Abstract
In the analysis of continuous data, researchers are often faced with the problem that statistical methods developed for single-point data (e.g., t test, analysis of variance) are not always appropriate for their purposes. Either methodological adaptations of single-point methods will need to be made, or confidence bands are the method of choice. In this article, we compare three prominent techniques to analyze continuous data (single-point methods, Gaussian confidence bands, and function-based resampling methods to construct confidence bands) with regard to their testing principles, prerequisites, and outputs in the analysis of continuous data. In addition, we introduce a new technique that combines the advantages of the existing methods and can be applied to a wide range of data. Furthermore, we introduce a method enabling a priori and a posteriori power analyses for experiments with continuous data.
Similar content being viewed by others
In behavioral science, researchers often deal with continuous data—for example, when comparing kinematic profiles of different movements or the same movement under different conditions. Differences that emerge in the varying profiles typically extend over a series of subsequent data points and are, thus, not limited to a single point. Although the general logic of statistically detecting differences between two streams of continuous data is similar to the analysis of single-point data, there are crucial differences in the concrete methodical steps of the analysis. The present article will describe and discuss three existing methods to analyze continuous data. We will then contrast these methods with a new method introduced here.
First, we need to elaborate the differences between single-point and continuous data and why it is important to differentiate them in terms of data analysis. In single-point data analysis, the attribute Φsingle is represented by one single data point Y. This is the case when measuring, for example, reaction times, jumping heights, or systolic blood pressure. In contrast to features that can be represented by a single value, other attributes Φmultiple can only be described by a number of sequential or parallel measurements of the observed quantity, producing multiple data points Y(1…n), where n is always greater than 1. Examples of these attributes are the course of the knee angle during the gait cycle or the neural activity associated with processing a certain stimulus (e.g., measured with electroencephalography [EEG] recordings). According to this logic, any multiple data recordings stored in a single vector could be considered as continuous data.Footnote 1 However, the methods discussed in the following paragraphs require that the consecutive data points be arranged in an ordered sequence in which each data index refers to the same subfeature of Φmultiple. Together, the ordered data vector describes a measured feature in the world. In an experiment, the manifestation of Φsingle or Φmultiple is usually measured and analyzed in different conditions—that is, for subsamples with different values of an independent variable. Thus, when measuring Φsingle for each subsample, the data set (DS) will consist of k measurements of Y values, where k is the number of recordings DSsingle = {Y1, …Yk}. When testing Φmultiple, however, the data set will contain k recordings of \( {Y}_{\left(1\dots n\right)},{DS}_{multiple}=\left\{{Y}_{\left(1\dots n\right)}^1,\dots, {Y}_{(1..n)}^k\right\} \). In this regard, it is important that the same indices j ∈ {1, .., n} of Y(1…n) represent the same aspect of Φmultiple in all k recordings. For example, when capturing the time course of joint angles during walking, the resulting data sets need to be time-locked with respect to appropriate (e.g., kinematic) landmarks of the gait cycle. Analogously, EEG signals are often synchronized with respect to specific trigger events, such as the appearance of a stimulus or the onset of a response, which are located at the same index j in all Y(1…n). Thus, the data must be synchronized and normalized in time before further analyses can be conducted.
Despite the different characteristics of single (Y) and continuous (Y(1…n)) data, the logical sequences of the data analysis are closely related for both types. The initial step in analyzing both single and continuous data is to calculate descriptive parameters (e.g., the mean μ and standard deviation σ) that provide information about the distribution and variation of the measured data. For continuous data, the internal dependence (ρ) of the data within Y(1…n) is an additional parameter that must be taken into account. It is typically quantified by a correlation coefficient ρ, ranging from 0 ≤ |ρ| ≤ 1.
To calculate μ, σ, and ρ, the question of how many of the data can maximally be used for a reliable estimation must be considered. As we mentioned before, the natural characteristic of continuous data is that subsequent data points are sampled continuously during data acquisition. The data segments that represent one trial (or one gait cycle, one response to a stimulus, etc.) are defined by start–stop signals or synchronization triggers. The corresponding time window containing these data points can be termed the sampling window (Fig. 1, red). However, using all the data within the sampling window (i.e., all data points of one recording) for the calculation of the descriptive parameters is not always appropriate because the sampling window may include data captured at a time when the hypothesized effect of the independent variable(s) is not (yet) observable. A specific time window should be defined to focus the analysis on the relevant part of the data set in which the expected effect will occur. The data within this time window, which we will refer to as the effect window (Fig. 1, dashed lines), can then be analyzed separately. The effect window is defined a priori on the basis of preliminary data or findings reported in the relevant literature. Note that this necessity to define a priori which part of the data one should look at is not a particular difficulty for the analyses described here. The deliberate choice of any dependent variable requires the same type of considerations.
Two types of time windows are important when analyzing continuous data. The sampling window (t1, . . . , tn) represents the whole data set, from the start until the end of the recording. The effect window (dashed) is the time range over which the hypothesized effect is expected. The effect window must be defined a priori by the researcher
With the help of the descriptive parameters, the data can be further analyzed. Typically, differences of Φ between the different conditions based on the corresponding measurements Y are analyzed by using inference statistical methods. To this end, probabilities (p values) are calculated with which the observed distribution of Y values would occur if the null hypothesis were true, using the descriptive parameters μ, σ, and ρ (ρ only for continuous data). On the basis of these probabilities, a decision to reject or accept the alternative hypothesis can be made. In addition to hypothesis testing, a power analysis can be used to evaluate the experimental design and the explanatory power of the results.
As we mentioned above, the general logic of statistical hypothesis testing is similar for both single and continuous data. A problem arises, however, when choosing adequate inference statistical methods to estimate the distribution of values under the assumption of the null hypothesis being true. A wide range of statistical methods can be applied to single data Y(k) (e.g., student’s t test and analysis of variance), but only a few approaches actually tackle the problem of implementing the corresponding analysis steps for continuous data.
In the following paragraphs, we will characterize four statistical methods that can be applied to continuous data. Two of them attempt to adjust commonly used local statistical methods in order to apply them to continuous data. The other two approaches are specifically designed to deal with the characteristics of continuous in contrast to single data points (overview in Fig. 2).
