Abstract
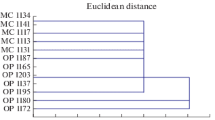
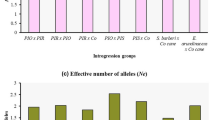
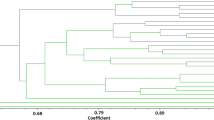
The goal of this study was a molecular genetic passportization of the initial breeding material of sugar beet (male sterile lines, synanthous pollinators, and their hybrids). The following SSR primers were used: Unigene 24 552, Unigene 2305, Unigene 17 623, Unigene 14 805, and Unigene 62 524. The length range of the obtained DNA fragments varied within 100–3000 bp, and up to 11 polymorphic bands per genotype were amplified. The maximum Polymorphic Information Content (PIC) value was revealed for the Unigene 17 623 (PIC = 0.88), Unigene 2305 (PIC = 0.84), and Unigene 14 805 (PIC = 0.85) loci that provided a possibility to differentiate sugar beet breeding material. Using five of the tested SSR markers, the molecular genetic passportization of 26 genotypes of valuable breeding samples of this crop was carried out that allowed their identification and certification for further use in a marker-assisted selection. The calculated genetic distances between male sterile forms and synanthous pollinators varied from 2.236 to 4.796. Parent samples located at considerable genetic distances from each other were recommended for use in creating heterotic hybrids.




Similar content being viewed by others
REFERENCES
Taški-Ajduković, K., Nagl, N., Ćurčić, Ž., and Zorić, M., Estimation of genetic diversity and relationship in sugar beet pollinators based on SSR markers, Electron. J. Biotechnol., 2017, vol. 27, pp. 1–7.
Abbasi, Z., Arzani, A., and Majidi, M., Evaluation of genetic diversity of sugar beet (Beta vulgaris L.) crossing parents using agro-morphological traits and molecular markers, J. Agric. Sci. Tech., 2014, vol. 16, pp. 1397–1411.
He, J., Zhao, X., Laroche, A., Lu, Z.-X., Liu, H., and Li, Z., Genotyping-by-sequencing (GBS), anultimate marker assisted selection (MAS) tool to accelerate plant breeding, Plant Genet. Genomics, 2014, vol. 5, pp. 1–7.
Holtgräwe, D., Rosleff, Th., Vieho, P., Schneider, J., Schulz, B., Borchardt, D., Kraft, Th., Himmelbauer, H., and Weisshaar, B., Polymorphisms and their application for extending the genetic map of sugar beet (Beta vulgaris), PLOS ONE, 2014, vol. 9, pp. 1–10.
Dohm, J.C., Minoche, A.E., Holtgrawe, D., Capella-Gutierrez, S., and Zakrzewski, F., The genome of the recently domesticated crop plant sugar beet (Beta vulgaris), Nature, 2014, vol. 505, pp. 546–549.
Shcherbina, O.M., Svirshchevskaya, A.M., and Kil’chevskii, A.V., Allelic diversity in microsatellite loci of diploid lines of sugar beet (Beta vulgaris L.), V s"ezd Vavilovskogo obshchestva genetikov i selektsionerov: Tezisy dokladov (V Congress of the Vavilov Society of Geneticists and Breeders: Abstracts), Moscow, 2009, part 1, p. 370.
Izzatullayeva, V., Akparov, Z., Babayeva, S., Ojaghi, J., and Abbasov, M., Efficiency of using RAPD and ISSR markers in evaluation of genetic diversity in sugar beet, Turk. J. Biol., 2014, vol. 38, pp. 429–438.
Curcic, Z., Taski-Ajdukovic, K., and Nagl, N., Relationship between hybrid performance and genetic variation in self-fertile and self-sterile sugar beet pollinators as estimated by SSR markers, Euphytica, 2017, vol. 213, no. 5, pp. 100–108.
Sandhu Surinder, K., Sarao Navraj, K., Meenakhsi, G., Uppal, S., Pritpal, S., Satveer, K., and Jaspreet, K., Profiling of sugar beet genotypes for agronomical, sugar quality and forage traits and their genetic diversity analysis using SSR markers, Electron. J. Plant Breed., 2016, vol. 7, pp. 253–266.
Broccanello, Ch., Chiodi, C., Funk, A., Mitchell McGrath, J., Panella Lee, and Stevanato, P., Comparison of three PCR-based assays for SNP genotyping in plants, Plant Methods, 2018, vol. 14, p. 28.
Simko, I., Eujayl, I., and van Hintum, T.J., Empirical evaluation of DArT, SNP, and SSR marker-systems for genotyping, clustering, and assigning sugar beet hybrid cultivars into populations, Plant Sci., 2012, vol. 184, pp. 54–62.
Srivastava, S., Pathak, A.D., Kumar, R., and Joshi, B.B., Genetic diversity of sugar beet genotypes evaluated by microsatellite DNA markers, J. Environ. Biol., 2017, vol. 38, pp. 777–783.
Mahuku, G.S., A simple extraction method suitable for PCR-based analysis of plant, fungal, and bacterial DNA, Plant Mol. Biol. Rep., 2004, vol. 22, pp. 71–81.
Hussein, A.S., Nalbandyan, A.A., Fedulova, T.P., and Bogacheva, N.N., Efficient and nontoxic DNA isolation method for PCR analysis, Russ. Agric. Sci., 2014, vol. 40, pp. 177–178.
Fugate, K., Fajardo, D., Schlautman, B., Ferrareze, J.P., Bolton, M.D., Campbell, L.G., Wiesman, E., and Zalapa, J., Generation and characterization of a sugarbeet transcriptome and transcript-based ssr markers, Plant Genome, 2014, vol. 7, no. 2, pp. 1–13.
Nei, M. and Roychoudhury, A.K., Sampling variances of heterozygosity and genetic distance, Genetics, 1974, vol. 76, pp. 379–390.
Nei, M. and Li, W.H., Mathematical model for studying genetic variation in terms of restriction endonucleases, Proc. Natl. Acad. Sci. U. S. A., 1979, vol. 76, pp. 5269–5273.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest. The authors declare that they have no conflict of interest.
Statement on the welfare of animals. This article does not contain any studies with animals performed by any of the authors.
Additional information
Translated by N. Statsyuk
About this article
Cite this article
Nalbandyan, A.A., Hussein, A.S., Fedulova, T.P. et al. Differentiation of Sugar Beet Varieties Using SSR Markers: A Tool to Create Promising Hybrids. Russ. Agricult. Sci. 46, 442–446 (2020). https://doi.org/10.3103/S1068367420050146
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.3103/S1068367420050146