800451.
Nguyen, T., et al. (2019). Uncovering the role of N-acetyl-aspartyl-glutamate as a glutamate reservoir in cancer. Cell Reports, 27(2), 491–501. e6.
Li, D., et al. (2015). Inhibition of glutamine metabolism counteracts pancreatic cancer stem cell features and sensitizes cells to radiotherapy. Oncotarget, 6(31), 31151–31163.
Cuyas, E., et al. (2018). Mitostemness. Cell Cycle, 17(8), 918–926.
Liu, P. P., et al. (2014). Metabolic regulation of cancer cell side population by glucose through activation of the Akt pathway. Cell Death and Differentiation, 21(1), 124–135.
Peiris-Pages, M., et al. (2016). Cancer stem cell metabolism. Breast Cancer Research, 18(1), 55.
Lonardo, E., et al. (2013). Metformin targets the metabolic Achilles heel of human pancreatic cancer stem cells. PLoS One, 8(10), e76518.
Sancho, P., et al. (2015). MYC/PGC-1alpha balance determines the metabolic phenotype and plasticity of pancreatic cancer stem cells. Cell Metabolism, 22(4), 590–605.
Reid, M. A., Dai, Z., & Locasale, J. W. (2017). The impact of cellular metabolism on chromatin dynamics and epigenetics. Nature Cell Biology, 19(11), 1298–1306.
Wainwright, E. N., & Scaffidi, P. (2017). Epigenetics and cancer stem cells: Unleashing, hijacking, and restricting cellular plasticity. Trends Cancer, 3(5), 372–386.
Mitani, M., et al. (1976). Salinomycin effects on mitochondrial ion translocation and respiration. Antimicrobial Agents and Chemotherapy, 9(4), 655–660.
Gupta, P. B., et al. (2009). Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell, 138(4), 645–659.
Zhi, Q. M., et al. (2011). Salinomycin can effectively kill ALDH (high) stem-like cells on gastric cancer. Biomedicine & Pharmacotherapy, 65(7), 509–515.
Zhang, G. N., et al. (2011). Combination of salinomycin and gemcitabine eliminates pancreatic cancer cells. Cancer Letters, 313(2), 137–144.
Lyakhovich, A., & Lleonart, M. E. (2016). Bypassing mechanisms of mitochondria-mediated cancer stem cells resistance to chemo- and radiotherapy. Oxidative Medicine and Cellular Longevity, 2016, 1716341.
Garcia-Heredia, J. M., & Carnero, A. (2015). Decoding Warburg’s hypothesis: Tumor-related mutations in the mitochondrial respiratory chain. Oncotarget, 6(39), 41582–41599.
Hirsch, H. A., et al. (2009). Metformin selectively targets cancer stem cells, and acts together with chemotherapy to block tumor growth and prolong remission. Cancer Research, 69(19), 7507–7511.
Dattilo, R., et al. (2020). Pyrvinium pamoate induces death of triple-negative breast cancer stem-like cells and reduces metastases through effects on lipid anabolism. Cancer Research, 80(19), 4087–4102.
Fiorillo, M., et al. (2016). Repurposing atovaquone: Targeting mitochondrial complex III and OXPHOS to eradicate cancer stem cells. Oncotarget, 7(23), 34084–34099.
Fiorillo, M., et al. (2016). Bedaquiline, an FDA-approved antibiotic, inhibits mitochondrial function and potently blocks the proliferative expansion of stem-like cancer cells (CSCs). Aging (Albany NY), 8(8), 1593–1607.
Sosa, V., et al. (2013). Oxidative stress and cancer: An overview. Ageing Research Reviews, 12(1), 376–390.
Redza-Dutordoir, M., & Averill-Bates, D. A. (2016). Activation of apoptosis signalling pathways by reactive oxygen species. Biochimica et Biophysica Acta, 1863(12), 2977–2992.
Fan, P. C., et al. (2019). Quantitative proteomics reveals mitochondrial respiratory chain as a dominant target for carbon ion radiation: Delayed reactive oxygen species generation caused DNA damage. Free Radical Biology & Medicine, 130, 436–445.
Yang, H., et al. (2018). The role of cellular reactive oxygen species in cancer chemotherapy. Journal of Experimental & Clinical Cancer Research, 37(1), 266.
Piao, L. S., et al. (2012). CD133+ liver cancer stem cells modulate radioresistance in human hepatocellular carcinoma. Cancer Letters, 315(2), 129–137.
Gomez-Casal, R., et al. (2013). Non-small cell lung cancer cells survived ionizing radiation treatment display cancer stem cell and epithelial-mesenchymal transition phenotypes. Molecular Cancer, 12(1), 94.
