Abstract
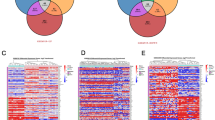
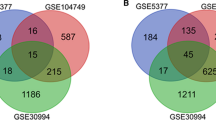
To identify novel genes in castration-resistant prostate cancer (CRPC), we downloaded three microarray datasets containing CRPC and primary prostate cancer in Gene Expression Omnibus (GEO). R packages affy and limma were performed to identify differentially expressed genes (DEGs) between primary prostate cancer and CRPC. After that, we performed functional enrichment analysis including gene ontology (GO) and Kyoto encyclopedia of genes and genomes (KEGG) pathway. In addition, protein-protein interaction (PPI) analysis was used to search for hub genes. Finally, to validate the significance of these genes, we performed survival analysis. As a result, we identified 53 upregulated genes and 58 downregulated genes that changed in at least two datasets. Functional enrichment analysis showed significant changes in the positive regulation of osteoblast differentiation pathway and aldosterone-regulated sodium reabsorption pathway. PPI network identified hub genes like cortactin-binding protein 2 (CTTNBP2), Rho family guanosine triphosphatase (GTPase) 3 (RND3), protein tyrosine phosphatase receptor-type R (PTPRR), Jagged1 (JAG1), and lumican (LUM). Based on PPI network analysis and functional enrichment analysis, we identified two genes (PTPRR and JAG1) as key genes. Further survival analysis indicated a relationship between high expression of the two genes and poor prognosis of prostate cancer. In conclusion, PTPRR and JAG1 are key genes in the CRPC, which may serve as promising biomarkers of diagnosis and prognosis of CRPC.
摘要
目的
鉴定去势抵抗性前列腺癌(CRPC)的关键基因.
创新点
(1)结合多个数据库数据,运用生物信息学方 法鉴定CRPC 的关键基因; (2)首次报道PTPRR 可能在CRPC 里起关键作用.
方法
通过下载三个GEO 数据库的mRNA 微阵列数据, 分析CRPC 和激素敏感前列腺癌之间的基因差 异,对筛选出的差异基因进行功能富集分析和蛋 白质间相互作用分析,最终筛选出两个有重要功 能的差异基因(PTPRR 和JAG1).通过在多个 其他数据库中进行表达量验证和生存分析,进一 步证明这些基因的重要作用.
结论
PTPRR 和JAG1 在CRPC 中显著增高,并与预后 差相关.因此,这两个基因有可能作为CRPC 的 诊断和预后的生物标志物.
Similar content being viewed by others
References
Artavanis-Tsakonas S, Rand MD, Lake RJ, 1999. Notch signaling: cell fate control and signal integration in development. Science, 284(5415):770–776. https://doi.org/10.1126/science.284.5415.770
Attard G, Parker C, Eeles RA, et al., 2016. Prostate cancer. Lancet, 387(10013):70–82. https://doi.org/10.1016/S0140-6736(14)61947-4
Chang CC, Huang RL, Wang HC, et al., 2014. High methylation rate of LMX1A, NKX6-1, PAX1, PTPRR, SOX1, and ZNF582 genes in cervical adenocarcinoma. Int J Gynecol Cancer, 24(2):201–209. https://doi.org/10.1097/igc.0000000000000054
Chen WQ, Zheng RS, Baade PD, et al., 2016. Cancer statistics in China, 2015. CA Cancer J Clin, 66(2):115–132. https://doi.org/10.3322/caac.21338
D’Antonio JM, Ma CQ, Monzon FA, et al., 2008. Longitudinal analysis of androgen deprivation of prostate cancer cells identifies pathways to androgen independence. Prostate, 68(7):698–714. https://doi.org/10.1002/pros.20677
Dennis G Jr., Sherman BT, Hosack DA, et al., 2003. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol, 4(5):P3. https://doi.org/10.1186/gb-2003-4-5-p3
Duś-Szachniewicz K, Woźniak M, Nelke K, et al., 2015. Protein tyrosine phosphatase receptor R and Z1 expression as independent prognostic indicators in oral squamous cell carcinoma. Head Neck, 37(12):1816–1822. https://doi.org/10.1002/hed.23835
Espinoza I, Pochampally R, Xing F, et al., 2013. Notch signaling: targeting cancer stem cells and epithelial-to-mesenchymal transition. Onco Targets Ther, 6:1249–1259. https://doi.org/10.2147/ott.s36162
