Abstract
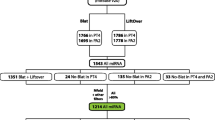
Mutation (substitution, deletion, insertion, etc.) in nucleotide acid causes the maximal sequence lengths of exact match (MALE) between paralogous members from a duplicate event to become shorter during evolution. In this work, MALE changes between members of 26 gene families from four representative species (Arabidopsis thaliana, Oryza sativa, Mus musculus and Homo sapiens) were investigated. Comparative study of paralogous’ MALE and amino acid substitution rate (dA<0.5) indicated that a close relationship existed between them. The results suggested that MALE could be a sound evolutionary scale for the divergent time for paralogous genes during their early evolution. A reference table between MALE and divergent time for the four species was set up, which would be useful widely, for large-scale genome alignment and comparison. As an example, detection of large-scale duplication events of rice genome based on the table was illustrated.
Similar content being viewed by others
References
Arratia, R., Gordon, L., Waterman, M.S., 1986. An extreme value theory for sequence matching.Ann. Stat.,14:971–993.
Blane, G., Wolfe, K.H., 2004. Widespread paleopolyploidy in model plant species inferred from age distributions of duplicate genes.Plant Cell,16:1667–1678.
Brown, T.A., 1999. Genomes. BIOS Scientific Publishers Limited, Oxford.
Delcher, A.L., Phillippy, A., Carlton, J., Salzberg, S.L., 2002. Fast algorithms for large-scale genome alignment and comparison.Nucleic Acids Res.,30:2478–2483.
Goff, S.A., Ricke, D., Lan, T.H., Presting, G., Wang, R., Dunn, M., Glazebrook, J., Sessions, A., Oeller, P., Varma, H.,et al., 2002. A draft sequence of the rice genome (Oryza sativa L. ssp.japonica).Science,296:92–100.
Karlin, S., Altschul, S.F., 1990. Methods for assessing the statistical significance of molecular sequence features by using general scoring schemes.Proc. Natl. Acad. Sci. USA,87(6):2264–2268.
Li, W.H., 1997. Molecular Evolution. Sinauer Associates. Sunderland, MA.
Lynch, M., Conery, J.S., 2000. The evolutionary fate and consequences of duplicate genes.Science,290: 1151–1155.
Ohno, S., 1970. Evolution by Gene Duplication. George Allen and Unwin, London.
Paterson, A.H., Bowers, J.E., Chapman, B.A., 2004. Ancient polyploidization predating divergence of the cereals, and its consequences for comparative genomics.Proc. Natl. Acad. Sci. USA,101:9903–9908.
Yang, Z., 1999. Phylogenetic Analysis by Maximum Likelihood (PAML). Version 2., London, UK.
Author information
Authors and Affiliations
Corresponding author
Additional information
Project supported by the National Natural Science Foundation of China (Grant Nos. 30270810, 90208022 and 30471067) and IBM Shared University Research (Life Science), China
Rights and permissions
About this article
Cite this article
Wen, X., Guo, Xy. & Fan, Lj. Maximal sequence length of exact match between members from a gene family during early evolution. J Zheijang Univ Sci B 6, 470–476 (2005). https://doi.org/10.1631/jzus.2005.B0470
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1631/jzus.2005.B0470
Key words
- Maximal length of exact match (MALE)
- Divergent time
- Gene family
- Minimal length of exact match (MILE)
- Genome alignment