Abstract
Poly(ADP-ribosyl)ation (PARylation) is a protein modification reaction regulating various diverse cellular functions ranging from metabolism, DNA repair and transcription to cell death. We set out to investigate the role of PARylation in wound healing, a highly complex process involving various cellular and humoral factors. We found that topically applied poly(ADP-ribose) polymerase (PARP) inhibitors 3-aminobenzamide and PJ-34 accelerated wound closure in a mouse model of excision wounding. Moreover, wounds also closed faster in PARP-1 knockout mice as compared with wild-type littermates. Immunofluorescent staining for poly(ADP-ribose) (PAR) indicated increased PAR synthesis in scattered cells of the wound bed. Expression of interleukin (IL)-6, tumor necrosis factor (TNF)-α, inducible nitric oxide synthase and matrix metalloproteinase-9 was lower in the wounds of PARP-1 knockout mice as compared with control, and expression of IL-1 β, cyclooxygenase-2, TIMP-1 and -2 also were affected. The level of nitrotyrosine (a marker of nitrating stress) was lower in the wounds of PARP-1 knockout animals as compared with controls. In vitro scratch assays revealed significantly faster migration of keratinocytes treated with 3-aminobenzamide or PJ34 as compared with control cells. These data suggest that PARylation by PARP-1 slows down the wound healing process by increasing the production of inflammatory mediators and nitrating stress and by slowing the migration of keratinocytes.
Similar content being viewed by others
Introduction
Physical injury to the skin triggers a well-coordinated series of events leading to wound closure. Wound healing occurs in three phases: inflammation, proliferation and remodeling. The inflammatory phase involves hemostatic activation (vasoconstriction, platelet aggregation and clot formation), vasodilation, inflammatory cell migration, phagocytosis and production of inflammatory mediators. In the proliferative phase, granulation tissue is formed, wound edges contract and epithelial cells migrate to cover the wound surface. During remodeling, tissue structure is reorganized to most closely resemble normal skin. This phase involves disappearance of the majority of inflammatory cells, deposition and maturation of proteoglycans and collagens replacing transient matrix elements. Many soluble factors such as inflammatory cytokines (tumor necrosis factor [TNF]-α, interleukin [IL]-1β, IL-6) and chemokines, growth factors (platelet-derived growth factor [PDGF], epidermal growth factor [EGF], fibroblast growth factor [FGF], transforming growth factor [TGF] β, keratinocyte growth factor [KGF7]) have been shown to regulate the wound healing process (1). Moreover, reactive oxygen and nitrogen species (ROS and RNS, respectively) are also abundantly produced during wound healing and regulate various aspects of the process (2).
Understanding complex regulatory circuitries operating in normal or pathological wound healing is of utmost importance for the successful treatment of wound healing abnormalities. These include impaired wound healing in diabetes, systemic sclerosis or chronic leg ulcers.
The treatment of chronic, nonhealing wounds represents a major health issue with treatment costs estimated to amount to 25 billion USD per year in the United States alone (3). Current modern or experimental treatments include dressings impregnated with collagenase, DNase, PDGF or nitric oxide-releasing compounds. In spite of the high number of novel and innovative approaches to cure nonhealing wounds, the efficient treatment of these diseases still represents an unmet medical need.
On the basis of available literature, PARylation appeared to be a likely candidate for being a regulator of the wound healing process (see below). PARylation is a covalent protein modification carried out by poly(ADP-ribose) polymerase (PARP) enzymes. PARPs cleave off nicotinamide from NAD+ and attach the remaining ADP-ribose moieties to suitable protein acceptors. By cleaving many NAD+ molecules and adding further ADP-ribose units to the protein-proximal first residue, these enzymes build a large, branched poly(ADP-ribose) (PAR) polymer on proteins. The polymer is degraded mostly by PAR glycohydrolase (PARG). PARP-1, -2 and -3 are DNA-damage-sensitive enzymes that become activated by DNA breaks. For example PARP-1, the main PARP enzyme, has been shown to become activated by oxidative stress-induced DNA damage and to facilitate DNA break repair. Moreover, PARP-1 is a cofactor of nuclear factor (NF)-κB, the master regulator of the transcription of inflammatory mediators. PARylation also regulates a wide variety of molecular events and cellular functions including genome organization, replication, protein stability and cell death.
On the basis of the involvement of many of these PARylation-related reactions (oxidative stress response, expression of inflammatory cytokines and chemokines, cell proliferation and migration) in the wound healing process, we set out to investigate the role of PARylation in a mouse model of excision wounding.
