Abstract
Neural image denoising is a promising approach for quality enhancement of ultra-low dose (ULD) CT scans after image reconstruction. The availability of high-quality training data is instrumental to its success. Still, synthetic noise is generally used to simulate the ULD scans required for network training in conjunction with corresponding normal dose scans. This reductive approach may be practical to implement but ignores any departure of the real noise from the assumed model. In this paper, we demonstrate the training of denoising neural networks with real noise. For this purpose, a special training set is created from a pair of ULD and normal-dose scans acquired on each subject. Accurate deformable registration is computed to ensure the required pixel-wise overlay between corresponding ULD and normal-dose patches. To our knowledge, it is the first time real CT noise is used for the training of denoising neural networks. The benefits of the proposed approach in comparison to synthetic noise training are demonstrated both qualitatively and quantitatively for several state-of-the art denoising neural networks. The obtained results prove the feasibility and applicability of real noise learning as a way to improve neural denoising of ULD lung CT.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
1 Introduction
Neural image denoising and its application to low-dose CT scans has become a dynamic field of research [1,2,3,4,5,6,7,8]. In [1], the patches sequentially extracted from the input slice by a 2-D sliding window are processed by a multi-layer perceptron and directly replaced by the denoised output patch. The denoised image is obtained by spatial averaging of the overlapping output patches. A fully-connected neural network is trained in [2] to denoise coronary CT angiography patches. The denoised image is generated by a locally-consistent non-local-means algorithm [9] applied to the denoised patches.
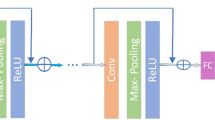
In [3], a convolutional neural network (CNN) is proposed for denoising. The CNN layers are purely convolutional, each one being followed by batch-normalization [10] and a (rectified linear unit) ReLU non-linearity. To improve denoising performance, the residual between ground-truth (GT) and input, thus representing a noise image, is learned by the network instead of the habitual mapping between noisy input and GT image.
In [4], a denoised instance of the input image is first computed by the classical BM3D algorithm [11]. Next, similar image patches from the denoised instance, are stacked together into blocks with noisy image patches extracted at the same position. The blocks are subsequently denoised by the network proposed in [3] and the output image is obtained by aggregation of the denoised patches.
In [5], a 3-layers fully CNN (FCNN) is trained on pairs of corresponding clean and noisy CT patches. The noisy images, from which the training patches are extracted, are generated by Poisson noise addition to the clean images in the sinogram domain. L2 distance metric is used for network loss computation. Leveraging the FCNN architecture, training on image patches allows for direct denoising of full size images.
A perceptual loss was proposed in [6] as an improvement to the commonly used L2 loss between denoised and original clean images. The perceptual loss is computed in a high-dimensional feature space generated by the well-known VGG network [12]. The proposed CNN is trained to minimize the L2 loss between the VGG features extracted from the denoised image and the same extracted from the GT clean image. In [7], a residual encoder-decoder architecture is proposed with skip connections. The encoding layers (convolutions) extract the image features, while the decoding layers successively reconstruct the denoised image from the extracted features (deconvolutions). Generative adversarial networks (GAN) [13] have also been proposed for low dose CT denoising in combination with a sharpness detection network to avoid oversmoothing of the output [8].
While previous works have proposed a variety of network architectures, they all share a basic limitation dictated by convenience: the networks are trained on synthetic data obtained by artificial addition of parametric noise, usually assumed to be Poisson-distributed, in the sinogram domain. Consequently, any departure from the assumed noise model is ignored. In this work, we propose the training of denoising networks with a dataset built on real ultra-low dose/normal-dose pairs of scans, acquired for each patient. The proposed approach allows, for the first time to our knowledge, the learning of real ultra-low dose (ULD) CT noise by denoising neural networks. The remainder of this paper is organized as follows: The construction of the real noise dataset is described in Sect. 2 and validated, both qualitatively and quantitatively, in Sect. 3. A discussion concludes the paper in Sect. 4.
