Abstract
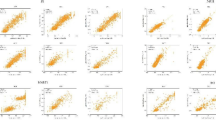
This paper presents the MRBF network, a new algorithm adapted from the RBF network, to construct the classifiers for predicting phenotypic resistance on 6 protease inhibitors. The performance of the prediction was measured by 10-fold cross-validation, The results show that MRBF gives the lowest average mean square error (MSE) when compared with the traditional RBF network and multiple linear regression analysis (REG). Moreover, it provides the best average predictive accuracy when compared with HIVdb, REG, and Support Vector Machines (SVM).
Chapter PDF
Similar content being viewed by others
References
J.M. Coffin, “HIV population dynamics in vivo: implications for genetic variation, pathogenesis, and therapy”, Science, vol. 267, 1995, pp. 483–489.
L. Demeter, R. Haubrich, “Phenotypic and genotypic resistance assays: methodology, reliability, and interpretations”, Journal of Acquired Immune Deficiency Syndromes, vol. 26, 2001, pp. S3–S9.
M. Robnik Sikonja and I. Kononenko, “An adaptation of relief for attribute estimation in regression”, Machine Learning: Proceedings of the Fourteenth International Conference (ICML’ 97), 1997, pp. 296–304.
R.W. Shafer, D.R. Jung, B.J. Betts, “Human immunodeficiency virus type 1 reverse transcriptase and protease mutation search engine for queries”, NAT Med, vol. 6, 2000, pp. 1290–1292.
JL. Meynard, M. Vray, L. Morand-Joubert, et al, “Phenotypic or genotypic resistance testing for choosing antiretroviral therapy after treatment failure: a randomized trial”, AIDS, vol. 16, 2002, pp. 727–736.
K. Van Laethem, A. Ke Luca, A, Antinori, et al. “A genotypic drug resistance interpretation algorithm that significantly predicts therapy response in HIV-1 infected patients”, Antiviral Ther, vol. 7, 2002, pp. 123–129.
C. Reid, R. Bassett, S. Day, et al, “A dynamic rules-based interpretation system derived by an expert panel is predictive of virological failure”, Antiviral Ther, vol. 7, 2002, pp. s91.
K. Wang, E. Jenwitheesuk, R. Samudrala, J.E. Mitter, “Simple linear model provides highly accurate genotypic predictions of HIV-1 drug resistance”, Antivir, Ther, vol. 9, 2004, pp. 343–352.
N. Beerenwinkel, B. Schmidt, H. Walter, R. Kaiser, T. Lengauer, D. Hoffmann, K. Korn, and J. Selbig, “Geno2pheno: interpreting genotypic HIV drug resistance test”, IEEE Intellig Syst, vol. 16, 2001, pp. 35–41.
N. Beerenwinkel, B. Schmidt, H. Walter, R. Kaiser, T. Lengauer, D. Hoffmann, K. Korn, and J. Selbig, “Diversity and complexity of HIV-1 drug resistance: a bioinformatics approach to prediction phenotype from genotype”, Proceedings of Natl Acad, Sc, USA, 2002, pp. 8271–8276.
N. Beerenwinkel, M. Daumer, M Oette, K. Korn, D. Hoffmann, R. Kaiser, T. Lengauer, J. Selbig, and H. Walter, “Geno2Pheno: estimating phenotypic drug resistance from HIV-1 genotypes”, Nucleic Acids Research, vol. 31, 2003, pp. 3850–3855.
D. Wang and B. Larder, “Enhanced prediction of lopinavir resistance from genotype by use of artificial neural networks”, Infectious Disease, vol. 188, 2003, pp. 653–660.
S. Draghici and B. Potter, “Predicting HIV drug resistance with neural networks”, Bioinformatics, vol. 19, 2003, pp. 98–107.
M. Powell, “Radial basis function for multivariable interpolation: A review”, Algorithms for approximation, 1987, pp. 143–167.
D. S. Broomhead and D. Lowe, “Mutivariable functional interpolation and adaptive networks”, Complex System 2, 1988, pp. 321–355.
J. Moody and C. J. Darken, “Fast learning in networks of locally-tuned processing units”, Neural Computation, 1(2), 1989, pp. 281–294.
S. Haykin, Neural Networks; A comprehensive foundation, Prentice Hall, New Jersey, 1999, pp. 256–312.
S. W. Robert, “Genotypic Testing for Human Immunodeficiency Virus Type 1 Drag Resistance”, Clinical Microbiology Regviews, vol. 15, 2002, pp. 247–277.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2006 International Federation for Information Processing
About this paper
Cite this paper
Srisawat, A., Kijsirikul, B. (2006). MRBF: A Method for Predicting HIV-1 Drug Resistance. In: Shi, Z., Shimohara, K., Feng, D. (eds) Intelligent Information Processing III. IIP 2006. IFIP International Federation for Information Processing, vol 228. Springer, Boston, MA. https://doi.org/10.1007/978-0-387-44641-7_34
Download citation
DOI: https://doi.org/10.1007/978-0-387-44641-7_34
Publisher Name: Springer, Boston, MA
Print ISBN: 978-0-387-44639-4
Online ISBN: 978-0-387-44641-7
eBook Packages: Computer ScienceComputer Science (R0)