Gautier L, Cope L, Bolstad BM, et al., 2004. affy—analysis of Affymetrix GeneChip data at the probe level. Bioinformatics, 20(3):307–315. https://doi.org/10.1093/bioinformatics/btg405
Gene Ontology Consortium, 2004. The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res, 32(S1):D258–D261. https://doi.org/10.1093/nar/gkh036
Guo YZ, Sun HH, Wang XT, et al., 2018. Transcriptomic analysis reveals key lncRNAs associated with ribosomal biogenesis and epidermis differentiation in head and neck squamous cell carcinoma. J Zhejiang Univ-Sci B (Biomed & Biotechnol), 19(9):674–688. https://doi.org/10.1631/jzus.B1700319
Heidenreich A, Bastian PJ, Bellmunt J, et al., 2014. EAU guidelines on prostate cancer. Part II: treatment of advanced, relapsing, and castration-resistant prostate cancer. Eur Urol, 65(2):467–479. https://doi.org/10.1016/j.eururo.2013.11.002
Kanehisa M, Goto S, 2000. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res, 28(1):27–30. https://doi.org/10.1093/nar/28.1.27
Kwon OJ, Zhang L, Wang JH, et al., 2016. Notch promotes tumor metastasis in a prostate-specific Pten-null mouse model. J Clin Invest, 126(7):2626–2641. https://doi.org/10.1172/jci84637
Laczmanska I, Karpinski P, Bebenek M, et al., 2013. Protein tyrosine phosphatase receptor-like genes are frequently hypermethylated in sporadic colorectal cancer. J Hum Genet, 58(1):11–15. https://doi.org/10.1038/jhg.2012.119
Li DM, Masiero M, Banham AH, et al., 2014. The Notch ligand Jagged1 as a target for anti-tumor therapy. Front Oncol, 4:254. https://doi.org/10.3389/fonc.2014.00254
Lin Y, Shen Z, Song X, et al., 2018. Comparative transcriptomic analysis reveals adriamycin-induced apoptosis via p53 signaling pathway in retinal pigment epithelial cells. J Zhejiang Univ-Sci B (Biomed & Biotechnol), 19(12):895–909. https://doi.org/10.1631/jzus.B1800408
Logothetis CJ, Lin SH, 2005. Osteoblasts in prostate cancer metastasis to bone. Nat Rev Cancer, 5(1):21–28. https://doi.org/10.1038/nrc1528
Menigatti M, Cattaneo E, Sabates-Bellver J, et al., 2009. The protein tyrosine phosphatase receptor type R gene is an early and frequent target of silencing in human colorectal tumorigenesis. Mol Cancer, 8:124. https://doi.org/10.1186/1476-4598-8-124
Munkley J, Lafferty NP, Kalna G, et al., 2015. Androgen-regulation of the protein tyrosine phosphatase PTPRR activates ERK1/2 signalling in prostate cancer cells. BMC Cancer, 15:9. https://doi.org/10.1186/s12885-015-1012-8
Noordman YE, Jansen PAM, Hendriks WJAJ, 2006. Tyrosine-specific MAPK phosphatases and the control of ERK signaling in PC12 cells. J Mol Signal, 1:4. https://doi.org/10.1186/1750-2187-1-4
Ritchie ME, Phipson B, Wu D, et al., 2015. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res, 43(7):e47. https://doi.org/10.1093/nar/gkv007
Santagata S, Demichelis F, Riva A, et al., 2004. JAGGED1 expression is associated with prostate cancer metastasis and recurrence. Cancer Res, 64(19):6854–6857. https://doi.org/10.1158/0008-5472.can-04-2500
Schmitt I, Bitoun E, Manto M, 2009. PTPRR, cerebellum, and motor coordination. Cerebellum, 8(2):71–73. https://doi.org/10.1007/s12311-009-0118-4
Sethi N, Dai XD, Winter CG, et al., 2011. Tumor-derived Jagged1 promotes osteolytic bone metastasis of breast cancer by engaging Notch signaling in bone cells. Cancer Cell, 19(2):192–205. https://doi.org/10.1016/j.ccr.2010.12.022
Shannon P, Markiel A, Ozier O, et al., 2003. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res, 13(11):2498–2504. https://doi.org/10.1101/gr.1239303
Shou JY, Ross S, Koeppen H, et al., 2001. Dynamics of Notch expression during murine prostate development and tumorigenesis. Cancer Res, 61(19):7291–7297.