Collins, A. T., et al. (2005). Prospective identification of tumorigenic prostate cancer stem cells. Cancer Research, 65(23), 10946–10951.
Al-Hajj, M., et al. (2003). Prospective identification of tumorigenic breast cancer cells. Proceedings of the National Academy of Sciences of the United States of America, 100(7), 3983–3988.
Dalerba, P., et al. (2007). Phenotypic characterization of human colorectal cancer stem cells. Proceedings of the National Academy of Sciences of the United States of America, 104(24), 10158–10163.
Ishimoto, T., et al. (2011). CD44 variant regulates redox status in cancer cells by stabilizing the xCT subunit of system xc(-) and thereby promotes tumor growth. Cancer Cell, 19(3), 387–400.
Diehn, M., et al. (2009). Association of reactive oxygen species levels and radioresistance in cancer stem cells. Nature, 458(7239), 780–783.
Schulz, A., et al. (2019). Cancer stem cells and radioresistance: DNA repair and beyond. Cancers (Basel), 11, 6.
Park, J. K., et al. (2021). The heterogeneity of lipid metabolism in cancer. Advances in Experimental Medicine and Biology, 1311, https://doi.org/10.1007/978-3-030-65768-0_3
Sun, M., & Yang, Z. (2019). Metabolomic studies of live single cancer stem cells using mass spectrometry. Analytical Chemistry, 91(3), 2384–2391.
Gouw, A. M., et al. (2019). The MYC oncogene cooperates with sterol-regulated element-binding protein to regulate lipogenesis essential for neoplastic growth. Cell Metabolism, 30(3), 556–572. e5.
Foster, D. W. (2012). Malonyl-CoA: The regulator of fatty acid synthesis and oxidation. The Journal of Clinical Investigation, 122(6), 1958–1959.
Begicevic, R. R., Arfuso, F., & Falasca, M. (2019). Bioactive lipids in cancer stem cells. World Journal of Stem Cells, 11(9), 693–704.
Dang, C. V., Le, A., & Gao, P. (2009). MYC-induced cancer cell energy metabolism and therapeutic opportunities. Clinical Cancer Research, 15(21), 6479–6483.
Le, A., & Dang, C. V. (2013). Studying Myc’s role in metabolism regulation. Methods in Molecular Biology, 1012, 213–219.
Hanai, J. I., et al. (2013). ATP citrate lyase knockdown impacts cancer stem cells in vitro. Cell Death & Disease, 4, e696.
Rios Garcia, M., et al. (2017). Acetyl-CoA carboxylase 1-dependent protein acetylation controls breast cancer metastasis and recurrence. Cell Metabolism, 26(6), 842–855. e5.
Corominas-Faja, B., et al. (2014). Chemical inhibition of acetyl-CoA carboxylase suppresses self-renewal growth of cancer stem cells. Oncotarget, 5(18), 8306–8316.
Yasumoto, Y., et al. (2016). Inhibition of fatty acid synthase decreases expression of stemness markers in glioma stem cells. PLoS One, 11(1), e0147717.
Tracz-Gaszewska, Z., & Dobrzyn, P. (2019). Stearoyl-CoA desaturase 1 as a therapeutic target for the treatment of cancer. Cancers (Basel), 11, 7.
Gruenbacher, G., & Thurnher, M. (2018). Mevalonate metabolism in cancer stemness and trained immunity. Frontiers in Oncology, 8, 394.
Chen, C. L., et al. (2016). NANOG metabolically reprograms tumor-initiating stem-like cells through tumorigenic changes in oxidative phosphorylation and fatty acid metabolism. Cell Metabolism, 23(1), 206–219.
Kuo, C. Y., & Ann, D. K. (2018). When fats commit crimes: Fatty acid metabolism, cancer stemness and therapeutic resistance. Cancer Communications (Lond), 38(1), 47.
Hoang, G., Udupa, S., & Le, A. (2019). Application of metabolomics technologies toward cancer prognosis and therapy. International Review of Cell and Molecular Biology, 347, 191–223.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2021 The Author(s)
About this chapter
Cite this chapter
Alvina, F.B., Gouw, A.M., Le, A. (2021). Cancer Stem Cell Metabolism. In: Le, A. (eds) The Heterogeneity of Cancer Metabolism. Advances in Experimental Medicine and Biology, vol 1311. Springer, Cham. https://doi.org/10.1007/978-3-030-65768-0_12
Download citation
DOI: https://doi.org/10.1007/978-3-030-65768-0_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-65767-3
Online ISBN: 978-3-030-65768-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)