Abstract
Background
During the last decade, the scientific community has begun to investigate the composition and role of gut microbiota in normal health and disease. These studies have provided crucial information on the relationship between gut microflora composition and intestinal parasitic infection, and have demonstrated that many enteric pathogen infections are associated with altered gut microflora composition. In this study, we investigated the effects of Cryptosporidium parvum infection (zoonotic protozoan affecting a large range of vertebrates) on both qualitative and quantitative composition of gut microbiota in a CD-1 neonatal mouse model.
Methods
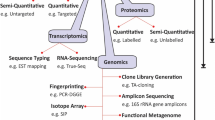
5-day-old neonate mice were experimentally infected with 105Cryptosporidium parvum Iowa oocysts by oesophageal gavage. The intestinal microbiota of both infected (Cp+) and uninfected (Cp−) mice groups was examined by high-throughput sequencing of the bacterial 16S rDNA gene V3–V4 hypervariable region.
Results
The most consistent change in the microbiota composition of Cp+ mice was the increased proportion of bacterial communities belonging to the Phylum Bacteroidetes. In contrast, the microbiota of Cp− mice was associated with increased proportions of several Firmicutes and Actinobacteria phyla members.
Conclusion
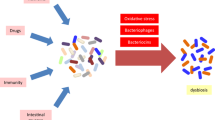
For the first time, our study provides evidence of an association between cryptosporidial infection and gut dysbiosis, thus contributing valuable knowledge to the as-yet little-explored field of Cryptosporidium–microbiota interactions in a neonatal mouse model.


Similar content being viewed by others
References
Anderson BC (1998) Cryptosporidiosis in bovine and human health. J Dairy Sci 81:3036–3041
Angly FE, Dennis PG, Skarshewski A, Vanwonterghem I, Hugenholtz P, Tyson GW (2014) CopyRighter: a rapid tool for improving the accuracy of microbial community profiles through lineage-specific gene copy number correction. Microbiome 2:11
Barash NR, Maloney JG, Singer SM, Dawson SC (2017) Giardia alters commensal microbial diversity throughout the murine gut. Infect Immun. https://doi.org/10.1128/iai.00948-16
Benson A, Pifer R, Behrendt CL, Hooper LV, Yarovinsky F (2009) Gut commensal bacteria direct a protective immune response against the human pathogen Toxoplasma gondii. Cell Host Microbe 6:187–196
Brook I (1991) Pathogenicity of Propionibacterium acnes in mixed infections with facultative bacteria. J Med Microbiol 34:249–252
Eren AM, Maignien L, Sul WJ, Murphy LG, Grim SL, Morrison HG, Sogin ML (2013) Oligotyping: differentiating between closely related microbial taxa using 16S rRNA gene data. Methods Ecol Evolut. https://doi.org/10.1111/2041-210x.12114
Eren AM, Morrison HG, Lescault PJ, Reveillaud J, Vineis JH, Sogin ML (2015) Minimum entropy decomposition: unsupervised oligotyping for sensitive partitioning of high-throughput marker gene sequences. ISME J 9:968–979
Fayer R, Gasbarre L, Pasquali P, Canals A, Almeria S, Zarlenga D (1998) Cryptosporidium parvum infection in bovine neonates: dynamic clinical, parasitic and immunologic patterns. Int J Parasitol 28:49–56
Finch GR, Daniels CW, Black EK, Schaefer FW, Belosevic M (1993) Dose response of Cryptosporidium parvum in outbred neonatal CD-1 mice. Appl Environ Microbiol 59:3661–3665
Foster JC, Glass MD, Courtney PD, Ward LA (2003) Effect of Lactobacillus and Bifidobacterium on Cryptosporidium parvum oocyst viability. Food Microbiol 20:351–357