2 Methods
A dataset of 20 cases was created by scanning each subject twice, once at normal dose and once at ULD. The normal-dose scan was acquired at 120 kVp under adaptive tube current modulation. The ULD scan was acquired at 120 kVp at a constant tube current of 10 or 20 mA for subjects with a body mass index (BMI) below or above 29, respectively. Prior written consent was obtained for each patient as required in the IRB authorization released by our institution. The cases consisted of adult subjects, over 18 years old for men, and 50 for women to avoid risks of undetected pregnancy. The subjects were recruited among patients referred to a standard chest CT, with or without contrast media. The reduction in radiation dose (R) between normal and ULD scans is given by (Eq. 1):
where \( DLP_{ULD} , \,\text{and}\;DLP_{normal} \) are the dose-length-product of the ULD and normal-dose scans, respectively. The average R across the dataset was about 94%. The dose report for a sample subject is shown in Fig. 1 with (green) \( DLP_{normal} = 317.4\,{\rm mGy.cm} \) and (red) \( DLP_{ULD} = 13.58\,{\rm mGy.cm} \), corresponding to R = 96%. Consequently, the additional radiation dose incurred by the patient undergoing the two scans is merely of 4%, keeping the overall radiation exposure (total exam DLP) at an acceptable level for a lung CT scan. The scans are performed at full inspiration under breath-hold. Since the total scan time may be too long for a single breath-hold, each scan is acquired under a separate one. However, as lung inflation is not exactly the same in both scans, a deformable registration step is required to ensure accurate pixel-wise overlay between the scans, so that pairs of corresponding normal-dose and ULD patches can be extracted to train the neural networks.
The registration process involves two images: a fixed image (IF) and a moving image (IM). It recovers a spatial transformation T(x) = x + u(x) such that IM(T(x)) is spatially aligned to IF(x). To compute T(x), the formulation of the Elastix package [14] was used with a third order b-spline parametrization. Minimization was performed for the cost function given by (Eq. 2):
where, S is a similarity function computed between IF and IM(T(x)) and P is a regularization function that constrains T(x). Mutual information was chosen as function S, and bending energy as function P [14]. The γ parameter balances between similarity and regularity in the cost function. To compensate for small changes in patient position and orientation, a 3-D rigid transform (6 degrees of freedom) was first computed between the ULD and the normal-dose scans, before the deformable step. The parameter values for both registration steps were chosen as in [15]. The normal-dose scan was taken as reference image (IF) for all the registration tasks. A sample case slice is shown in Fig. 2. The ULD image (a) is shown alongside the normal-dose (b) and co-registered ULD (T-ULD) (c) images. Highlighted ROIs (red, green, blue) are zoomed-in in (d): ULD (left column), normal-dose (middle) and T-ULD (right). The ULD ROIs (left) are clearly misaligned with the normal-dose ones (middle), while the latter are well aligned with the T-ULD ROIs. Before using the pairs of aligned patches to train neural networks, we perform a patch selection step in order to remove pairs with insufficient registration accuracy from the generated training set. We define the selected patch dataset D as (Eq. 3):
Sample case slice: (a) ULD image; (b) normal-dose image; (c) co-registered ULD (T-ULD). (d) Zoom-in ROIs (red, green, yellow): ULD (left column), normal-dose (middle) and T-ULD (right). The ULD ROIs (left) are clearly misaligned with the normal-dose ones (middle), while the latter are well aligned with the T-ULD ROIs. (Color figure online)
Where, pi and qi are normal-dose and T-ULD patch pairs, respectively, SIM is a similarity measure and TSIM is a threshold. We used the structural similarity index (SSIM) [16] as the similarity measure and set empirically TSIM to be 0.4. The underlying idea being that improved alignment accuracy will result in increased content similarity between the compared patches, leading to higher SSIM values. After selection, the resulting real-noise database contained about 700,000 patch pairs of size 45 × 45 pixels.
3 Experiments
For the validation of the proposed approach, four state-of-the-art denoising CNN were implemented: LD-CNN [5], DnCNN [3], RED-CNN [7] and CNN-VGG [6]. Each architecture was trained in two different version: a first using the real-noise database, and a second using synthetic noise. The synthetic noise database was created using the ASTRA toolbox [17] by adding Poisson noise with I0 = 22 × 104 to the computed sinogram of each normal-dose image in order to simulate R ~ 90% [2].