Materials and Methods
Animals
Animal experiments were approved by the Institutional Animal Care and Use Committee of University of Debrecen (protocol number 9/2008/DEMÁB) and were carried out in accordance with the EU guidelines on animal protection (4).
Adult male BALB/c mice (6 to 8 wks old) were used for assessing the effect of the PARP inhibitors 3-aminobenzamide (3-AB) and PJ34 (Sigma-Aldrich, St. Louis, MO, USA) on wound closure. In addition, male (6 to 8 wks old) homozygous PARP-1 knockout mice (PARP-1−) and their respective wild-type (PARP-1+/+) littermates (5) were also used. All animals were housed individually under controlled conditions with 12-h light-dark cycle, and were allowed free access to water and diet.
Wound Healing Model
Wound incision was performed as described previously in (6). Mice were anaesthetized with isoflurane (Abbott Animal Health, Abbott Park, IL, USA). The fur was removed from the back of mice by applying a commercial depilatory cream and hairless skin was wiped with 70% ethanol. Mice were kept warm during anesthesia and surgery using a heat lamp. Two full thickness 4 mm punch biopsies were made on the dorsal surface of BALB/c mice and the wound beds were wiped with povidone-iodine to prevent infection. The PARP inhibitors PJ-34 and 3-aminobenzamide (3-AB) were applied locally on one of the wounds in hydrophilic cream in 100 µmol/L and 50 µmol/L concentrations, respectively. The control wound was treated with hydrophilic cream only. Treatments were repeated daily for 7 d and wounds were covered with a surgical dressing to keep the cream in place.
Wounding and treatments of PARP-1−/− and PARP-1+/+ mice (n = 6 for each genotype) was performed the same way as detailed above.
Quantification of Wound Closure
Wounds from individual mice were digitally photographed daily for 7 d. All photographs were taken from an angle perpendicular to the wound with a sterile ruler placed adjacent to the wounds for standardization (7). The wound area was quantified using NIH ImageJ software (NIH, Bethesda, MD, USA); and wound closure was expressed as the ratio of actual wound area to initial wound size.
Tissue Harvesting
Tissue samples were collected from a separate cohort of mice (n = 6 per time point) on d 0, 1, 2, 4 and 8 after wounding. Under isoflurane anesthesia, wound tissue was harvested using a circular blade (6 mm). After wounds had been collected, animals were euthanized. One of the two wound samples in each animal was processed for histology while the other was snap frozen in liquid nitrogen for RNA and protein analysis (8).
Histological Examination
PAR was detected by immunostaining as formerly described in (9). In brief, immunofluorescent staining was performed on formalin-fixed skin tissues by use of a PAR-specific monoclonal antibody (clone 10H; Enzo Life Sciences, Farmingdale, NY, USA) and a fluorescein-based mouse-on-mouse Immunodetection kit (Vector Laboratories, Peterborough, UK). Nuclei were counterstained with DAPI (Invitrogen, Life Technologies [Thermo Fisher Scientific Inc., Waltham, MA, USA]); and stainings were visualized with a Zeiss Axiolab fluorescent microscope.
RNA Extraction and Real-Time Quantitative PCR Analysis (RT-qPCR)
RT-qPCR was carried out as described in (10). In brief, total RNA from frozen skin tissue was isolated with TRIzol reagent (Invitrogen, Life Technologies [Thermo Fisher Scientific Inc.]) according to the manufacturer’s instructions. The concentration of isolated RNA was determined spectrophotometrically and finally adjusted to 2 µg for the reverse transcription (RT) step. By use of a high capacity-RT kit (Applied Biosystems [Thermo Fisher Scientific Inc.]), RT was performed in a mixture of 2 µL RT buffer, 0.8 µL dNTP, 2 µL random primer, 1 µL reverse transcriptase and diethylpyrocarbonate (DEPC) H2O up to 10 µL. PCR cycles (Veriti 96-Well Thermal Cycler, Thermo Fisher Scientific Inc.) were set as follows: 25°C for 10 min, 42°C for 60 min, 95°C for 5 min, 94°C for 5 min, followed by 35 cycles 94°C for 30 s, 67°C for 1 min and 74°C for 1 min. SYBR Green (Applied Biosystems [Thermo Fisher Scientific Inc.]) qPCR reactions were run in ABI 7500 thermal cycler (Applied Biosystems [Thermo Fisher Scientific Inc.]). Sequences of primers are given in Supplementary Table 1. Expression values were normalized to the control gene glyceraldehyde 3-phosphate dehydrogenase (GAPDH) expression.