Galan M, Razzauti M, Bard E et al (2016) 16S rRNA amplicon sequencing for epidemiological surveys of bacteria in wildlife. mSystems. https://doi.org/10.1128/msystems.00032-16
Geurden T, Thomas P, Casaert S, Vercruysse J, Claerebout E (2008) Prevalence and molecular characterisation of Cryptosporidium and Giardia in lambs and goat kids in Belgium. Vet Parasitol 155:142–145
Glendinning L, McLachlan G, Vervelde L (2017) Age-related differences in the respiratory microbiota of chickens. PLoS One. https://doi.org/10.1371/journal.pone.0188455
de Graaf DC, Vanopdenbosch E, Ortega-Mora LM, Abbassi H, Peeters JE (1999) A review of the importance of cryptosporidiosis in farm animals. Int J Parasitol 29:1269–1287
Harp JA, Wannemuehler MW, Woodmansee DB, Moon HW (1988) Susceptibility of germfree or antibiotic-treated adult mice to Cryptosporidium parvum. Infect Immunol 56:2006–2010
Hijjawi N (2010) Cryptosporidium: new developments in cell culture. Exp Parasitol 124:54–60
Honda K, Littman DR (2012) The microbiome in infectious disease and inflammation. Annu Rev Immunol 30:759–795
Hufeldt MR, Nielsen DS, Vogensen FK, Midtvedt T, Hansen AK (2010) Variation in the gut microbiota of laboratory mice is related to both genetic and environmental factors. Comp Med 60:336–347
Iebba V, Santangelo F, Totino V, Pantanella F, Monsia A, Di Cristanziano V, Di Cave D, Schippa S, Berrilli F, D’Alfonso R (2016) Gut microbiota related to Giardia duodenalis, Entamoeba spp. and Blastocystis hominis infections in humans from Côte d’Ivoire. J Infect Dev Ctries 10:1035–1041
Korich DG, Marshall MM, Smith HV, O’Grady J, Bukhari Z, Fricker CR, Rosen JP, Clancy JL (2000) Inter-laboratory comparison of the CD-1 neonatal mouse logistic dose-response model for Cryptosporidium parvum oocysts. J Eukaryot Microbiol 47:294–298
Kreisinger J, Bastien G, Hauffe HC, Marchesi J, Perkins SE (2015) Interactions between multiple helminths and the gut microbiota in wild rodents. Philos Trans R Soc B 370(1675):20140295. https://doi.org/10.1098/rstb.2014.0295
Lacroix-Lamandé S, Guesdon W, Drouet F, Potiron L, Lantier L, Laurent F (2014) The gut flora is required for the control of intestinal infection by poly(I:C) administration in neonates. Gut Microbes 5:533–540
Lantier L, Drouet F, Guesdon W, Mancassola R, Metton C, Lo-Man R, Werts C, Laurent F, Lacroix-Lamandé S (2014) Poly(I:C)-induced protection of neonatal mice against intestinal Cryptosporidium parvum infection requires an additional TLR5 signal provided by the gut flora. J Infect Dis 209:457–467
Levi Mortera S, Del Chierico F, Vernocchi P et al (2016) Monitoring perinatal gut microbiota in mouse models by mass spectrometry approaches: parental genetic background and breastfeeding effects. Front Microbiol. https://doi.org/10.3389/fmicb.2016.01523
Mammeri M, Chevillot A, Thomas M, Polack B, Julien C, Marden J-P, Auclair E, Vallée I, Adjou KT (2018) Efficacy of chitosan, a natural polysaccharide, against Cryptosporidium parvum in vitro and in vivo in neonatal mice. Exp Parasitol 194:1–8
Mariat D, Firmesse O, Levenez F, Guimarăes V, Sokol H, Doré J, Corthier G, Furet J-P (2009) The Firmicutes/Bacteroidetes ratio of the human microbiota changes with age. BMC Microbiol 9:123
Martín-Gómez S, Alvarez-Sánchez M, Rojo-Vázquez F (2006) A newborn mouse Cryptosporidium parvum infection model: its application to the study of therapeutic and prophylactic measures for controlling cryptosporidiosis in ruminants. Parasitol Res 99:1–6