The nature of any local method is that single data points Y are analyzed. Accordingly, the application of local methods to continuous data implies the selection of a single data point at one specific moment in time or in reference to a predetermined landmark, and thus a reduction to a single-data analysis. The major benefit of such an approach is that it allows the use of a wide range of existing inferential statistical methods (e.g., confidence intervals, p value estimates for different distributions [t, F, χ2, . . .], analysis of variance, power analysis, etc.). However, there are critical reasons why the simple application of local methods to continuous data is often not reasonable. First, there is the problem of validity because a single data point is chosen to represent the effect of the independent variable in the continuous data. In some cases it is not easy to adequately select this representative data point, to identify its position in the data stream, or even to find a valid representative at all, meaning that some (truly) multiple feature Φmultiple of the data cannot adequately be described by a single value. Second, reliability might be reduced since the value of a single point measure can be affected by stochastic fluctuations. As a consequence, the tested effect might be over- or underestimated.
The local Gaussian method (Fig. 2) is an alternative that combines the statistical options of the local methods (e.g., t test) with the inclusion of more than one data point in the analysis. More specifically, the local Gaussian method applies the same local test repeatedly on each index within the effect window. As a result, confidence intervals are calculated for each index which together form a confidence band (Fig. 2, broken curves). Two major issues need to be taken into account when repeatedly applying methods designed for single data points to continuous data. First, every data point of the tested curve must belong to a specific symmetric parametric family (Gaussian distribution). This prerequisite must be assured before testing. However, the data often do not fulfill this requirement of a normal distribution of data points at every index of the data set. As a consequence, the true coverage probability of the confidence bands is not even close to the desired nominal level in most cases. Second, this method does not consider the internal dependence ρ of the data points within a continuous data set. In the case, for instance, that the internal dependence of the analyzed data is ρ = 1, any single value of Y1 − Yn carries the same information, and the analysis is thus equivalent to the single-point case. If, on the other hand, it is the case that ρ = 0, a Bonferroni correction could be applied. However, with an increasing number of data points to be included in the confidence band, the width of the band increases so that down-sampling or up-sampling would severely affect the results. In most cases, the local method would be incorrect and Bonferroni corrections would be extremely conservative because the true internal dependence is, unbeknownst to the observer, located somewhere in-between 0 and 1 (Lenhoff et al., 1999). Thus, if not every data point is normally distributed and/or the value of ρ is neither close to 0 nor 1, as it is commonly the case, continuous data should not be analyzed using local Gaussian methods.
Alternatives to the local methods are methods that use simultaneous confidence bands to test deviations in continuous data curves. For instance, Lenhoff et al. (1999) applied a bootstrap procedure to construct simultaneous confidence bands of continuous gait data. The bootstrap procedure is a nonparametric resampling technique, and hence Gaussian distribution is not a prerequisite (Efron, 1979; Tukey, 1958). Moreover, it takes into account the inner dependency ρ of continuous data by describing the variance in the data as a variation of the coefficients of the fitting function (SEFunc) and evaluates its variance–covariance matrix. Therefore, this method is suitable for computing simultaneous confidence intervals for several subsequent data points of continuous signals. An essential part of the method introduced by Lenhoff and colleagues is that they fit gait cycle curves with mathematical functions so that the interdependence of the data points would be reflected by these functions. With the help of a correction factor C, which is determined using a resampling technique (e.g., bootstrap), their method can assure that the true coverage probability of the confidence bands comes close to the desired nominal level. Thus, this function-based resampling technique (FBRT), as we call it, is an advanced tool that can be applied even to data with asymmetric distributions. Although the FBRT seems to be the appropriate tool for analyzing continuous data in general, it still has limitations that have to be considered. One issue can arise from the high computational cost of this procedure. The ideal bootstrap sample consists of mm subsamples (where m stands for the sample size). As m becomes large, it becomes unfeasible to calculate all possible subsamples. However, an approximation can be used. Our simulations show that the estimation variability of the correction factor C saturates with a reasonable number of calculations. A more critical limitation of the FBRT procedure, as the authors described it, is that it cannot be applied to all kinds of continuous data. Lenhoff and colleagues fitted gait data with mathematical functions representing the data as closely as possible. However, recorded data cannot always be represented by adequate fits—for instance, when no function family exists that is capable of fitting all the entities (see Appendix 2). EEG data, for example, are characterized by rapid changes in voltage polarity that cannot reasonably be represented by a single mathematical function prototype. Another limitation of the FBRT, as the authors describe it, is that the C factor is calculated using the data from only one sample (mostly representing the data of a null/control condition). By looking at a single data point analysis, for example the t test, the estimation of the variance parameter would integrate the data from all measured samples, which should in consequence lead to a more reliable estimation.
In conclusion, all of the described methods to statistically analyze continuous data have inherent problems that limit their scopes of application. The FBRT of Lenhoff et al. (1999) seems to be the most appropriate and useful approach with respect to continuous data that are not normally distributed and that can be represented by mathematical functions. For cases in which the data cannot be fitted, however, and therefore are not suited for the FBRT procedure, we developed a method combining a point-by-point variance estimate with a resampling technique in order to calculate confidence bands according to the FBRT. In addition, the new method includes the calculation of a C factor that integrates the data of all samples. We call this method the point-based resampling technique (PBRT); and it will be explained in the following sections. For illustration purposes, we will demonstrate the application of this method using exemplary EEG data from an event-related potential (ERP) analysis. Although we demonstrate the PBRT in a between-groups comparison, this procedure can also be used with other continuous data and different experimental designs (e.g., within-subjects comparisons, as is described in Appendix 3).
Method
Exemplary data
For our exemplary data, we recorded EEG signals while participants practiced a virtual throwing task. The specific EEG potential of interest was the so-called error-related negativity (ERN, Ne), which is characterized as a negative deflection that appears shortly after the onset of an erroneous motor response to an imperative stimulus. The measured activation voltage of the ERN is significantly more negative than the signal from correct motor responses in the same temporal interval (Falkenstein, Hohnsbein, Hoormann, & Blanke, 1991; Gehring, Goss, Coles, Meyer, & Donchin, 1993).