A 5-folds cross-validation scheme was adopted. In each fold, 4 cases were used for testing and all the remaining, for training. In total, 40 different networks were created (5 folds × 4 architectures × 2 training sets). The quantitative comparison was performed between denoised T-ULD and normal-dose scans for SSIM and PSNR scores. Here again, to avoid affecting the denoising scores by registration inaccuracies, SSIM and PSNR were only computed at locations where (1) was satisfied. Note that the SSIM computed in (1) for patch selection is between normal-dose and T-ULD, whereas the SSIM of the validation is between denoised T-ULD and normal-dose. In Fig. 3, quantitative comparison of the PSNR (a–d) and SSIM (e–h) for the LD-CNN, DnCNN, RED-CNN and CNN-VGG networks is shown for each fold, following real (blue) and synthetic (red) noise training. The networks trained with real-noise, outperformed the SSIM (e–h) of their synthetic noise counterpart for all the folds and architectures. For PSNR (a–d), the same situation is observed, except for the third fold, in which CNN-VGG was the only network with higher PSNR for real-noise training. Also, it is interesting to note that although LD-CNN has a simple architecture, consisting of only three convolutional layers, it generated the best PSNR and SSIM scores in most of the cases. A possible reason may be that wider and shallow networks are better in capturing pixel-distribution features [18], a property which is critical in the case of real noise learning. Denoising results for a sample slice are shown in Fig. 4. The highlighted ROIs (red, green, yellow) in the ULD (a) and normal-dose images (b) are denoised and zoomed-in (c–f) by DnCNN (c), LD-CNN (d), RED-CNN (e) and CNN-VGG (f) for real (left columns) and synthetic (right columns) noise training. The results obtained for real-noise training appear sharper, with more fine details visible in comparison to the synthetic training results, which exhibit some over-smoothing. This is further verified by computing the overall image sharpness from the approximate sharpness map, S3(x), defined in [19] by (Eq. 4):
A sample result slice. The highlighted ROIs (red, green, yellow) in the ULD (a) and normal-dose images (b) are denoised and zoomed-in (c–f) by DnCNN (c), LD-CNN (d), RED-CNN (e) and CNN-VGG (f) for real (left columns) and synthetic (right columns) noise training. The reader is encouraged to zoom in for a better assessment of sharpness. (Color figure online)
Where, S1(x) is a spectral-based sharpness map and S2(x) is a spatial based sharpness map. The overall image sharpness is given by the average of the top 1% values in S3(x) and is denoted by \( \varvec{S}_{3} \) [19]. The average \( \varvec{S}_{3} \) index computed on the denoised images for all the networks trained with real-noise (blue) and synthetic-noise (red) are shown in Fig. 5 for the 5 folds. In all, but one, fold of a single algorithm (CNN-VGG), the overall sharpness index was better for real-noise training than for synthetic, confirming the visual impression of Fig. 4.
The average \( \varvec{S}_{3} \) values [19] of the denoising results obtained by networks trained with real (blue) and synthetic (red) noise. (Color figure online)
In a last experiment, the local contrast of a tiny 2.5 mm lung nodule was compared after denoising with real and synthetic noise training. Figure 6 shows the ROI containing the nodule (a, top) zoomed-in after denoising (red arrow) by the four state-of-the-art CNNs (b–e). The nodule appears with stronger contrast for real-noise training (top row images) than for synthetic noise (bottom row images). This is further confirmed in Table 1, by the Michelson contrast, MC, [20] which computes the normalized difference between maximal and minimal intensities measured and averaged along radial profiles (green lines, (a)-bottom) of the nodule.
Tiny lung nodule (a, top) in red ROI zoomed-in after denoising (red arrow) by the four state-of-the-art CNNs (b–e). The nodule appears with stronger contrast for real-noise training (top row images) than for synthetic noise (bottom row images). The Michelson contrast is computed along the green sampling lines (a, bottom) on the nodule. (Color figure online)
4 Conclusions
In this paper, the learning of real noise was proposed for neural network denoising of ULD lung CT. While synthetic noise has been widely used in prior works, it is the first time, to our knowledge, that real CT noise, extracted from real pairs of co-registered ULD and normal-dose scans, is used for training denoising neural networks. In particular, the proposed combination of deformable registration and double scanning enables the computation of quantitative denoising scores in real denoising tasks whereas these were only computed for synthetic noise in previous works. The benefits of the proposed approach were demonstrated on four state-of-the-art neural denoising architectures. Beside the improvement in PSNR and SSIM, a substantial improvement in image sharpness was observed for the images denoised using real-noise trained networks. Local contrast improvement was also demonstrated for a tiny lung nodule following denoising by the proposed approach. The obtained results suggest that the proposed approach is a promising way to improve neural denoising of ULD lung CT scans.