Siegel RL, Miller KD, Jemal A, 2019. Cancer statistics, 2019. CA Cancer J Clin, 69(1):7–34. https://doi.org/10.3322/caac.21551
Su L, Song X, Xue Z, et al., 2018. Network analysis of microRNAs, transcription factors, and target genes involved in axon regeneration. J Zhejiang Univ-Sci B (Biomed & Biotechnol), 19(4):293–304. https://doi.org/10.1631/jzus.B1700179
Su PH, Lin YW, Huang RL, et al., 2013. Epigenetic silencing of PTPRR activates MAPK signaling, promotes metastasis and serves as a biomarker of invasive cervical cancer. Oncogene, 32(1):15–26. https://doi.org/10.1038/onc.2012.29
Su Q, Zhang B, Zhang L, et al., 2017. Jagged1 upregulation in prostate epithelial cells promotes formation of reactive stroma in the Pten null mouse model for prostate cancer. Oncogene, 36(5):618–627. https://doi.org/10.1038/onc.2016.232
Sun YT, Wang BE, Leong KG, et al., 2012. Androgen deprivation causes epithelial-mesenchymal transition in the prostate: implications for androgen-deprivation therapy. Cancer Res, 72(2):527–536. https://doi.org/10.1158/0008-5472.can-11-3004
Szklarczyk D, Morris JH, Cook H, et al., 2017. The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res, 45(D1):D362–D368. https://doi.org/10.1093/nar/gkw937
Tang ZF, Li CW, Kang BX, et al., 2017. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res, 45(W1):W98–W102. https://doi.org/10.1093/nar/gkx247
Terada N, Shimizu Y, Kamba T, et al., 2010. Identification of EP4 as a potential target for the treatment of castration-resistant prostate cancer using a novel xenograft model. Cancer Res, 70(4):1606–1615. https://doi.org/10.1158/0008-5472.can-09-2984
Watson PA, Arora VK, Sawyers CL, 2015. Emerging mechanisms of resistance to androgen receptor inhibitors in prostate cancer. Nat Rev Cancer, 15(12):701–711. https://doi.org/10.1038/nrc4016
Weinstein JN, Collisson EA, Mills GB, et al., 2013. The Cancer Genome Atlas Pan-Cancer analysis project. Nat Genet, 45(10):1113–1120. https://doi.org/10.1038/ng.2764
Woźniak M, Gamian E, Łaczmańska I, et al., 2014. Immunohistochemical and Western blot analysis of two protein tyrosine phosphatase receptors, R and Z1, in colorectal carcinoma, colon adenoma and normal colon tissues. Histol Histopathol, 29(5):635–639. https://doi.org/10.14670/HH-29.10.635
Yong T, Sun A, Henry MD, et al., 2011. Down regulation of CSL activity inhibits cell proliferation in prostate and breast cancer cells. J Cell Biochem, 112(9):2340–2351. https://doi.org/10.1002/jcb.23157
Zhu H, Zhou XC, Redfield S, et al., 2013. Elevated Jagged-1 and Notch-1 expression in high grade and metastatic prostate cancers. Am J Transl Res, 5(3):368–378.
Zhu H, Li Y, Wang M, et al., 2019. Analysis of cardiovascular disease-related NF-κB-regulated genes and microRNAs in TNFα-treated primary mouse vascular endothelial cells. J Zhejiang Univ-Sci B (Biomed & Biotechnol), 20(10):803–815. https://doi.org/10.1631/jzus.B1800631
Acknowledgments
The authors would like to thank The Cancer Genome Atlas (TCGA) and the Gene Expression Omnibus (GEO) programs for providing high quality data.
Author information
Authors and Affiliations
Contributions
Guo-ping REN and Ji-li WANG conceived and designed the study. Ji-li WANG and Yan WANG collected the data and analyzed the data. Ji-li WANG wrote the original draft. Guo-ping REN and Yan WANG reviewed and edited the manuscript. All authors have read and approved the final manuscript. All authors have full access to all the data in the study and have responsibility for the integrity and security of the data.
Corresponding author
Additional information
List of electronic supplementary materials
Table S1 Clinical characteristics of samples in this study Fig. S1 Expression of the PTPRR in Grasso PCa
Data availability
The data that support the findings of this study are openly available in Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo), and the code for analysis in this article can be obtained by emailing the corresponding author.
Compliance with ethics guidelines
Ji-li WANG, Yan WANG, and Guo-ping REN declare that they have no conflict of interest.
This article does not contain any studies with human or animal subjects performed by any of the authors.
Electronic supplementary materials
11585_2020_414_MOESM1_ESM.pdf
Identification of PTPRR and JAG1 as key genes in castration-resistant prostate cancer by integrated bioinformatics methods
Rights and permissions
About this article
Cite this article
Wang, Jl., Wang, Y. & Ren, Gp. Identification of PTPRR and JAG1 as key genes in castration-resistant prostate cancer by integrated bioinformatics methods. J. Zhejiang Univ. Sci. B 21, 246–255 (2020). https://doi.org/10.1631/jzus.B1900329
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1631/jzus.B1900329
Key words
- Bioinformatics
- Protein tyrosine phosphatase receptor-type R (PTPRR)
- Jagged1 (JAG1)
- Differentially expressed genes (DEGs)
- Castration-resistant prostate cancer (CRPC)
- Functional enrichment