Abstract
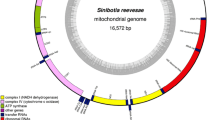
Cobitis macrostigma, a small-sized benthic fish species, is an endemic species distributing in the middle and lower reaches of the Yangtze River. In this study, the complete mitochondrial genome of C. macrostigma was first sequenced and the phylogenetic relationship with its closely related species was estimated. It was 16,573 bp in length, consisting of 13 protein-coding genes (PCGs), 2 ribosomal RNAs, 22 tRNAs and a control region. The total base composition was A 29.6%, T 28.1%, C 25.7%, G 16.6%, with A + T content of 57.7%. This species shared the same gene composition and structural arrangement of mitogenome with other Cobitidae fish. Phylogenetic analyses showed that C. macrostigma has a relatively close relationship with other 18 Cobitinae species and was closer to Cobitis qujiangensis.


Similar content being viewed by others
References
Avise JC, Neigel JE, Arnold J (1984) Demographic influences on mitochondrial DNA lineage survivorship in animal populations. J Mol Evol 20:99–105. https://doi.org/10.1007/BF02257369
Bai J, Xu S, Nie Z, Wang Y, Zhu C, Wang Y, Min W, Cai Y, Zou J, Zhou X (2018) The complete mitochondrial genome of Huananpotamon lichuanense (Decapoda: Brachyura) with phylogenetic implications for freshwater crabs. Gene 646:217–226. https://doi.org/10.1016/j.gene.2018.01.015
Bernt M, Donath A, Juhling F, Externbrink F, Florentz C, Fritzsch G, Putz J, Middendorf M, Stadler PF (2013) MITOS: improved de novo metazoan mitochondrial genome annotation. Mol Phylogenet Evol 69:313–319. https://doi.org/10.1016/j.ympev.2012.08.023
Chen Y (1998) Zhongguo dong wu zhi. Ying gu yu gang. Li xing mu, Ke xue chu ban she: Xin hua shu dian. Beijing fa xing suo fa xing, Beijing
Cui Z, Liu Y, Li CP, You F, Chu KH (2009) The complete mitochondrial genome of the large yellow croaker, Larimichthys crocea (Perciformes, Sciaenidae): unusual features of its control region and the phylogenetic position of the Sciaenidae. Gene 432:33–43. https://doi.org/10.1016/j.gene.2008.11.024
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models,new heuristics and parallel computing. Nat Methods 9:772. https://doi.org/10.1038/nmeth.2109
de Meo PD, D'Antonio M, Griggio F, Lupi R, Borsani M, Pavesi G, Castrignano T, Pesole G, Gissi C (2012) MitoZoa 2.0: a database resource and search tools for comparative and evolutionary analyses of mitochondrial genomes in Metazoa. Nucleic Acids Res 40:D1168–D1172. https://doi.org/10.1093/nar/gkr1144
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Huang S, Tian X, Wang W, Song W, Zhang X, Bai X, Cao X (2015) The complete mitochondrial genome of natural Misgurnus bipartitus (Cypriniformes: Cobitidae). Mitochondr DNA 26:680–681. https://doi.org/10.3109/19401736.2013.840605
Kartavtsev YP, Jung SO, Lee YM, Byeon HK, Lee JS (2007) Complete mitochondrial genome of the bullhead torrent catfish, Liobagrus obesus (Siluriformes, Amblycipididae): genome description and phylogenetic considerations inferred from the Cyt b and 16S rRNA genes. Gene 396:13–27. https://doi.org/10.1016/j.gene.2007.01.027
Kim KY, Lee SY, Bang IC, Nam YK (2008) Complete mitogenome sequence of an endangered freshwater fish, Iksookimia choii (Teleostei; Cypriniformes; Cobitidae). Mitochondr DNA 19:438-445. https://doi.org/10.1080/19401730802449188
Kim IC, Kweon HS, Kim YJ, Kim CB, Gye MC, Lee WO, Lee YS, Lee JS (2004) The complete mitochondrial genome of the javeline goby Acanthogobius hasta (Perciformes, Gobiidae) and phylogenetic considerations. Gene 336:147–153. https://doi.org/10.1016/j.gene.2004.04.009
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Lanfear R, Calcott B, Ho SYW, Guindon S (2012) PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol 29:1695–1701. https://doi.org/10.1093/molbev/mss020
Laslett D, Canback B (2008) ARWEN: a program to detect tRNA genes in metazoan mitochondrial nucleotide sequences. Bioinformatics 24:172–175. https://doi.org/10.1093/bioinformatics/btm573
Li P, Yang CZ, Tu FY, Liu GX (2012) The complete mitochondrial genome of the Elongate loach (Cypriniformes: Cobitidae) . Mitochondr DNA 23:352–354. https://doi.org/10.3109/19401736.2012.690754
Liu SQ, Mayden RL, Zhang JB, Yu D, Tang QY, Deng X, Liu HZ (2012) Phylogenetic relationships of the Cobitoidea (Teleostei: Cypriniformes) inferred from mitochondrial and nuclear genes with analyses of gene evolution. Gene 508:60–72. https://doi.org/10.1016/j.gene.2012.07.040
Liu T, Jin X, Wang R, Xu T (2013) Complete sequence of the mitochondrial genome of Odontamblyopus rubicundus (Perciformes: Gobiidae): genome characterization and phylogenetic analysis. J Genet 92:423–432. https://doi.org/10.1007/s12041-013-0283-6