References
Burger, H.C., Schuler, C.J., Harmeling, S.: Image denoising: can plain neural networks compete with BM3D? In: IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2012)
Green, M., Marom, E.M., Kiryati, N., Konen, E., Mayer, A.: A neural regression framework for low-dose Coronary CT Angiography (CCTA) denoising. In: Wu, G., Munsell, B.C., Zhan, Y., Bai, W., Sanroma, G., Coupé, P. (eds.) Patch-MI 2017. LNCS, vol. 10530, pp. 102–110. Springer, Cham (2017). https://doi.org/10.1007/978-3-319-67434-6_12
Zhang, K., Zuo, W., Chen, Y., Meng, D., Zhang, L.: Beyond a gaussian denoiser: residual learning of deep CNN for image denoising. IEEE Trans. Image Process. 26, 3142–3155 (2017)
Ahn, B., Cho, N.I.: Block-matching convolutional neural network for image denoising. arXiv preprint arXiv:1704.00524 (2017)
Chen, H., et al.: Low-dose CT via convolutional neural network. Biomed. Opt. Express 8(2), 679–694 (2017)
Yang, Q., Yan, P., Kalra, M.K., Wang, G.: CT image denoising with perceptive deep neural networks. arXiv preprint arXiv:1702.07019 (2017)
Chen, H., et al.: Low-dose CT with a residual encoder-decoder convolutional neural network (RED-CNN). arXiv preprint arXiv:1702.00288 (2017)
Yi, X., Babyn, P.: Sharpness-aware low dose CT denoising using conditional generative adversarial network. arXiv preprint arXiv:1708.06453 (2017)
Green, M., Marom, E.M., Kiryati, N., Konen, E., Mayer, A.: Efficient low-dose CT denoising by locally-consistent non-local means (LC-NLM). In: Ourselin, S., Joskowicz, L., Sabuncu, M.R., Unal, G., Wells, W. (eds.) MICCAI 2016. LNCS, vol. 9902, pp. 423–431. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-46726-9_49
Ioffe, S., Szegedy, C.: Batch normalization: accelerating deep network training by reducing internal covariate shift. In: International Conference on Machine Learning (2015)
Dabov, K., Foi, A., Katkovnik, V., Egiazarian, K.: Image denoising by sparse 3-D transform-domain collaborative filtering. IEEE Trans. Image Process. 16(8), 2080–2095 (2007)
Simonyan, K., Zisserman, A.: Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556 (2014)
Goodfellow, I., et al.: Generative adversarial nets. In: Advances in Neural Information Processing Systems (2014)
Klein, S., Staring, M., Murphy, K., Viergever, M.A., Pluim, J.P.: Elastix: a toolbox for intensity-based medical image registration. IEEE Trans. Med. Imaging 29(1), 196–205 (2010)
Sokooti, H., Saygili, G., Glocker, B., Lelieveldt, B.P.F., Staring, M.: Accuracy estimation for medical image registration using regression forests. In: Ourselin, S., Joskowicz, L., Sabuncu, M.R., Unal, G., Wells, W. (eds.) MICCAI 2016. LNCS, vol. 9902, pp. 107–115. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-46726-9_13
Wang, Z., Bovik, A.C., Sheikh, H.R., Simoncelli, E.P.: Image quality assessment: from error visibility to structural similarity. IEEE Trans. Image Process. 13(4), 600–612 (2004)
van Aarle, W., et al.: Fast and flexible X-ray tomography using the ASTRA toolbox. Opt. Express 24(22), 25129–25147 (2016)
Liu, P., Fang, R.: Wide inference network for image denoising. arXiv preprint arXiv:1707.05414 (2017)
Vu, C.T., Phan, T.D., Chandler, D.M.: S3: a spectral and spatial measure of local perceived sharpness in natural images. IEEE Trans. Image Process. 21(3), 934–945 (2012)
Michelson, A.A.: Studies in Optics. Courier Corporation (1995)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Switzerland AG
About this paper
Cite this paper
Green, M., Marom, E.M., Konen, E., Kiryati, N., Mayer, A. (2018). Learning Real Noise for Ultra-Low Dose Lung CT Denoising. In: Bai, W., Sanroma, G., Wu, G., Munsell, B., Zhan, Y., Coupé, P. (eds) Patch-Based Techniques in Medical Imaging. Patch-MI 2018. Lecture Notes in Computer Science(), vol 11075. Springer, Cham. https://doi.org/10.1007/978-3-030-00500-9_1
Download citation
DOI: https://doi.org/10.1007/978-3-030-00500-9_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-00499-6
Online ISBN: 978-3-030-00500-9
eBook Packages: Computer ScienceComputer Science (R0)