Western Blot Analysis
The method of lysate preparation and immunoblotting was carried out as previously reported by Kioka et al. (11). Briefly, frozen skin tissue (approximately 5 mg) was homogenized in 300 µL RIPA buffer (20 mmol/L Tris-HCL pH7.4, 150 mmol/L NaCl, 1 mmol/L EDTA, 1% Triton-X100, 1% sodium deoxycholate, 0.1% SDS) with PMSF (1 mmol/L) and protease inhibitor cocktail (5 mmol/L) added immediately before use. Protein concentrations were determined using BCA kit (Pierce, Rockford, IL, USA). Proteins were separated on 8% SDS-polyacrylamide gels, and then transferred onto nitrocellulose membranes. Nonspecific protein binding was blocked by incubating the membranes in 5% w/v BSA in Tris-buffered saline with Tween® 20 (TBS-T; 50 mmol/L Tris-HCl, pH 7.6, 150 mmol/L NaCl, and 0.1% v/v Tween-20). Membranes were incubated overnight at 4°C with mouse monoclonal anti-nitrotyrosine antibody (Cayman, Ann Arbor, MI, USA) or rabbit polyclonal anti-FGF antibody (Abcam, Cambridge, UK) at dilutions of 1:1000 and 1:500, respectively. After washes, membranes were incubated with either HRP-conjugated anti-mouse or anti-rabbit antibody at a dilution of 1:10,000 for 1 h at room temperature. Detection of mouse monoclonal β-actin (Sigma-Aldrich) (1:5000 dilution, 1-h-room-emperature incubation) served for normalization. Protein bands were detected using the ECL Plus kit (Pierce) and densitometry was carried out using NIH ImageJ software.
Scratch Assay
Confluent cultures of primary human keratinocytes seeded in 96-well plates were scratched with a small pipette tip utilizing a Tecan Freedom EVO liquid handling robot (Tecan Group Ltd., Männedorf, Switzerland). Damaged cells were removed by washing with PBS solution. A triplicate of freshly wounded samples was dried and used as control (0% healed samples). Cells were treated with PJ-34 or 3-AB and incubated for 48 h. Cell culture medium was replaced with Coomassie staining solution and cells were stained for 20 min. Samples were then washed twice with PBS and photos were taken under a Zeiss AxioVision microscope (Carl Zeiss Microscopy GmbH, Jena, Germany). Images were analyzed with Tscratch software. Ratio of covered area was compared with freshly wounded samples.
Statistical Analysis
All values are presented as means ± SEM. Statistical analysis of experimental data was performed with one-way analysis of variance (ANOVA), followed by Dunnett test for multiple comparisons. Results were considered statistically significant when p < 0.05.
All supplementary materials are available online at https://doi.org/www.molmed.org .
Results
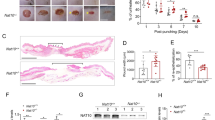
To investigate the role of PARylation in wound healing, we set up a mouse model of excision wounding with two wounds made on the back of each mice. One of the wounds on each mouse was treated with a hydrophilic crème whereas the other wound was covered with hydrophilic crème containing the PARP inhibitor 3-AB (50 µmol/L). We followed wound closure with time and found that 3-AB significantly accelerated the closure of wounds (Figure 1).
Two full thickness cutaneous wounds (4-mm diameter) were made on the back of male BALB/c mice, and wounds were evaluated by daily digital photography. Wounds were treated topically with either vehicle (hydrophilic cream, left wound) or PJ34 (100 µmol/L in hydrophilic cream; right wound) for 7 consecutive d; starting 24 h after excision. The line graph represents the mean of percent change relative to original wound size ± SEM (n = 8). *P < 0.05, **P < 0.01, ***P < 0.001: significantly different as compared between PJ34-treated (solid line) and vehicle-treated (dashed line) wounds in the same time point.
To confirm our results obtained with 3-AB, we also applied another PARP inhibitor, PJ-34. Similarly to 3-AB, the application of PJ-34 has also stimulated the wound-healing process (Figure 2).