During data acquisition, we measured ten healthy participants. The participants’ task was to throw a virtual ball presented on a projection screen around an obstacle in order to hit a target object on the other side of the obstacle (for details, see Müller & Sternad, 2004). They were instructed to hit the target as frequently as possible. EEG data were collected while participants executed the task. In the following sections, we will describe how the PBRT was applied to these exemplary data in order to test whether there was a difference in neural activation between hit \( {DS}_{hit}=\left\{{Y}_{(1..n)}^1,\dots, {Y}_{(1..n)}^k\right\} \) and miss \( {DS}_{miss}=\left\{{X}_{(1..n)}^1,\dots, {X}_{(1..n)}^k\right\} \) trials. Further information about the data acquisition and processing can be found in Appendix 1.
Selecting an appropriate analysis method
To decide which methods are most appropriate for the analysis of a given data set in the context of a particular hypothesis, particular limitations and benefits of the methods need to be discussed. For our EEG example, local methods could be excluded from further consideration since the inner dependency ρ was neither 0 nor 1 and thus could not be considered adequately. When ρ < 1, a reduction of the continuous data (neural activation over time) to one single representative value (e.g., a peak or mean amplitude) goes along with a loss of information and is therefore limited. On the other hand, EEG data are also not completely independent from index to index (|ρ| > 0). Hence, rendering Bonferroni corrections is inadequate. The methods that fully integrate all data points adequately would be the FBRT and the PBRT. The application of the FBRT, however, challenges us with finding a function prototype whose parameters could be adjusted in order to represent each of the curves in the data set. Since every data curve can be represented by a sum of sine wave terms by applying a Fourier transform, one might think that this should solve the problem. However, this is not the case since the coefficients resulting from Fourier transforms of the different EEG curves cannot reasonably be used to estimate the variance within the data set. This can be demonstrated by applying Fourier transforms sequentially to all \( {Y}_{(1..n)}^k \) within \( {DS}_{hit}=\left\{{Y}_{(1..n)}^1,\dots, {Y}_{(1..n)}^k\right\} \). An example highlighting this issue is shown in Appendix 2. In this case and a wide range of comparable situations, no valid fitting function can be found, and hence the FBRT cannot be applied to the data set. The PBRT, however, is still applicable. We will describe the implementation of the PBRT in the following sections by using our exemplary EEG data. A more detailed and formalized description can be found in Appendix 3 and in the MATLAB script in the supplementary material.
Application of the point-based resampling technique (PBRT)
Preparation of the data before analysis
First, we synchronized the data, in our case, to the respective moments of ball release. In the next step, data from hits and misses were baseline-corrected and cut into segments of equal length (i.e., all resulting segments containing an equal amount of data points/indices). Two average curves were then calculated for each data set, yielding \( {\overline{DS}}_{\left(1\dots n\right)}^{hit} \) and \( {\overline{DS}}_{\left(1\dots n\right)}^{miss} \). Next, the temporal location and width of the effect window was determined on the basis of preliminary findings (e.g., Joch, Hegele, Maurer, Müller, & Maurer, 2017; Maurer, Maurer, & Müller, 2015). In our example, we set the effect window from 200 to 350 ms after ball release.
Construction of confidence bands using the PBRT
In our example, the aim was to test whether the average curve of miss trials \( {\overline{DS}}_{\left(1\dots n\right)}^{miss} \) was different from the average curve of hit trials \( {\overline{DS}}_{\left(1\dots n\right)}^{hit} \) within the effect window. To this end, a confidence band for the data points within the effect window was constructed on the basis of the data set that represents the null hypothesis (i.e., \( {DS}_{hit}=\left({Y}_{(1..n)}^1,\dots, {Y}_{(1..n)}^k\right) \)). To provide evidence that the effect is present on the miss trials, \( {\overline{DS}}_{\left(1\dots n\right)}^{miss} \) should fall outside the confidence band (which is constructed on the basis of the hit trial data) within the effect window.
Construction of the confidence bands involved two steps: (1) estimation of the local distribution parameters and (2) determination of a factor C to correct for the inner dependencies in the curves.
-
Step 1 (Estimation of local distribution): The calculation of the averageFootnote 2 curves (e.g., \( {\overline{DS}}_{\left(1\dots n\right)}^{hit} \) from \( {DS}_{hit}=\left({Y}_{(1..n)}^1,\dots, {Y}_{(1..n)}^k\right) \) ) resulted in only one average curve for each of the conditions (hit and miss trials, respectively). To estimate the distribution parameters of these average curves \( \left({SE}_{{\overline{DS}}_{\left(1\dots n\right)}^{hit}}\right) \), we simulated pseudosamples using the bootstrap resampling method described by Efron (1979), yielding a sample containing pseudo-average curves. For each of the 900 data points (i.e., the data points within the effect window), standard deviation was then computed across all pseudo-average curves.
Before applying Step 1, the researcher has to determine the number of required bootstrap samples and need to decide what measure of central tendency is adequate for the analysis. The ideal number of bootstrap samples is mm, where m represents the number of data curves available for analyzing (m = 10). The size of this ideal bootstrap sample becomes a computational issue when m is large. However, a simulation calculating the reliability of the estimates can help to choose an appropriate amount of bootstrap samples. In our example, the standard deviation of the estimates drops proportionally to the square root of the number of runs (see Fig. 3). We, therefore, stopped resampling after 400 runs where the variability of estimations of the correction factor C dropped below 5%. Alternatively, the number of bootstrap samples can be chosen without a simulation by calculating as many samples as reasonably possible.
