Abstract
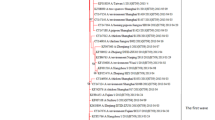
The principle of the present study was to determine the evolution of pandemic novel influenza A/H1N1 2009 virus (NIV) by phylogenetic, comparative and statistical analyses. The phylogenetic trees of eight genomic segments illustrate that, so far, the sequences of the NIVs (outbreak group A) are relatively homogeneous and derived by the event of multiple genetic reassortment of Eurasian and North American swine, avian and human viruses (group B). It implies that some of the influenza viruses in group B had higher potential to evolve and getting the ability to transmit from human-to-human after animal-to-human cross-species transmission. The second analysis shows that NIV had attempted a little evolutionary change among humans and before introduction into human it had long evolutionary history. Statistical analysis shows that viruses from both outbreak and nearest group have homologous genes in their genomes which might be reflecting the phylogenetic relationship of strains, and also the presence of unique mutations between groups A-B may associate with increased virulence of NIVs. Both phylogenetic and cluster analyses confirm that the gene exchange takes place between viruses originated from different species and it could be generated NIV with unpredictable pandemic potential. Hence, we conclude that an extensive study should be made to recognize, which reassortment groups are closely related to NIVs, and to determine the sites in the genes of NIV under greatest or least selection pressure, which will ultimately be important in the effective design of vaccine and drugs for ‘swine flu’.
Article PDF
Similar content being viewed by others
Abbreviations
- GD:
-
genetic distance
- HA:
-
haemagglutinin
- MCL:
-
maximum composite likelihood
- MP:
-
matrix protein
- NA:
-
neuraminidase
- NIV:
-
novel influenza A/H1N1 2009 virus
- NJ:
-
neighbour joining
- NP:
-
nucleoprotein
- NS:
-
non-structural protein
- PA:
-
RNA polymerase
- PB1:
-
catalytic RNA polymerase
- PB2:
-
cap-binding RNA polymerase
References
Antonovics J., Hood M.E. & Baker C.H. 2006. Molecular virology: was the 1918 flu avian in origin? Nature 440: E9.
Babakir-Mina M., Dimonte S., Perno C.F. & Ciotti M. 2009. Origin of the 2009 Mexico influenza virus: a comparative phylogentic analysis of the principal external antigens and matrix protein. Arch. Virol. 154: 1349–1352.
Benson D.A., Karsch-Mizrachi I., Lipman D.J., Ostell J. & Sayers E.W. 2011. GenBank. Nucleic Acids Res. 39: D32–D37.
Centers for Disease Control and Prevention (CDC). 2009a. Swine influenza A(H1N1) infection in two children — Southern California, March–April 2009. Morbidity and Mortality Weekly Report MMWR 58: 400–402.
Centers for Disease Control and Prevention (CDC). 2009b. Update swine influenza A(H1N1) infections — California and Texas, April 2009. Morbidity and Mortality Weekly Report MMWR 58: 1–3.
Dawood F.S., Jain S., Finelli L., Shaw M.W., ... & Zipprich J. 2009. Emergence of a novel swine-origin influenza A(H1N1) virus in humans. N. Engl. J. Med. 360: 2605–2615.
Ding N., Wu N., Xu Q., Chen K. & Zhang C. 2009. Molecular evolution of novel swine-origin A/H1N1 influenza viruses among and before human. Virus Genes 39: 293–300.
Fraser C., Donnelly C.A., Cauchemez S., Hanage W.P., ... & Roth C. 2009. Pandemic potential of a strain of influenza A (H1N1): early findings. Science 324: 1557–1561.
Gambotto A., Barratt-Boyes S.M., Jong M.D., Neumann G. & Kawaoka Y. 2008. Human infection with highly pathogenic H5N1 influenza virus. Lancet 371: 1464–1475.
Garten R.J., Davis C.T., Russell C.A., Shu B., ... & Cox N.J. 2009. Antigenic and genetic characteristics of swine-origin 2009 A(H1N1) influenza viruses circulating in humans. Science 325: 197–201.
Gibbs M. & Gibbs A. 2006. Molecular virology: was the 1918 pandemic caused by a bird flu? Nature 440: E8.
Hammer O., Harper D.A.T. & Ryan P.D. 2001. PAST: paleontological statistics software package for education and data analysis. Palaeontologia Electronica 4: art. 4.
Hatta M., Hatta Y., Kim J.H., Watanabe S., Shinya K., Nguyen T., Lien P.S., Le Q.M. & Kawaoka Y. 2007. Growth of H5N1 influenza A viruses in the upper respiratory tracts of mice. PLoS Pathog. 3: e133.
Kryazhimskiy S., Bazykin G.A. & Dushoff J. 2008. Natural selection for nucleotide usage at synonymous and nonsynonymous sites in influenza A virus genes. J. Virol. 82: 4938–4945.
Li Y.G., Chittaganpitch M., Waicharoen S., Kanai Y., Bai G.R., Kameoka M., Takeda N., Ikuta K. & Sawanpanyalert P. 2010. Characterization of H5N1 influenza viruses isolated from humans in vitro. Virol. J. 7: 112.