Two full thickness cutaneous wounds (4-mm diameter) were made on the back of male BALB/c mice and wounds were evaluated by daily digital photography. Wounds were treated topically with either vehicle (hydrophilic cream, left wound) or 3-AB (50 µmol/L in hydrophilic cream; right wound) for 7 consecutive days; starting 24 h after excision. Each time point represents mean of eight measurements ± SEM (*P < 0.05, **P < 0.01, ***P < 0.001)
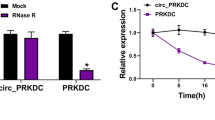
Since PARP-1 is responsible for the bulk of PAR synthesis in oxidative stress situations such as wound healing, we hypothesized that the effects of PARP inhibitors is mainly due to inhibition of PARP-1. Indeed wounds on PARP-1 knockout animals healed faster than wounds on wild-type mice (Figure 3), confirming that activation of PARP-1 leads to delayed wound healing. Expression of PARP-1 mRNA increased between d 0 and d 4, but declined by d 8 (Figure 4A). Basal expression of PARP-2 was much lower than that of PARP-1, and it also showed a moderate increase up to d 4 (Figure 4B). PARylation, however, is not primarily regulated at the expression level but mostly at the activity level. Appearance of PAR, the product of PARP-catalyzed reaction in the woundbed (Figure 4C) indicates that PARylation indeed takes place during wound healing.
Relative expression of (A) PARP-1/GAPDH mRNA (in PARP-1+/+ mice), (B) PARP-2/GAPDH mRNA (in PARP-1+/+ and PARP-1−/− mice) were determined from wound tissue samples using RT-qPCR at different time points. Error bars represent SEM of six independent samples. (**P < 0.01). (C) Immunofluorescent staining of formalin-fixed, paraffin-embedded skin tissues (wound beds) for PAR formation (green). Nuclei were counterstained with DAPI (blue).
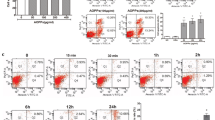
Inflammation is an essential phenomenon accompanying the wound healing process. Excision wounding triggered the expression of various inflammatory mediators such as IL-1β, TNF-α, IL-6, cyclooxygenase-2 (COX-2) and inducible nitric oxide synthase (iNOS) (Figure 5A-E). PARP-1 deficiency affected the expression of genes encoding most of these inflammatory mediators with the most significant reduction seen in IL-6, TNF-α and iNOS expression. Production of much nitric oxide may lead to the formation of peroxynitrite, a powerful protein-nitrating agent. Protein nitration was most prominent on a 57-kDa protein of the wound homogenate. Consistent with lower iNOS expression in PARP-1 knockout mice, the intensity of tyrosine nitration was also lower in the wound lysates of PARP-1-deficient animals (Figure 5F).
Expression of inflammatory cytokines (A–C), iNOS (D) and COX-2 (E) were quantified using RT-qPCR. Expression values were normalized to GAPDH. (F) Immunoblot analysis of nitrotyrosine in PARP-1+/+ and PARP-1−/− mice. All blots were normalized β-actin. Values are expressed as mean ± SEM. (n = 6). (*P < 0.05, **P < 0.01, ***P < 0.001)
Matrix metalloproteinases (MMPs) and their tissue inhibitors (TIMPs) also play important roles in wound healing (12). We have determined the expression of MMP-9, TIMP-1 and TIMP-2 mRNA in homogenates of wounds from wild-type and PARP-1 knockout mice. Expression of MMP-9, TIMP-1 and -2 increased until d 4 but declined by d 8 (Supplementary Figure S1). The expression profile of TIMP-2 in PARP-1 knockout mice did not significantly differ from that of wildtype mice. However, expression of MMP-9 on d 1 and d 4 was significantly lower in the PARP-1-deficient animals, whereas the expression of TIMP-1 in PARP-1 knockout mice was lower only on d 4.
Proliferation and migration of fibroblasts as well as keratinocytes is important for granulation tissue formation and epithelialization. In in vitro scratch assays, inhibition of PARylation by either 3-aminobenzamide or PJ-34 facilitated migration of primary human keratinocytes and thus the repopulation of the scratched surface (Figure 6).
PARP inhibition stimulates keratinocyte migration. Confluent cultures of keratinocytes were scratched with a pipette tip fixed to a liquid handling robot arm followed by washing and incubation of cells with medium (control), with PJ-34 (10 µmol/L) or 3-AB (3 mmol/L). Microphotographs were taken after 48 h. Representative images are presented (top) and repopulation of the scratched areas also was quantified by image analysis.
Discussion
The wound-healing process requires a precisely orchestrated series of tightly controlled cellular events and it has four main phases: hemostasis, inflammation, proliferation and remodeling. Any disease condition (for example, diabetes, circulatory problems) or external factor (for example, cytostatic drugs) disturbing this highly complex process may lead to the development of chronic nonhealing wounds. On the basis of the pleiotropic biological functions of PARylation by PARP-1 we hypothesized that it may regulate wound healing. Our main finding is that PARP-1 activity slows down the wound-healing process. Both the PARP-1 knockout phenotype and treatment with PARP inhibitors accelerated the healing of incision wounds. Accompanying accelerated wound healing, a suppression of inflammatory cytokines (IL-1β, TNF-α, IL-6), COX-2, MMP-9 and iNOS could also be observed. The somewhat surprising finding of accelerated wound healing upon genetic inactivation of the parp-1 gene or inhibition of PARP activity can be due to a combination of effects on various cellular events required for wound healing as discussed below.