Development of the estimation variability of the scaling factor C with an increasing number of bootstrap samples (BSs). The coefficient of variation drops from almost 30% when using only ten BSs to under 3% when using 1,600 BSs. Such simulations can be used to determine the minimum number of BSs when using the bootstrap resampling procedure
In many single-point analyses, the central tendency of the data set is represented by the mean. Please note that in some cases the mean is not a robust measure of central tendency. Alternative estimators can be, for example, the median or the trimmed mean (Wilcox, 2012). Which of the measures of central tendency best fits the data at hand will depend on the type of the distribution to be analyzed. The PBRT works with any of these measures. To demonstrate the influence of different measures of central tendency when using the PBRT, we calculated the α and β error rates while applying the PBRT using the mean, median, and trimmed mean of artificially generated EEG data (please see Appendix 4 for details about the artificially generated EEG data). The outcome of this parameter comparison can be found in the Results section.
-
Step 2 (Calculation of scaling factor C): Following the usual logic, the confidence limits (CIs) are set at a specific radius from the mean in order to exclude the intended percentage of cases. In the case of normally distributed data, the limits would be set to the mean plus/minus a given factor times the standard deviation (CI = ± C ⋅ std), with this factor usually being C = 1.96, in order to include 95% of the cases. In addition, subsequent analyses would also require the scaling factor to be corrected according to the amount of the inner dependency ρ of the curves, as we mentioned earlier. Since it is very difficult to quantify ρ appropriately, the necessary correction factor cannot be determined directly. We therefore used the method introduced by Lenhoff et al. (1999). This method is based on the distribution of critical distances dcrit. For testing our hypothesis, the crucial concern was whether at least a single index exists for which the curve in question, \( {\overline{DS}}_{\left(1\dots n\right)}^{miss} \), would lie outside the confidence band. In general, a curve Y(1. . n) is inside the confidence band if the distance to the mean \( {d}_j=\left|{y}_j-\overline{y_j}\right| \) is smaller than C ⋅ stdj for every j = 1…n —that is, \( {d}_{crit}=\underset{j}{\max}\left(\frac{d_j}{std_j}\right)<C \). In our case, we calculated dcrit values for the instances from the resampling runs within the effect window according to the procedure of Lenhoff et al. (1999). A C value was then selected that cut off 5% of the distribution of the dcrit values.
This was done for the effect window only. The scaling factor is used to regulate the location of the upper and lower limits of the bands (see Appendix 3). Furthermore, this factor defines which multiple of the standard errors is added to or subtracted from the population mean so that the upper and lower endpoints of the confidence band will meet the desired coverage probability.
Once the confidence band for the null hypothesis (\( {DS}_{hit}=\left\{{Y}_{(1..n)}^1,\dots, {Y}_{(1..n)}^k\right\} \), in our case) is constructed, the average curve of the data set representing the alternative hypothesis can be tested within this band. Note that the confidence band was constructed so that if only a single data point at any index within the effect window lay outside the confidence band, a significant difference between the tested data sets would be suggested. The result of the test in our example can be found in the Results section.
Post-hoc analyses
Validation of the calculated C value
After the confidence band was calculated, the band had to be tested in terms of its true prediction coverage. A validation of the confidence band can be achieved by using cross-validation techniques. To do so, the average curves of the m bootstrap samples were used. Starting with m1, one curve was removed from the sample and a confidence band was computed with the remaining curves. In each run, we checked whether the removed curve stayed within the statistical limits over all data points 1 . . . n. This procedure was repeated for every value of m. In our case, the procedure was done 400 times. The relative number of cases in which the removed curve falls outside of the confidence band should be close to the desired α error probability. To emphasize the advantage of the PBRT over the local Gaussian method, we calculated the true prediction coverages of both methods for data sets with different distributions (varying the skewness) using the artificially generated EEG data (see Appendix 4). The results of this calculation can be found in the Results section and are illustrated in Fig. 6.
Power analysis
An important aspect of data analysis, aside from hypothesis testing, is the verification of the statistical test itself. Using the local methods (see Fig. 2), it is possible to quantify the power of an applied test in a post hoc analysis. Furthermore, a priori calculations of required sample sizes based on given effects can be done. However, these analyses are not limited to the single-point data case. We now present a way to validate the computed confidence bands and apply a power analysis to continuous data as well.
Our application of a power analysis method in the context of analyzing continuous data is based on effects from previous findings and simulated test curves for power analysis of the computed confidence band. Concretely, we used the effect (voltage differences between hit and miss trials) within the effect window found in a previous study (Maurer et al., 2015) and added those values to each of the 400 resampled average hit curves, which represent our null hypothesis. In other words, we constructed artificial error curves (AECs) from the hit curves. The power of the confidence band analysis was quantified by testing how many of the AECs fell outside the confidence band, which had been constructed on the basis of the hit curves. The number of AECs that stayed inside the confidence band of the hit curves throughout the complete effect window would be categorized incorrectly—that is, a curve carrying the effect would be judged as showing no effect. The relative frequency of this type of error would represent the ß error probability. Conversely, the relative frequency of AECs leaving the confidence limits for at least one index could be taken as the power (1–ß). For better illustration, the procedure of the power analysis is presented as a schematic diagram in Fig. 4. A detailed description of the calculation of the power analysis can also be found in Appendix 3.
Schematic illustration of the confidence band (CB) power analysis. The power analysis starts with the resampled data (A). The effect found in previous experiments (B; i.e., the effect within effect window) is then added to the data supporting the null hypothesis (Α) to produce artificial error curves (AECs). (C) Result of the effect addition A + B = AEC. (D) CBs are computed using the resampled data shown in A. (E) Tests of the AECs (C) within the CB (D), calculated from the resampled data. The relative frequency of AECs that do not leave the CB at any index within the effect window can be used as an estimate of β error probability
Results
Testing for differences with confidence bands
The aim in our example was to test two EEG data sets for differences in the means within a preset effect window. One data set included the hit trials \( {DS}_{hit}=\left\{{Y}_{(1..n)}^1,\dots, {Y}_{(1..n)}^k\right\} \) and the other included the miss trials \( {DS}_{miss}=\left\{{X}_{(1..n)}^1,\dots, {X}_{(1..n)}^k\right\} \) acquired from a goal-oriented throwing task. Therefore, the null hypothesis would be formulated as \( {H}_0:{\overline{DS}}_{\left(1\dots n\right)}^{hit}={\overline{DS}}_{\left(1\dots n\right)}^{miss} \). (H1 : ¬ H0). For significance testing, a confidence band was calculated using the data set that was supposed to show no effect in the EEG data (DShit). The test for differences in the mean curves was done by placing the average curve of the data set in which an effect was expected (\( {\overline{DS}}_{\left(1\dots n\right)}^{miss} \)) within the confidence band around \( {\overline{DS}}_{\left(1\dots n\right)}^{hit} \). If \( {\overline{DS}}_{\left(1\dots n\right)}^{miss} \) were to leave the band for at least one single index, a significant difference between \( {\overline{DS}}_{\left(1\dots n\right)}^{hit} \) and \( {\overline{DS}}_{\left(1\dots n\right)}^{miss} \) would be indicated at all indices leaving the band.