Ma W., Vincent A.L., Lager K.M., Janke B.H., Henry S.C., Rowland R.R.R., Hesse R.A. & Richt J.A. 2010. Identification and characterization of a highly riulent triple reassortant H1N1 swine influenza virus in the United States. Virus Genes 40: 28–36.
Munster V.J., de Wit E., van den Brand J.M.A., Herfst S., ... & Fouchier R.A. 2009. Pathogenesis and transmission of swineorigin 2009 A(H1N1) influenza virus in ferrets. Science 325: 481–483.
Nei M. & Kumar S. 2000. Molecular Evolution and Phylogenetics. Oxford University Press, Inc., New York.
Olsen C.W., Karasin A. & Erickson G. 2003. Characterization of a swine-like reassortant H1N2 influenza virus isolated from a wild duck in the United States. Virus Res. 93: 115–121.
Olsen W.C. 2002. The emergence of novel swine influenza viruses in North America. Virus Res. 85: 199–210.
Palese P. 2004. Influenza: old and new threats. Nat. Med. 10: 82–87.
Qu Y., Zhang R., Cui P., Song G., Duan Z. & Lei F. 2011. Evolutionary genomics of the pandemic 2009 H1N1 influenza viruses (pH1N1v). Virol. J. 8: 250.
Reid A.H., Taubenberger J.K. & Fanning T.G. 2004. Evidence of an absence: the genetic origins of the 1918 pandemic influenza virus. Nat. Rev. Microbiol. 2: 909–914.
Reis M.D., Hay A.J. & Goldstein R.A. 2009. Using nonhomogeneous models of nucleotide substitution to identify host shift events: application to the origin of the 1918 ’spanish’ influenza pandemic virus. J. Mol. Evol. 69: 333–345.
Schafer J.R., Kawaoka Y., Bean W.J., Suss J., Senne D. & Webster R.G. 1993. Origin of the pandemic 1957 H2 influenza A virus and the persistence of its possible progenitors in the avian reservoir. Virology 194: 781–788.
Schnitzler S.U. & Schnitzler P. 2009. An update on swine-origin influenza virus A/H1N1: a review. Virus Genes 39: 279–292.
Scholtissek C. 1990. Pigs as ‘mixing vessels’ for the creation of new pandemic influenza A viruses. Med. Principles Pract. 2: 65–71.
Smith G.J.D., Bahl J., Vijaykrishna D., Zhang J.Z., Poon L.L., Chen H., Webster R.G., Peiris J.S. & Guan Y. 2009a. Dating the emergence of pandemic influenza viruses. Proc. Natl. Acad. Sci. USA 106: 11709–11712.
Smith G.J., Vijaykrishna D., Bahl J., Lycett S.J., Worobey M., Pybus O.G., Ma S.K., Cheung C.L., Raghwani J., Bhatt S., Peiris J S., Guan Y. & Rambaut A. 2009b. Origin and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature 459: 1122–1126.
Suzuki Y. 2006. Natural selection on the influenza virus genome. Mol. Biol. Evol. 23: 1902–1911.
Tamura K., Dudley J., Nei M. & Kumar S. 2007. MEGA 4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24: 1596–1599.
Tamura K. & Nei M. 1993. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol. Biol. Evol. 10: 512–526.
Tamura K., Nei M. & Kumar S. 2004. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc. Natl. Acad. Sci. USA 101: 11030–11035.
Tamuri A.U., Reis D.M., Hay A.J. & Goldstein R.A. 2009. Identifying changes in selective constraints: host shifts in influenza. PLoS Comput. Biol. 5: e1000564.
Taubenberger J.K. 2006. The origin and virulence of the 1918 “Spanish” influenza virus. Proc. Amer. Phil. Soc. 150: 86–112.
Taubenberger J.K., Reid A.H., Lourens R.M., Wang R., Jin G. & Fanning T.G. 2005. Characterization of the 1918 influenza virus polymerase genes. Nature 437: 889–893.
Thompson J.D., Higgins D.G. & Gibson T.J. 1994. Clustal W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22: 4673–4680.
Van-Reeth K. & Nicoll A. 2009. A human case of swine influenza virus infection in Europe — implications for human health and research. Eurosurveillance. 14: art. 19124.
Webby R.J. & Webster R.G. 2001. Emergence of influenza A viruses. Phil. Trans. R. Soc. B. 356: 1817–1828.
Webster R.G., Bean W.J., Gorman O.T., Chambers T.M. & Kawaoka Y. 1992. Evolution and ecology of influenza A viruses. Microbiol. Rev. 56: 152–179.
Webster R.G., Peiris M., Chen H. & Guan Y. 2006. H5N1 outbreaks and enzootic influenza. Emerg. Infect. Dis. 12: 3–8.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Arunachalam, R., Paulkumar, K. & Annadurai, G. Phylogenetic analysis of pandemic influenza A/H1N1 virus. Biologia 67, 14–31 (2012). https://doi.org/10.2478/s11756-011-0163-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2478/s11756-011-0163-6