One possible explanation underlying the stimulatory effect of PARP inhibition or PARP-1 deficiency may be improved cell metabolism or maintenance of general cellular “health.” It has been demonstrated in a high number of cellular and animal models characterized by oxidative stress that PARP-1 becomes activated by DNA breakage caused by ROS and RNS (13). Intense or sustained PARP-1 activation may lead to a compromised cellular energetic state due to depletion of NAD (the substrate of PARPs) and ATP. Compromised cellular energetics may lead to cell dysfunction or even to (necroptotic) cell death (14). One plausible hypothesis explaining our current findings is that PARP-1-mediated cell dysfunction may disturb the complex network of wound healing regulation. ROS-induced keratinocyte or fibroblast injury has been repeatedly demonstrated to occur via PARP activation (15–17), and it cannot be excluded that this pathway also operates in excision wounds. In fact, detection of PAR polymer in the wound bed (Figure 4C) illustrates that this might indeed be the case. Slowing down wound healing by PARP activation may even serve a good purpose: considering the role of PARylation in single strand break sensing (and consequently repair) it may slow down cell proliferation (required for granulation tissue formation and epithelialization) and thus provide a sufficient amount of time for the DNA repair machinery to restore the integrity of DNA.
Another interesting possibility is that a suppressed inflammatory response in PARP-1 knockout animals or in PARP inhibitor-treated mice may result in accelerated wound healing. The role of inflammation in wound healing is somewhat controversial but most reports underline the beneficial effects of antiinflammatory therapies. One common feature of most nonhealing wounds is persistent inflammation which is mostly due to macrophage dysregulation leading to an M1 dominated (inflammatory) phenotype over M2 (wound healing) phenotype (18). Moreover, application of mesenchymal stem cells, a type of multipotent stem cells with a strong antiinflammatory effect has been shown repeatedly to improve wound healing (19–21). Furthermore, scarless fetal wound healing is also characterized by a reduced inflammatory response (22), indicating that inflammation affects not only the dynamics but also the quality of wound healing.
Our finding that expression levels of TNF-α and, to a lesser extent, IL-1β is lower in PARP-1 knockout mice may be one of the factors underlying improved wound healing. It has been suggested that targeting these two inflammatory cytokines may be beneficial for the wound healing process. For example Mori et al. (23) reported accelerated wound healing in TNF receptor-deficient mice. Moreover, Infliximab, a chimeric monoclonal antibody against TNF-α used to treat autoimmune diseases, has been found to improve healing of chronic wounds (24). Interestingly, despite promising opportunities, the therapeutic potential of targeting IL-1β in inflammatory conditions (for example, autoimmune diseases) has not yet been explored in such a detail as for TNF-α. As for wound healing, it has been reported that disruption of IL-1β signaling by knocking out IL-1 receptor type I improved the quality of wound healing (lower levels of collagen and improved restoration of normal skin architecture) identifying IL-1β signaling as a potential target in wound healing (25). Moreover, blocking the Nod-like receptor protein 3 (NLRP-3)-caspase-1-IL-1β pathway also was demonstrated to improve wound healing in diabetic mice (18). Of note, expression of IL-1β in our current model did not differ very much between wildtype and PARP-1-knockout animals, suggesting that IL-1β is not the main factor contributing to the effects of PARP-1 inhibition/knockout in this model.
IL-6, the level of which also was found to be lower in PARP-1 knockout mice (Figure 5), is also considered as a key inflammatory cytokine. However, its role in wound healing appears to be opposite to those of TNF-α or IL-1β. For example, treatment with IL-6 augmented wound healing in immunosuppressed mice (26). In line with this, Lin et al. (27) reported delayed wound healing in IL-6-deficient mice. Thus, reduced IL-6 levels are not likely to explain accelerated wound healing in PARP-1-deficient mice.