In the example plotted in Fig. 5, the average curve of the miss trials leaves the confidence band from about 250 until 295 ms. Hence, a significant mean difference was found within the effect window and we would reject H0 and accept H1. With respect to the exemplary research question, we found a significant negative deflection in the miss trials—that is an error-related negativity.
Comparison of different measures of central tendency within the PBRT
As already mentioned in the description of the PBRT (Step 1: Estimation of local distribution), the method can be used with different measures of central tendency. Wilcox (2012) discusses in his book on robust estimation and hypothesis testing that prominent measures such as the mean, median, and trimmed mean can be more or less robust depending on the type of data distribution (e.g., the level of skewness). Thus, we tested the influence of choosing any of the three different measures on the actual prediction coverage of the resulting confidence bands (where the desired nominal level was set to α = 5%) when increasing the level of skewness in our artificial data curves (please see Appendix 4 for details about the artificial data).
As can be seen in Fig. 6, the increase in skewness does not lead to relevant deviations from the desired nominal level for any of the tested measures of central tendency integrated in the PBRT. Nevertheless, it is indispensable to individually decide what measure fits the data at hand best in order to use this measure with the PBRT.
Comparison of three measures of central tendency in terms of the α error probability of the confidence band with increasing levels of skewness. All tested measures (mean, median, and trimmed mean 20%) using the PBRT seem to be robust to different levels of skewness and deliver the desired α error probability of 5%. The α error probability using the local Gaussian method is clearly different from the desired nominal level and increases slightly with an increasing level of skewness, from 27% to 34%. The distributions below the x-axis visualize the skewness of the analyzed data
Discussion
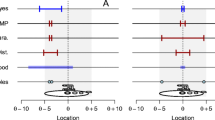
In this article, we have discussed differences between single and continuous data regarding data analysis. We compared different statistical methods with respect to their applicability to continuous data and introduced a method based on bootstrap resampling. An overview in Fig. 7 helps to visualize the key arguments for or against the methods discussed in this article.
Local methods are designed to analyze data sets composed of k single data points (Fig. 7, column A; e.g., reaction times of k participants). These techniques can be adapted to the analysis of continuous data by treating the data at each index j = 1…n as a single data point and running n single-data analyses in parallel. However, this procedure is only accurate for data with an internal dependence ρ close to either 1 or 0 (using a Bonferroni correction in the latter case) because other ρ values cannot be modeled adequately within the procedure (Fig. 7, column A4). Thus, using local methods on continuous data can lead to extremely progressive (assuming ρ = 1) or conservative (assuming ρ = 0) results. This is supported by the observation that the true prediction coverage in the analysis of continuous EEG data is far from the nominal level if we apply a local Gaussian method, which is a point-by-point (n-by-n) application of a local method to each index j = 1…n (Fig. 7, column B; αnominal = 5%, αtrue ranging from 27% to 34%, depending on the skewness of the analyzed distribution). This finding is in line with the results of Lenhoff et al. (1999) who compared the true coverage probability of their FBRT (Fig. 7, column C) to the local Gaussian method and came to a similar result. On the basis of these findings, we do not recommend using either local methods or local Gaussian methods for the analysis of continuous data unless (1) a normal distribution of every data point can be assured and (2) ρ is either 0 or 1.
In contrast to this, we described the FBRT as a tool that includes mathematical fits and a scaling factor for the confidence bands and that is appropriate to analyze effects represented by more than one data point (i.e., continuous data; Fig. 7, column C1, C4). The major benefit of this technique results from the scaling factor C used to adjust the prediction coverage so that it closely approaches the desired nominal level, which was confirmed by a cross-validation in the study by Lenhoff et al. (1999). Using mathematical functions to fit the data, and thus to model the inner dependency represented by the coefficients of the fitting function, has one advantage as compared to bootstrap resampling techniques without functional fits. Such functions usually require less computational effort because the resampling is only done with the function coefficients and not with all data points j ∈ {1, .., n}, thus decreasing the necessary computational effort substantially (Fig. 7, column C5). In the case that the functions fit the data sufficiently well, the resulting confidence band will only be marginally influenced by the fitting error, which will not substantially change the researcher’s decision in the hypothesis testing. However, the mathematical fitting of the data is also a strong limitation of the FBRT, as this procedure cannot be applied to a wide range of continuous data sets (e.g., high-frequency data such as EEG signals) as it is not possible to find a family of functions appropriate to fit the data curves reasonably well. In most cases, a function can be found to fit one of the k data curves in a data set. This function, however, does often not sufficiently represent the characteristics of the other curves in the data set, which makes it unfeasible to apply the FBRT in these cases (for an example of this problem, see Appendix 2).
As an alternative to the existing methods, we introduced a nonparametric approach to analyzing continuous data by combining some aspects of the FBRT with the point-by-point analysis of local methods (we call this a point-based resampling technique, PBRT; Fig. 7, column D). Integrating bootstrap resampling allowed us to statistically analyze two average curves (which is often needed when, e.g., analyzing ERPs in EEG or comparing two population mean curves in general) through the use of confidence bands. This was done by simulating multiple pseudo-average curves on the basis of real, nonfitted data in order to compute a simultaneous confidence band, which can then be used to estimate the distribution of the average curves. For demonstration purposes, we examined the EEG signals of ten subjects in order to test for differences between two EEG time series (hit vs. miss trials in a throwing task)—more specifically, with respect to the appearance of an ERP that correlates with erroneous motor executions. After calculating the confidence band from the data set representing the null hypothesis (hit trials) for the indices within the preset effect window, we tested the average curve of the miss trials within the confidence band (see Fig. 5). We found that the average curve of the miss trials left the confidence band within the effect window, which indicated significant differences between hit and miss trials within this time interval. Hence, the results of the PBRT suggested rejecting the null hypothesis in this example.
