Abstract
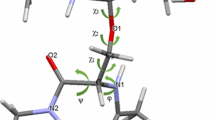
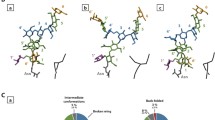
Analysis of the conformational variety of the oligosaccharide fragments of the human glycan receptors LSTa (α-D-Neu5Ac-(2-3)-β-D-Gal-(1-3)-β-D-GlcNAc-(1-3)-β-D-Gal(1-4)-D-Glc) and LSTc (α-D-Neu5Ac-(2-6)-β-D-Gal-(1-3)-β-D-GlcNAc-(1-3)-β-D-Gal(1-4)-D-Glc) in aqueous solution has been performed with the comprehensive use of molecular modeling and statistical data processing followed by determination of major and minor stabilized conformers and selection of relevant topologies. The sialic acid ring conformational free energy landscape for both pentasaccharides has been reconstructed and analyzed giving a specification of the most probable distorted ring conformations of the basic chair 1C4 structure. The obtained results are in a good agreement with experimental data generated by nuclear magnetic resonance spectroscopy and X-ray crystallography.
Similar content being viewed by others
References
Geissner, A. and Seeberger, P.H., Annu. Rev. Anal. Chem., 2016, vol. 9, p. 223.
Agre, P., Bertozzi, C., Bissell, M., Campbell, K.P., Cummings, R.D., Desai, U.R., and Ginsburg, D., J. Clin. Invest., 2016, vol. 126, no. 2, p. 405.
Domenech, M., Garcia, E., and Moscoso, M., Microb. Biotechnol., 2012, vol. 5, no. 4, p. 455.
Manco, S., Hernon, F., Yesilkaya, H., Paton, J.C., Andrew, P.W., and Kadioglu, A., Infect. Immun., 2006, vol. 74, no. 7, p. 4014.
Smith, D.F. and Cummings, R.D., Curr. Opin. Virol., 2014, vol. 7, p. 79.
Chandrasekaran, A., Srinivasan, A., Raman, R., Viswanathan, K., Raguram, S., Tumpey, T.M., and Sasisekharan, R., Nat. Biotechnol., 2008, vol. 26, no. 1, p. 107.
Case, D.A., Berryman, J.T., Betz, R.M., Cerutti, D.S., Cheatham, T.E., Darden, T.A., Duke, R.E., Giese, T.J., Smith, W., Swails, J., Walker, R.C., Wang, J., Wolf, R.M., Wu, X., York, D.M., and Kollman, P.A., AMBER 2015, San Francisco, 2015.
Bonomi, M., Branduardi, D., Bussi, G., Camilloni, C., Provasi, D., Raiteri, P., and Parrinello, M., Comput. Phys. Commun., 2009, vol. 180, no. 10, p. 1961.
Cremer, D. and Pople, J.A., J. Am. Chem. Soc., 1975, vol. 97, no. 6, p. 1354.
Iglesias-Fernández, J., Raich, L., Ardèvol, A., and Rovira, C., Chem. Sci., 2015, vol. 6, no. 2, p. 1167.
Costa, S.I., Torezzan, C., Campello, A., and Vaishampayan, V.A., IEEE Information Theory and Applications Workshop (ITA), 2013, p. 1.
Straub, J., Campbell, T., How, J.P., and Fisher, J.W., IEEE Conf. on Computer Vision and Pattern Recognition (CVPR), 2015, p. 334.
Pierce, L.C., Salomon-Ferrer, R., Augusto F. de Oliveira, C., McCammon, J.A., and Walker, R.C., J. Chem. Theory Comput., 2012, vol. 8, no. 9, p. 2997.
Laio, A. and Parrinello, M., Proc. Natl. Acad. Sci. U. S. A., 2002, vol. 99, no. 20, p. 12562.
Sassaki, G.L., Elli, S., Rudd, T.R., Macchi, E., Yates, E.A., Naggi, A., and Guerrini, M., Biochemistry, 2013, vol. 52, no. 41, p. 7217.
Sabesan, S., Bock, K., and Paulson, J.C., Carbohydr. Res., 1991, vol. 218, p. 27.
Poppe, L., Dabrowski, J., Lieth, C.W., Numata, M., and Ogawa, T., FEBS J., 1989, vol. 180, no. 2, p. 337.
Ohrui, H., Nishida, Y., Itoh, H., and Meguro, H., J. Org. Chem., 1991, vol. 56, no. 5, p. 1726.
Xu, D., Newhouse, E.I., Amaro, R.E., Pao, H.C., Cheng, L.S., Markwick, P.R., and Arzberger, P.W., J. Mol. Biol., 2009, vol. 387, no. 2, p. 465.
Thompson, A.J., Dabin, J., Iglesias-Fernandez, J., Ardevol, A., Dinev, Z., Williams, S.J., and Smith, D.K., Angew. Chem., Int. Ed., 2012, vol. 51, no. 44, p. 10997.
Chan, J., Lu, A., and Bennet, A.J., J. Am. Chem. Soc., 2011, vol. 133, no. 9, p. 2989.
Sadovnichy, V., Tikhonravov, A., Voevodin, V., and Opanasenko, V., “Lomonosov”: Supercomputing at Moscow State University, in Contemporary High Performance Computing: From Petascale toward Exascale, Boca Raton: CRC, 2013.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © E.M. Kirilin, V.K. Švedas, 2018, published in Vestnik Moskovskogo Universiteta, Seriya 2: Khimiya, 2018, No. 2, pp. 117–124.
About this article
Cite this article
Kirilin, E.M., Švedas, V.K. Study of the Conformational Variety of the Oligosaccharide Substrates of Neuraminidases from Pathogens using Molecular Modeling. Moscow Univ. Chem. Bull. 73, 39–45 (2018). https://doi.org/10.3103/S0027131418020050
Received:
Published:
Issue Date:
DOI: https://doi.org/10.3103/S0027131418020050