Here we also report reduced expression of iNOS in the wounds of PARP-1-deficient mice. Expression of iNOS is regarded as a general feature of the inflammatory response. Its role in wound healing, however, appears to be controversial with iNOS deficiency reported to either delay (28) or have no effect on wound healing (29) depending on the type of wounds investigated. Impaired collagen accumulation also has been observed in iNOS-deficient mice (30), further strengthening the hypothesis that iNOS regulates wound healing. Nitration of proteins also was found to occur in the wound tissue with nitration of an unknown 57-kDa protein being less intense in the PARP-1-deficient tissue (see Figure 5.). PARP inhibition or PARP-1 deficiency have been shown to reduce both oxidative and nitrative stress in different animal models of tissue injury and inflammation ranging from arthritis to sepsis, ischemia-reperfusion or burn injury (13,31). Nitric oxide (NO) and superoxide need to be produced in near equimolar amounts to form peroxynitrite, one of the most prominent nitrating agents. PARP inhibition has been shown to reduce production of both NO and ROS in various diverse models of tissue injury and inflammation and this was shown to result in suppressed nitration of proteins, lipids and DNA (13). Decreased ROS production may also suppress activation of NF-κB, the master regulator of the transcription of inflammatory mediators which may culminate in reduced production of inflammatory cytokines and chemokines (see above). Protein-protein interaction between NF-κB and PARP-1 also may be important, but direct stimulation of NF-κB by PARP-1 may only be relevant in PARP-1-deficient animals, but not in mice treated with PARP inhibitors since the interaction between these two proteins does not require PARP-1 activity (32).
Matrix metalloproteinases (MMPs) are also important, especially in the remodeling phase of wound healing. Here we found reduced expression of MMP-9 in the wounds of PARP-1-deficient mice and expression of some TIMPs was also dysregulated, but to a lower extent. In line with our finding, Kauppinen and Swanson also reported (33) that PARP inhibition suppressed MMP-9 expression in TNF-α-stimulated microglia cells. Increased MMP9 has been suggested to predict poor wound healing in diabetic foot ulcers (34). Therefore, we think it is possible that suppressed MMP-9 expression may have contributed to accelerated wound healing in PARP-1 deficient mice.
An interesting finding in our current paper was that keratinocytes migrated faster in the presence of a PARP inhibitor. Repopulation of the scratched area requires both proliferation and cell migration. The role of PARP-1/PARylation in cell proliferation is somewhat controversial. On the one hand, PARP-1 has been shown to be part of the multiprotein replication complex (35) while on the other hand various studies identified it as an inhibitor of replication (36) and a negative regulator of cell proliferation (37). Its role in cell migration has not yet been systematically investigated. Thus identifying critical molecular control points of accelerated proliferation/migration of PARP-inhibited cells requires further investigation.
We should note a limitation of our study: while the various stages of wound healing may take several weeks to be completed, our study focused on the initial events taking place in the first days after wounding. Thus, to get a comprehensive view on the role of PARylation in the complete wound-healing process, we should also consider factors such as angiogenesis and remodeling. Of note, PARP-1 has been shown to be required for both angiogenesis (38–40) and fibrosis (41). These observations may have implications for the later stages of wound healing, which might be worth investigating.
Overall, our data presented here are in line with the plethora of previous studies reporting the inflammation-promoting and tissue-damage-mediator roles of PARylation/PARP-1 as described in diverse biological models ranging from various forms of shock, sepsis, ischemia-reperfusion injury (13), arthritis (15), contact hypersensitivity (42), experimental allergic encephalomyelitis (16) and many others (13). Common features of PARP inhibition/knockout in these in vivo models include suppression of the expression of inflammatory mediators, reduced migration of inflammatory cells to the site of inflammation, inhibition of oxidative and nitrative stress and prevention of cell dysfunction and cell death. Some of these changes might also be relevant in explaining our current findings.
Conclusion
In summary, we report here that excision wounds heal faster if PARylation is inhibited or PARP-1 gene is knocked out, indicating that PARylation by PARP-1 delays wound healing. Owing to the high complexity or regulatory circuitries involved in wound healing, it is impossible to trace the effect of PARylation and PARP-1 on wound healing to a single molecular event. On the basis of our current data, reduced expression of inflammatory mediators with special regard to TNF-α, IL-1β and iNOS, suppressed nitrative stress and accelerated keratinocyte migration may be the key elements underlying accelerated wound healing in PARP-1-deficient animals or in mice treated with PARP inhibitors.
Disclosure
The authors declare they have no competing interests as defined by Molecular Medicine, or other interests that might be perceived to influence the results and discussion reported in this paper.
References
Behm B, Babilas P, Landthaler M, Schreml S. (2012) Cytokines, chemokines and growth factors in wound healing. J. Eur. Acad. Dermatol. Venereol. 26:812–20.
Bryan N, Ahswin H, Smart N, Bayon Y, Wohlert S, Hunt JA. (2012) Reactive oxygen species (ROS)—a family of fate deciding molecules pivotal in constructive inflammation and wound healing. Eur. Cell. Mater. 24:249–65.