In further simulations we showed that the true prediction coverage of the PBRT was very close to the desired nominal level of αnominal = 5% (5.3% ≤ αtrue≤ 5.8%). Furthermore, we could show that this result was robust to the degree of skewness in the analyzed data distribution. With respect to robust estimators, we tested the influences of different measures of central tendency on the prediction coverage of the confidence band and found no relevant differences when using the mean, median, and trimmed mean (20%; see also Fig. 6), at least in our example. However, the influences of different measures of central tendency can change when analyzing different data sets.
To conclude, the PBRT can be applied to nonnormally distributed continuous data that cannot be fitted by a basic mathematical function. In addition, our method provides hypothesis testing in specific time windows (effect windows). Nevertheless, the definition of an effect window—that is its location and size—is only reasonable when the onset of the hypothesized effect is clearly known. The generated confidence band can also be used for a power analysis, as we described in the Method section. In fact, it can also be used as a tool to specify required sample sizes a priori. Thus, we recommend this specific confidence band analysis as an alternative to the method of Lenhoff et al. (1999) when analyzing continuous behavioral data curves that cannot be fitted by an appropriate mathematical function family (e.g., with EEG data).
Although the introduced PBRT can be applied to many types of continuous data, individual decisions must be made before every analysis: first, the appropriate estimators for the data to be analyzed; second, the minimum number of iterations to be used for the bootstrap subsamples. Our calculations show that accuracy improves to asymptotically approximate zero error with an increasing number of subsamples. However, it should be noted that the decision of which confidence band accuracy will be sufficient will heavily depend on the particular data set and the hypothesized effect. The accuracy of the resulting confidence limits can be roughly estimated from multiple calculation runs; but it is still up to the experimenter to decide what level of confidence band accuracy is enough. Finally, the selection of adequate estimation parameters, the location and size of the effect window, and the experimental design itself will all depend on informed decisions by the researcher. Note that these requirements are not a particular disadvantage of the methods described here since such decisions are required in any type of statistical testing.
Notes
Continuous data are not “continuous“ in a strict sense. The data we discuss consist of n discrete values which together form a continuous-like signal.
Note that this step is not restricted to the mean; other measures of central tendency can be applied as well.
References
Efron, B. (1979). Bootstrap methods: Another look at the jackknife. Annals of Statistics, 7, 1–26. https://doi.org/10.1214/aos/1176344552
Falkenstein, M., Hohnsbein, J., Hoormann, J., & Blanke, L. (1991). Effects of crossmodal divided attention on late ERP components: II. Error processing in choice reaction tasks. Electroencephalography and Clinical Neurophysiology, 78, 447–455. https://doi.org/10.1016/0013-4694(91)90062-9
Gehring, W. J., Goss, B., Coles, M. G. H., Meyer, D. E., & Donchin, E. (1993). A neural system for error detection and compensation. Psychological Science, 4, 385–390. https://doi.org/10.1111/j.1467-9280.1993.tb00586.x
Joch, M., Hegele, M., Maurer, H., Müller, H., & Maurer, L. K. (2017). Brain negativity as an indicator of predictive error processing : The contribution of visual action effect monitoring. Journal of Neurophysiology, 18, 486–495. https://doi.org/10.1152/jn.00036.2017
Lenhoff, M. W., Santner, T. J., Otis, J. C., Peterson, M. G. E., Williams, B. J., & Backus, S. I. (1999). Bootstrap prediction and confidence bands: A superior statistical method for analysis of gait data. Gait and Posture, 9, 10–17. https://doi.org/10.1016/S0966-6362(98)00043-5
Makeig, S. (1993). Auditory event-related dynamics of the EEG spectrum and effects of exposure to tones. Electroencephalography and Clinical Neurophysiology, 86, 283–293. https://doi.org/10.1016/0013-4694(93)90110-H
Makeig, S., Bell, A. J., Jung, T.-P., & Sejnowski, T. J. (1996). Independent component analysis of electroencephalographic data. In D. Touretzky, M. Moser, & M. Hasselmo (Eds.), Advances in neural information processing systems (Vol. 8, pp. 145–151). Cambridge, MA: MIT Press. https://doi.org/10.1109/ICOSP.2002.1180091
Makeig, S., Jung, T. P., Bell, A. J., Ghahremani, D., & Sejnowski, T. J. (1997). Blind separation of auditory event-related brain responses into independent components. Proceedings of the National Academy of Sciences, 94, 10979–10984.
Maurer, L. K., Maurer, H., & Müller, H. (2015). Neural correlates of error prediction in a complex motor task. Frontiers in Behavioral Neuroscience, 9, 209:1–8. https://doi.org/10.3389/fnbeh.2015.00209
Müller, H., & Sternad, D. (2004). Decomposition of variability in the execution of goal-oriented tasks: Three components of skill improvement. Journal of Experimental Psychology: Human Perception and Performance, 30, 212–233. https://doi.org/10.1037/0096-1523.30.1.212
Tukey, J. W. (1958). Bias and confidence in not-quite large samples. Annals of Mathematical Statistics, 29, 614. https://doi.org/10.2307/2237363
Wilcox, R. R. (2012). Introduction to robust estimation and hypothesis testing (3rd ed.). Amsterdam, The Netherlands: Academic Press. Retrieved from http://linkinghub.elsevier.com/retrieve/pii/B9780123869838000019
Author note
We thank Heiko Maurer for his methodological and mathematical advice. This research was funded by the Deutsche Forschungsgemeinschaft via the Collaborative Research Center on “Cardinal Mechanisms of Perception” (SFB-TRR 135) and the research projects MU 1374/3-1 and MU 1374 /5-1.