Lohse M, Drechsel O, Bock R (2007) OrganellarGenomeDRAW (OGDRAW): a tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Curr Genet 52:267–274. https://doi.org/10.1007/s00294-007-0161-y
Lohse M, Drechsel O, Kahlau S, Bock R (2013) OrganellarGenomeDRAW—a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucleic Acids Res 41:W575–W581. https://doi.org/10.1093/nar/gkt289
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25:955–964. https://doi.org/10.1093/nar/25.5.955
Moritz C, Dowling TE, Brown WM (1987) Evolution of animal mitochondrial DNA: relevance for population biology and systematics. Annu Rev Ecol Syst 18:269–292. https://doi.org/10.1146/annurev.es.18.110187.001413
Nass MMK, Nass S (1963a) Intramitochondrial fibers with DNA characteristics. 1. Fixation and electron staining reactions. J Cell Biol 19:593. https://doi.org/10.1083/Jcb.19.3.593
Nass MMK, Nass S (1963b) Intramitochondrial fibers with DNA characteristics. 2. Enzymatic and other hydrolytic treatments. J Cell Biol 19:613–629. https://doi.org/10.1083/Jcb.19.3.613
Oh DJ, Kim JY, Lee JA, Yoon WJ, Park SY, Jung YH (2007) Complete mitochondrial genome of the rock bream Oplegnathus fasciatus (Perciformes, Oplegnathidae) with phylogenetic considerations. Gene 392:174–180. https://doi.org/10.1016/j.gene.2006.12.007
Perdices A, Bohlen J, Šlechtová V, Doadrio I (2016) Molecular evidence for multiple origins of the European spined loaches (Teleostei, Cobitidae). PLoS One 11:e0144628. https://doi.org/10.1371/journal.pone.0151228
Perna NT, Kocher TD (1995) Patterns of nucleotide composition at fourfold degenerate sites of animal mitochondrial genomes. J Mol Evol 41:353–358. https://doi.org/10.1007/BF00186547
Qi D, Chao Y, Zhao L, Shen Z, Wang G (2013) Complete mitochondrial genomes of two relatively closed species from Gymnocypris (Cypriniformes: Cyprinidae): genome characterization and phylogenetic considerations. Mitochondr DNA 24:260–262. https://doi.org/10.3109/19401736.2012.744975
Shen Y, Dai W, Gao Z, Yan G, Gan X, He S (2017a) Molecular phylogeny and divergence time estimates using the mitochondrial genome for the hadal snailfish from the Mariana trench. Sci Bull 62:1106–1108. https://doi.org/10.1016/j.scib.2017.07.010
Shen Y, Kou Q, Zhong Z, Li X, He L, He S, Gan X (2017b) The first complete mitogenome of the South China deep-sea giant isopod Bathynomus sp. (Crustacea: Isopoda: Cirolanidae) allows insights into the early mitogenomic evolution of isopods. Ecol Evol 7:1869–1881. https://doi.org/10.1002/ece3.2737
Song HY, Mabuchi K, Satoh TP, Moore JA, Yamanoue Y, Miya M, Nishida M (2014) Mitogenomic circumscription of a novel percomorph fish clade mainly comprising “Syngnathoidei” (Teleostei). Gene 542:146–155. https://doi.org/10.1016/j.gene.2014.03.040
Stamatakis A, Hoover P, Rougemont J (2008) A rapid bootstrap algorithm for the RAxML web servers. Syst Biol 57:758–771. https://doi.org/10.1080/10635150802429642
Tang Q, Freyhof J, Xiong B, Liu H (2008) Multiple invasions of Europe by east Asian cobitid loaches (Teleostei: Cobitidae). Hydrobiologia 605:17–28. https://doi.org/10.1007/s10750-008-9296-1
Wang C, Chen Q, Lu G, Xu J, Yang Q, Li S (2008) Complete mitochondrial genome of the grass carp (Ctenopharyngodon idella, Teleostei): insight into its phylogenic position within Cyprinidae. Gene 424:96–101. https://doi.org/10.1016/j.gene.2008.07.011
Yang W, Zhang Y, Feng S, Liu L, Li Z (2018) The first complete mitochondrial genome of the Japanese beetle Popillia japonica (Coleoptera: Scarabaeidae) and its phylogenetic implications for the superfamily Scarabaeoidea. Int J Biol Macromol 118:1406–1413. https://doi.org/10.1016/j.ijbiomac.2018.06.131
Zardoya R, Meyer A (1996) The complete nucleotide sequence of the mitochondrial genome of the lungfish (Protopterus dolloi) supports its phylogenetic position as a close relative of land vertebrates. Genetics 142:1249–1263
Zhong L, Wang M, Li D, Tang S, Zhang T, Bian W, Chen X (2018) Complete mitochondrial genome of freshwater goby Rhinogobius cliffordpopei (Perciformes, Gobiidae): genome characterization and phylogenetic analysis. Genes Genom 40:1137–1148. https://doi.org/10.1007/s13258-018-0669-1
Acknowledgments
We are grateful for our crew help in collecting specimens. This study was supported by the research start-up fund of the Chongqing Normal University (Nos. 18XLB007).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 22 kb)
Rights and permissions
About this article
Cite this article
Yang, N., Li, Y., Liu, Z. et al. The complete mitochondrial genome of Cobitis macrostigma (Cypriniformes: Cobitidae: Cobitinae) and a phylogenetic implication for its closely related species. Biologia 75, 393–399 (2020). https://doi.org/10.2478/s11756-019-00309-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2478/s11756-019-00309-9