Staylor A. (2009) Wound care devices: growth amid uncertainty. MedTech Insight. 11:32–47.
Directive 2010/63/EU of the European Parliament and of the Council of 22 September 2010 on the protection of animals used for scientific purposes. 2010 O.J. L 276/33.
de Murcia JM, et al. (1997) Requirement of poly(ADP-ribose) polymerase in recovery from DNA damage in mice and in cells. Proc. Natl. Acad. Sci. U. S. A. 94:7303–7.
Muller-Decker K, Hirschner W, Marks F, Furstenberger G. (2002) The effects of cyclooxygenase isozyme inhibition on incisional wound healing in mouse skin. J. Invest. Dermatol. 119:1189–95.
Sullivan SR, et al. (2007) Topical application of laminin-332 to diabetic mouse wounds. J. Dermatol. Sci. 48:177–88.
Lim Y, Levy M, Bray TM. (2004) Dietary zinc alters early inflammatory responses during cutaneous wound healing in weanling CD-1 mice. J. Nutr. 134:811–6.
Munoz-Gamez JA, et al. (2009) PARP-1 is involved in autophagy induced by DNA damage. Autophagy. 5:61–74.
Szabo E, et al. (2001) Peroxynitrite production, DNA breakage, and poly(ADP-ribose) polymerase activation in a mouse model of oxazolone-induced contact hypersensitivity. J. Invest. Dermatol. 117:74–80.
Kioka N, et al. (2010) Crucial role of vinexin for keratinocyte migration in vitro and epidermal wound healing in vivo. Exp. Cell Res. 316:1728–38.
Gill SE, Parks WC. (2008) Metalloproteinases and their inhibitors: regulators of wound healing. Int. J. Biochem. Cell Biol. 40:1334–47.
Virag L, Szabo C. (2002) The therapeutic potential of poly(ADP-ribose) polymerase inhibitors. Pharmacol. Rev. 54:375–429.
Virag L, Robaszkiewicz A, Rodriguez-Vargas JM, Oliver FJ. (2013) Poly(ADP-ribose) signaling in cell death. Mol. Aspects Med. 34:1153–67.
Szabo C, et al. (1998) Protection against peroxynitrite-induced fibroblast injury and arthritis development by inhibition of poly(ADP-ribose) synthase. Proc. Natl. Acad. Sci. U. S. A. 95:3867–72.
Scott GS, Hake P, Kean RB, Virag L, Szabo C, Hooper DC. (2001) Role of poly(ADP-ribose) synthetase activation in the development of experimental allergic encephalomyelitis. J. Neuroimmunol. 117:78–86.
Virag L, et al. (2002) Nitric oxide-peroxynitrite-poly(ADP-ribose) polymerase pathway in the skin. Exp. Dermatol. 11:189–202.
Mirza RE, Fang MM, Weinheimer-Haus EM, Ennis WJ, Koh TJ. (2014) Sustained inflammasome activity in macrophages impairs wound healing in type 2 diabetic humans and mice. Diabetes. 63:1103–14.
Falanga V, et al. (2007) Autologous bone marrow-derived cultured mesenchymal stem cells delivered in a fibrin spray accelerate healing in murine and human cutaneous wounds. Tissue Eng. 13:1299–312.
Chen JS, Wong VW, Gurtner GC. (2012) Therapeutic potential of bone marrow-derived mesenchymal stem cells for cutaneous wound healing. Front. Immunol. 3:192.
Singer NG, Caplan AI. (2011) Mesenchymal stem cells: mechanisms of inflammation. Annu. Rev. Pathol. 6:457–78.
Zgheib C. (2014) Fetal skin wound healing is characterized by a reduced inflammatory response. Adv. Wound Care (New Rochelle). 3:344–55.
Mori R, Kondo T, Ohshima T, Ishida Y, Mukaida N. (2002) Accelerated wound healing in tumor necrosis factor receptor p55-deficient mice with reduced leukocyte infiltration. FASEB J. 16:963–74.
Streit M, Beleznay Z, Braathen LR. (2006) Topical application of the tumour necrosis factor-alpha antibody infliximab improves healing of chronic wounds. Int. Wound J. 3:171–9.
Thomay AA, et al. (2009) Disruption of interleukin-1 signaling improves the quality of wound healing. Am. J. Pathol. 174:2129–36.
Gallucci RM, et al. (2001) Interleukin-6 treatment augments cutaneous wound healing in immunosuppressed mice. J. Interferon Cytokine Res. 21:603–9.
