Abstract
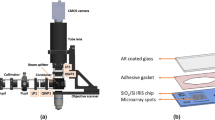
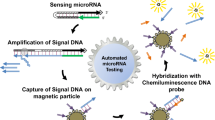
MicroRNAs (miRNAs) are attracting considerable attention as potential biomarkers for the early diagnosis of cancer. We have been developing a detection method for miRNAs on a microfluidic chip with external-power-free fluid pumping and enzyme-free amplification. The assay is completed within 20 min. Here, we describe the specificity of this miRNA detection method. First, the specificity against mismatched sequences was investigated. The nonspecific detection of a 2-nucleotide mismatched sequence was negligible, while that of a 1-nucleotide mismatched sequence was observed to a reasonable extent. Next, the disturbance in mature miRNA detection by existence of its precursor miRNA was evaluated. One precursor miRNA out of four tested showed significant nonspecific responses at 1 nM or higher concentrations. However, those responses were much lower than that of the target mature miRNA at 0.1 nM. Finally, we tried to detect three endogenous miRNAs, which are known to be potential cancer biomarkers, in human leucocyte total RNA. The measured concentraions of these miRNAs agreed well with those obtained by quantitative reverse transcription polymerase chain reaction. These results indicate that the on-chip miRNA detection method has good specificity, which is promising for applications to real biological samples.
Similar content being viewed by others
References
R. C. Friedman, K. K. H. Farh, C. B. Burge, and D. P. Bartel, Genome Res., 2009, 19, 92.
M. V. Iorio, M. Ferracin, C. G. Liu, A. Veronese, R. Spizzo, S. Sabbioni, E. Magri, M. Pedriali, M. Fabbri, M. Campiglio, S. Menard, J. P. Palazzo, A. Rosenberg, P. Musiani, S. Volinia, I. Nenci, G. A. Calin, P. Querzoli, M. Negrini, and C. M. Croce, Cancer Res., 2005, 65, 7065.
G. A. Calin and C. M. Croce, Nat. Rev. Cancer, 2006, 6, 857.
P. S. Mitchell, R. K. Parkin, E. M. Kroh, B. R. Fritz, S. K. Wyman, E. L. Pogosova-Agadjanyan, A. Peterson, J. Noteboom, K. C. O’Briant, A. Allen, D. W. Lin, N. Urban, C. W. Drescher, B. S. Knudsen, D. L. Stirewalt, R. Gentleman, R. L. Vessella, P. S. Nelson, D. B. Martin, and M. Tewari, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 10513.
J. A. Weber, D. H. Baxter, S. L. Zhang, D. Y. Huang, K. H. Huang, M. J. Lee, D. J. Galas, and K. Wang, Clin. Chem., 2010, 56, 1733.
A. Turchinovich, L. Weiz, A. Langheinz, and B. Burwinkel, Nucleic Acids Res., 2011, 39, 7223.
K. Wang, Y. Yuan, J. H. Cho, S. McClarty, D. Baxter, and D. J. Galas, PLoS One, 2012, 7, e41561
Y. Yamamoto, N. Kosaka, M. Tanaka, F. Koizumi, Y. Kanai, T. Mizutani, Y. Murakami, M. Kuroda, A. Miyajima, T. Kato, and T. Ochiya, Biomarkers, 2009, 14, 529.
N. Kosaka, H. Iguchi, and T. Ochiya, Cancer Sci., 2010, 101, 2087.
M. Tsujiura, D. Ichikawa, S. Komatsu, A. Shiozaki, H. Takeshita, T. Kosuga, H. Konishi, R. Morimura, K. Deguchi, H. Fujiwara, K. Okamoto, and E. Otsuji, Brit. J. Cancer, 2010, 102, 1174.
J. Zhou, L. Yu, X. Gao, J. Hu, J. P. Wang, Z. Dai, J. F. Wang, Z. Y. Zhang, S. H. Lu, X. W. Huang, Z. Wang, S. J. Qiu, X. Y. Wang, G. H. Yang, H. C. Sun, Z. Y. Tang, Y. Wu, H. G. Zhu, and J. Fan, J. Clin. Oncol., 2011, 29, 4781.
C. Zhu, C. Ren, J. Han, Y. Ding, J. Du, N. Dai, J. Dai, H. Ma, Z. Hu, H. Shen, Y. Xu, and G. Jin, Brit. J. Cancer, 2014, 110, 2291.
M. Baker, Nat. Methods, 2010, 7, 687.
B. N. Johnson and R. Mutharasan, Analyst, 2014, 139, 1576.
R. M. Graybill and R. C. Bailey, Anal. Chem., 2016, 88, 431.
T. Ueno and T. Funatsu, PLoS One, 2014, 9, e90920
H. Lee, R. L. Srinivas, A. Gupta, and P. S. Doyle, Angew. Chem., Int. Ed., 2015, 54, 2477.
S. Hofmann, Y. W. Huang, P. Paulicka, A. Kappel, H. A. Katus, A. Keller, B. Meder, C. F. Stahler, and W. Gumbrecht, Anal. Chem., 2015, 87, 12104.
H. Arata, H. Komatsu, A. Han, K. Hosokawa, and M. Maeda, Analyst, 2012, 137, 3234.
H. Arata, H. Komatsu, K. Hosokawa, and M. Maeda, PLoS One, 2012, 7, e48329.
R. Ishihara, K. Hasegawa, K. Hosokawa, and M. Maeda, Anal. Sci., 2015, 31, 573.
K. Hosokawa, K. Sato, N. Ichikawa, and M. Maeda, Lab Chip, 2004, 4, 181.
K. Hosokawa, M. Omata, and M. Maeda, Anal. Chem., 2007, 79, 6000.
K. Hosokawa and M. Maeda, Lab Chip, 2009, 9, 464.
R. I. Gregory and R. Shiekhattar, Cancer Res., 2005, 65, 3509.
“UNAFold” web site, http://unafold.rna.albany.edu/, accessed July 11, 2016.
“miRBase” web site, http://www.mirbase.org/index.shtml, accessed July 11, 2016.
Acknowledgments
This work was suppoerted by KAKENHI (25350581), JST Center of Innovation Program, and RIKEN Junior Research Associate Program.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Hasegawa, K., Negishi, R., Matsumoto, M. et al. Specificity of MicroRNA Detection on a Power-free Microfluidic Chip with Laminar Flow-assisted Dendritic Amplification. ANAL. SCI. 33, 171–177 (2017). https://doi.org/10.2116/analsci.33.171
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2116/analsci.33.171