Author information
Authors and Affiliations
Corresponding author
Appendices
Appendix 1: EEG recording and data preprocessing
EEG was recorded using a 16-channel AC/DC amplifier with Ag/AgCl active scalp electrodes (V-Amp, Brain Products). EEG electrodes were placed with an electrode cap (actiCap, Brain Products) according to the international 10–20 system on F3, Fz, F4, FCz, C3, Cz, C4, P3, Pz, and P4. An electrooculogram (EOG) was recorded with electrodes on the external canthi of both eyes as well as above and below the right eye. All electrodes were referenced online to the left mastoid and re-referenced after data collection to the averaged mastoids. The electrode impedance was kept below 20,000,Ω which is appropriate for active electrodes. The data were digitized at a sampling rate of 500 Hz. The EEG and EOG data were filtered online with a high-pass filter at 0.01 Hz. EEG data were additionally filtered offline with a phase-shift free Butterworth filter (0.2–30 Hz bandpass). Ocular artifacts were removed from the continuous EEG signal using infomax independent component analysis (Makeig, Bell, Jung, & Sejnowski, 1996; Makeig, Jung, Bell, Ghahremani, & Sejnowski, 1997) and were segmented into epochs around the release, beginning 600 ms before release and ending 1,200 ms after release. The segments were baseline-corrected by subtracting the average amplitude of the complete signal. Single segments were then visually inspected to remove epochs that contained artifacts.
Appendix 2: The issue of estimating variance from function coefficients
To illustrate why it can be problematic to find a general function prototype for certain types of continuous data (such as EEG), we fit two example data curves from our EEG data set using a sum of sines model, defined as
where a represents the amplitude, b the frequency, and c the phase for each sine wave term. We calculated a sum of the eight sine wave terms for each of the two EEG curves in order to yield a reliable fit. The coefficients of the eight terms a1–8, b1–8, and c1–8 are shown in Table 1. The rationale for fitting the data with mathematical functions in the FBRT is that the coefficients of the fitting functions are used to estimate the variance within the data. Thus, the next step in our example will be to identify sine wave terms in Curve 1 that correspond with the sine wave terms of Curve 2. In other words, the terms have to be sorted so that corresponding terms represent the same feature of a curve. This correspondence between the sine wave terms of the different curves is essential for the estimation of the variance. To find a reasonable order, let us look at the frequency component (b) of each term in Curve 1 (C1) and try to find a corresponding frequency coefficient in Curve 2 (C2). The sorting could also be based on the ai or ci coefficients, but focusing on bi is sufficient to demonstrate the issue. Starting with C1(b1) = 0.006, an obvious corresponding frequency coefficient of C2 would be C2(b1). Hence, we match the first sine wave term of C1 to the first of C2. Next, C1(b2) = 0.095. The best match would be C2(b7) = 0.115. However, the frequencies described in C1(b2) and C2(b7) already differ by 18%. Proceeding, C1(b3) = 0.057 matches best to C2(b5), but C2(b5) would also be the only good match for C1(b4). Choosing C2(b4) as the next best match for C1(b4) highlights the problem of finding corresponding terms in the curves since the difference in the frequencies included in C1(b4) and C2(b4) is already 47.5%. Please note that this example is constructed using two curves. The problem of finding the corresponding terms in the other curves of the data set becomes increasingly difficult as the sample size becomes larger.
A variation of this fitting method was used by Lenhoff et al. (1999). They also used a sum of sine terms to fit their data, but they defined fixed frequencies to the terms. When fixing the frequency components (bi) of the sine wave terms, it becomes clear which terms in multiple curves are corresponding. We tried to apply this logic to our EEG data, but we identified two problems that prevented us from generating valid fits. First, the chosen fixed frequencies did not fully reflect the frequency components of the original data curves. Concretely, it was not possible to choose the exact frequencies that the original curve was composed of. Even a Fourier transform cannot solve this problem due to its resolution constraints. This frequency offset would lead to less accurate fits. Second, and more importantly, the limits of functions as representations of data curves become apparent when a “singular” (i.e., unique and temporarily limited) event, such as an effect (e.g., an ERP component in the EEG), is manifested within the data curve. The frequency components that might be able to fit the general course of the data will not be sufficient to adequately fit the effect. In consequence, the effect is likely to be reduced, and therefore the construction of confidence bands based on these fits is not recommended.
Appendix 3: Calculation of confidence bands using the PBRT
The data used for processing must be synchronized and, if necessary, normalized in time so that the temporal correspondence between data points is assured across curves. It is also important that every data curve has the same length (n = number of values) within the effect window (WEff). The WEff is the time interval in which an effect is expected to occur, and its length can be defined on the basis of preliminary data or from previous findings in the literature.
The PBRT is constructed to compare two data sets (DS) of continuous data—for example, \( {DS}_A=\left\{{Y}_{\left(1\dots n\right)}^1,\dots, {Y}_{\left(1\dots n\right)}^k\right\} \) and \( {DS}_B=\left\{{X}_{\left(1\dots n\right)}^1,\dots, {X}_{\left(1\dots n\right)}^k\right\} \), where k is the number of continuous data curves in a data set. Please note that the sample sizes may differ in DSA and DSB in between-subjects comparisons. However, to simplify the notation we have used the same k for both samples.
In our EEG example, DSA stands for the hit trials and DSB represents the miss trials. To be able to analyze the difference between \( {\overline{DS}}_{\left(1\dots n\right)}^A \) and \( {\overline{DS}}_{\left(1\dots n\right)}^B \) in a preset effect window WEff [1, …, n], a confidence band was calculated for the data set representing the null hypothesis. In the example of our EEG data, it is reasonable to assume that the variances of the data in DSA and DSB might differ substantially (see Makeig, 1993). Therefore, in our case, the calculation of the confidence band was based on the data included in DSA because it represents our null condition (i.e., hit trials). The calculation of a confidence band given similar variances in both data sets is shown at the end of this technical description. The continuous data curves in DSA were resampled with replacement in order to yield m pseudo-bootstrap samples (BS) \( {BS}_A^1,\dots, {BS}_A^m \), each of which contained k randomly chosen data curves.