Maurice CF, Knowles SCL, Ladau J, Pollard KS, Fenton A, Pedersen AB, Turnbaugh PJ (2015) Marked seasonal variation in the wild mouse gut microbiota. ISME J 9:2423–2434
Nguyen TLA, Vieira-Silva S, Liston A, Raes J (2015) How informative is the mouse for human gut microbiota research? Dis Model Mech 8:1–16
O’Donoghue PJ (1995) Cryptosporidium and cryptosporidiosis in man and animals. Int J Parasitol 25:139–195
Puiu D, Enomoto S, Buck GA, Abrahamsen MS, Kissinger JC (2004) CryptoDB: the cryptosporidium genome resource. Nucleic Acids Res 32(Database issue):D329–D331
Raetz M, Hwang S-H, Wilhelm CL et al (2013) Parasite-induced T H 1 cells and intestinal dysbiosis cooperate in IFN-γ-dependent elimination of Paneth cells. Nat Immunol 14:136–142
Ramirez NE, Ward LA, Sreevatsan S (2004) A review of the biology and epidemiology of cryptosporidiosis in humans and animals. Microbes Infect Inst Pasteur 6:773–785
Ras R, Huynh K, Desoky E, Badawy A, Widmer G (2015) Perturbation of the intestinal microbiota of mice infected with Cryptosporidium parvum. Int J Parasitol 45:567–573
Sherwood D, Angus KW, Snodgrass DR, Tzipori S (1982) Experimental cryptosporidiosis in laboratory mice. Infect Immun 38:471–475
Shkoporov AN, Chaplin AV, Khokhlova EV, Shcherbakova VA, Motuzova OV, Bozhenko VK, Kafarskaia LI, Efimov BA (2015) Alistipes inops sp. nov. and Coprobacter secundus sp. nov., isolated from human faeces. Int J Syst Evol Microbiol 65:4580–4588
Stockinger S, Hornef MW, Chassin C (2011) Establishment of intestinal homeostasis during the neonatal period. Cell Mol Life Sci 68:3699–3712
Thoene-Reineke C, Fischer A, Friese C, Briesemeister D, Göbel UB, Kammertoens T, Bereswill S, Heimesaat MM (2014) Composition of intestinal microbiota in immune-deficient mice kept in three different housing conditions. PLoS One 9:e113406
Verma AK, Verma R, Ahuja V, Paul J (2012) Real-time analysis of gut flora in Entamoeba histolytica infected patients of Northern India. BMC Microbiol 12:183
Walsh CJ, Guinane CM, O’Toole PW, Cotter PD (2014) Beneficial modulation of the gut microbiota. FEBS Lett 588:4120–4130
Walter J (2008) Ecological role of Lactobacilli in the gastrointestinal tract: implications for fundamental and biomedical research. Appl Environ Microbiol 74:4985–4996
Zhang M, Zhang M, Zhang C, Du H, Wei G, Pang X et al (2009) Pattern extraction of structural responses of gut microbiota to rotavirus infection via multivariate statistical analysis of clone library data. FEMS Microbiol Ecol 70:177–185
Acknowledgements
M. Mammeri was the grateful recipient of a Cifre (Industrial Research Training Agreement) grant. He would like to thank the Phileo Lesaffre Animal Care centre (France) and the ANRT (National Association for Technical Research), Ministry of Research. The authors thank Alain Bernier and Océane Le Bidel for their technical expertise and skilful animal husbandry; Muriel Vayssier and Jean-François Cosson for their constructive discussions on the use of 16S rDNA high-throughput sequencing strategy, and Illumina Miseq platform at Eurofins Genomics (Germany) for performing the 16S rDNA high-throughput sequencing.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicting interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mammeri, M., Chevillot, A., Thomas, M. et al. Cryptosporidium parvum-Infected Neonatal Mice Show Gut Microbiota Remodelling Using High-Throughput Sequencing Analysis: Preliminary Results. Acta Parasit. 64, 268–275 (2019). https://doi.org/10.2478/s11686-019-00044-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2478/s11686-019-00044-w