Lin ZQ, Kondo T, Ishida Y, Takayasu T, Mukaida N. (2003) Essential involvement of IL-6 in the skin wound-healing process as evidenced by delayed wound healing in IL-6-deficient mice. J. Leukoc. Biol. 73:713–21.
Yamasaki K, et al. (1998) Reversal of impaired wound repair in iNOS-deficient mice by topical adenoviral-mediated iNOS gene transfer. J. Clin. Invest. 101:967–71.
Most D, Efron DT, Shi HP, Tantry US, Barbul A. (2002) Characterization of incisional wound healing in inducible nitric oxide synthase knockout mice. Surgery. 132:866–76.
Park JE, Abrams MJ, Efron PA, Barbul A. (2013) Excessive nitric oxide impairs wound collagen accumulation. J. Surg. Res. 183:487–92.
Avlan D, Unlu A, Ayaz L, Camdeviren H, Nayci A, Aksoyek S. (2005) Poly (ADP-ribose) synthetase inhibition reduces oxidative and nitrosative organ damage after thermal injury. Pediatr. Surg. Int. 21:449–55.
Hassa PO, Covic M, Hasan S, Imhof R, Hottiger MO. (2001) The enzymatic and DNA binding activity of PARP-1 are not required for NF-kappa B coactivator function. J. Biol. Chem. 276:45588–97.
Kauppinen TM, Swanson RA. (2005) Poly(ADP-ribose) polymerase-1 promotes microglial activation, proliferation, and matrix metalloproteinase-9-mediated neuron death. J. Immunol. 174:2288–96.
Liu Y, et al. (2009) Increased matrix metalloproteinase-9 predicts poor wound healing in diabetic foot ulcers. Diabetes Care. 32:117–9.
Simbulan-Rosenthal CM, et al. (1996) The expression of poly(ADP-ribose) polymerase during differentiation-linked DNA replication reveals that it is a component of the multiprotein DNA replication complex. Biochemistry. 35:11622–33.
Eki T. (1994) Poly (ADP-ribose) polymerase inhibits DNA replication by human replicative DNA polymerase alpha, delta and epsilon in vitro. FEBS Lett. 356:261–6.
Pagano A, Metrailler-Ruchonnet I, Aurrand-Lions M, Lucattelli M, Donati Y, Argiroffo CB. (2007) Poly(ADP-ribose) polymerase-1 (PARP-1) controls lung cell proliferation and repair after hyperoxia-induced lung damage. Am. J. Physiol. Lung Cell. Mol. Physiol. 293: L619–29.
Rajesh M, et al. (2006) Pharmacological inhibition of poly(ADP-ribose) polymerase inhibits angiogenesis. Biochem. Biophys. Res. Commun. 350:352–7.
Tentori L, et al. (2007) Poly(ADP-ribose) polymerase (PARP) inhibition or PARP-1 gene deletion reduces angiogenesis. Eur. J. Cancer. 43:2124–33.
Pyriochou A, Olah G, Deitch EA, Szabo C, Papapetropoulos A. (2008) Inhibition of angiogenesis by the poly(ADP-ribose) polymerase inhibitor PJ-34. Int. J. Mol. Med. 22:113–8.
Mukhopadhyay P, et al. (2014) Poly (ADP-ribose) polymerase-1 is a key mediator of liver inflammation and fibrosis. Hepatology. 59:1998–2009.
Bai P, et al. (2009) Poly(ADP-ribose) polymerase mediates inflammation in a mouse model of contact hypersensitivity. J. Invest. Dermatol. 129:234–8.
Acknowledgments
This research was supported by the European Union and the State of Hungary, cofinanced by the European Social Fund in the framework of TAMOP 4.2.4.A/2-11/1-2012-0001 National Excellence Program. Direct costs of this study were supported by the Hungarian Science Research Fund (OTKA K82009, K112336 and K108308) and by the Faculty of Medicine, University of Debrecen (Bridging Fund).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, and provide a link to the Creative Commons license. You do not have permission under this license to share adapted material derived from this article or parts of it.
The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this license, visit (https://doi.org/creativecommons.org/licenses/by-nc-nd/4.0/)
About this article
Cite this article
El-Hamoly, T., Hegedűs, C., Lakatos, P. et al. Activation of Poly(ADP-Ribose) Polymerase-1 Delays Wound Healing by Regulating Keratinocyte Migration and Production of Inflammatory Mediators. Mol Med 20, 363–371 (2014). https://doi.org/10.2119/molmed.2014.00130
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2119/molmed.2014.00130