To estimate what number of BSs would be sufficient, we calculated the development of the variability in the estimation of C for repeated runs of the bootstrap algorithm with different numbers of BSs (m). Figure 3 shows that the estimation variability decreases with an increasing number m of BSs. On the basis of this variability, we chose to calculate m = 400 BSs in our example because the estimation variability dropped below 5% of the magnitude of C at this point. This shows that a reasonable accuracy can be achieved with a manageable number of BSs. However, the actual number of BSs for a specific future study must be chosen individually, depending on the type of data and the precision requirements.
To compute confidence bands based on the distribution of the BSs, average curves were calculated by averaging the k curves for every \( {BS}_A^1,\dots, {BS}_A^m \), yielding m curves \( {\overline{BS}}_{A,\dots,}^1{\overline{BS}}_A^m \). For data within the effect window WEff (indices 1, …, n), means of \( {\overline{BS}}_{A,\dots,}^1{\overline{BS}}_A^m \) (\( {\mu}_{\overline{BS}} \), Eq. 1) and standard errors (SE, Eq. 2) of the m average curves were needed for further processing.
To reach a desired confidence level (1 − α), where α is the family-wise error rate, it is necessary to determine a constant scaling factor C to calculate the lower (L–) and upper (L+) limits of the confidence band using Eq. 3.
The final scaling factor C is chosen so that the relative frequency p, with which the single curves of a pseudosample stayed within the upper and lower limits of the confidence band throughout the complete effect window, was p = 1 − α (4).
An appropriate constant C can be determined by calculating the critical distances dcrit (maximum relativized distances) for every BS (1,…, m) between \( {\mu}_{\overline{BS}}(n) \) and the average curves \( {\overline{BS}}_{A,\dots,}^1{\overline{BS}}_A^m \). The (1 – α) percentile of the resulting dcrit distribution meets the criterion formulated in Eq. 4.
Calculation for data sets with similar variances
If similar variances can be assumed for both data sets (DSA and DSB) to be analyzed, it is reasonable to estimate the variation of the data on the basis of both data sets to yield a more reliable estimation. In the example above, the scaling factor C was calculated by using data from DSA only. To adapt the PBRT to similar variances in both data sets, an adjusted C factor can be calculated as follows:
In addition to the C factor that was calculated for DSA (i.e., CA), a second C is calculated using the data from DSB (i.e., CB), with the same procedure that was used to yield CA. Subsequently, CA and CB are averaged to yield \( \overline{C} \), as is shown in Eq. 5.
If the sample sizes in DSA and DSB differ, a weighted average should be used. Finally, the confidence limits at each index are calculated as shown in Eq. 6.
Within-subjects comparisons using the PBRT
In accordance with the logic of the dependent-samples t test, within-subjects comparisons can be conducted by testing differences between the conditions against zero. For instance, given the data \( {DS}_A=\left\{{Y}_{\left(1\dots n\right),}^1\dots {Y}_{\left(1\dots n\right)}^k\right\} \) from a pretest and \( {DS}_B=\left\{{X}_{\left(1\dots n\right),}^1\dots {X}_{\left(1\dots n\right)}^k\right\} \) representing the posttest data for the same participants, the alternative hypothesis would be formalized as H1 : (DSA − DSB) ≠ 0, and the null hypothesis as H0 : ¬ H1. In style of the dependent-samples t test, a confidence band would be generated on the basis of the difference data by applying the PBRT procedure as we have described it above. The resulting confidence band can subsequently be used for hypothesis testing. If the value of zero falls outside the confidence band for at least a single index, a signficant difference between pre- and posttest would be suggested at all indices leaving the band.
Power analysis
Since power describes the expressiveness of a statistical test with the help of the β error, it is necessary to quantify β.
For the estimation of β a known effect is required. For instance, when doing a power analysis for confidence bands, the effect must be represented by an effect curve (e.g., the curve that can be seen in Fig. 4B).
The first step is to add the postulated effect to each of a set of m data curves, which represent the null hypothesis. As a result, m artificial effect curves (AEC) are constructed. The data curves that represent the null hypothesis are then used in Step 2 to compute a confidence band (CB) as shown above. The relative number of AECs that do not leave the CB within the effect window can be taken as an estimate of the probability of a β (Type II) error. The power is then calculated as is shown in Eq. 7.
Appendix 4: Generation of the artificial data curves for simulation
To test the influence of different measures of central tendency when using the PBRT, we generated artificial data curves that had characteristics similar to those of our real data curves. To create these artificial EEG-like curves, we used a sinusoidal function with 24 coefficients to replicate the main characteristics of the original hit curves. The fit was applied to the mean curve and was also detrended, to avoid fitting any trend present in the original data. This fit was analogously labeled as the mother curve and was used as source for all our artificial curves. In the process of creating individual curves, different trends were readded and noise was applied to the mother curve in order to gain realistic patterns. The noise consisted of variability in the coefficients of the function and of local random noise added to each data point. In the last step, we added the effect found in a preliminary study to half of the curves in order to generate artificial error curves. We validated our created data curves in two ways. First, we examined the artificial data visually and subjectively compared them to the original data. Second, we compared the variability and the autocorrelation profiles of the original and artificial curves. The results of both these subjective and objective validations can be seen in Fig. 8.
Comparison between the original EEG data curves and the artificially generated data curves used for the simulations. (A) Original data curves recorded during data acquisition. (B) Artificial data curves. Comparisons are based on such objective parameters as the autocorrelation profile (C) and the standard deviation profile (D). In all diagrams, the original data are printed in black, and the artificially generated data are colored gray
Rights and permissions
About this article
Cite this article
Joch, M., Döhring, F.R., Maurer, L.K. et al. Inference statistical analysis of continuous data based on confidence bands—Traditional and new approaches. Behav Res 51, 1244–1257 (2019). https://doi.org/10.3758/s13428-018-1060-5
Published:
Issue Date:
DOI: https://doi.org/10.3758/s13428-018-1060-5