Abstract
X-ray Bragg coherent diffraction imaging is a powerful technique for operando and in situ materials characterization and provides a unique means of quantifying the influence of one-dimensional (1D) and two-dimensional (2D) material defects on material response. However, obtaining full images from raw x-ray diffraction data is nontrivial and computationally intensive, precluding real-time experimental feedback. Here, we present a machine learning approach to identify the presence of crystalline line defects (edge and screw) in samples from the raw, 2D, coherent diffraction data without the need for image reconstruction through iterative phase retrieval. We compare different approaches to designing neural networks for this application and demonstrate the potential of automated ML (autoML) approaches.
Impact statement
The need for automated processing of coherent diffraction data is strongly motivated by the advent of fourth-generation synchrotron x-ray sources, where coherent diffraction data will be generated at a tremendous rate and human interaction with data analysis, and especially iterative phase retrieval image reconstruction, will become untenable. Our approach provides a path to dealing with this necessary improvement in data processing efficiency. We expect that this work, which demonstrates the applicability of automated machine learning to x-ray analysis, will be of broad interest to scientists and users of synchrotron and XFEL facilities.
Graphical abstract





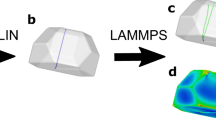
taken from the starting array. (c) Represents a simulated perfect edge defect placed into the structure. (d) The LAMMPS relaxed version of (c, e) and (g) are examples of the simulated 3D Bragg coherent diffraction imaging (BCD) patterns for the given simulated crystals. (f, h) Two-dimensional central slices of the simulated 3D BCDI patterns.
Similar content being viewed by others
Data availability
Trained networks, code, raw data, and tutorials can be found at https://github.com/WilliamJudge94/CDI_AI. In turn, tutorials on the crystal simulations, LAMMPS relaxation, and PyNX simulations can be found at https://crystal-simulation.readthedocs.io/en/latest/index.html.
Code availability
Trained networks, code, raw data, and tutorials can be found at https://github.com/WilliamJudge94/CDI_AI. In turn, tutorials on the crystal simulations, LAMMPS relaxation, and PyNX simulations can be found at https://crystal-simulation.readthedocs.io/en/latest/index.html .
References
A. Singer, M. Zhang, S. Hy, D. Cela, C. Fang, T.A. Wynn, B. Qiu, Y. Xia, Z. Liu, A. Ulvestad, N. Hua, J. Wingert, H. Liu, M. Sprung, A.V. Zozulya, E. Maxey, R. Harder, Y.S. Meng, O.G. Shpyrko, Nucleation of dislocations and their dynamics in layered oxide cathode materials during battery charging. Nat. Energy 3, 641 (2018)
H. Guo, Z. Wei, K. Jia, B. Qiu, C. Yin, F. Meng, Q. Zhang, L. Gu, S. Han, Y. Liu, H. Zhao, W. Jiang, H. Cui, Y. Xia, Z. Liu, Abundant nanoscale defects to eliminate voltage decay in Li-rich cathode materials. Energy Storage Mater. 16, 220 (2019)
D. Kim, M. Chung, S. Kim, K. Yun, W. Cha, R. Harder, H. Kim, Defect dynamics at a single Pt nanoparticle during catalytic oxidation. Nano Lett. 19, 5044 (2019)
R. Chattot, P. Bordet, I. Martens, J. Drnec, L. Dubau, F. Maillard, Building practical descriptors for defect engineering of electrocatalytic materials. ACS Catal. 10, 9046 (2020)
X. Shi, R. Harder, Z. Liu, O. Shpyrko, E. Fullerton, B. Kiefer, E. Fohtung, Nanoscale mapping of heterogeneous strain and defects in individual magnetic nanocrystals. Curr. Comput. Aided Drug Des. 10, 658 (2020). https://doi.org/10.3390/cryst10080658
M. Dupraz, G. Beutier, D. Rodney, D. Mordehai, M. Verdier, Signature of dislocations and stacking faults of face-centred cubic nanocrystals in coherent x-ray diffraction patterns: A numerical study. J. Appl. Crystallogr. 48, 621 (2015)
J. Wang, L. Liu, C. Chen, X. Dong, Q. Wang, L. Alfilfil, M.R. AlAlouni, K. Yao, J. Huang, D. Zhang, Y. Han, Engineering effective structural defects of metal–organic frameworks to enhance their catalytic performances. J. Mater. Chem. A 8, 4464 (2020)
L. David, R.E. Ruther, D. Mohanty, H.M. Meyer, Y. Sheng, S. Kalnaus, C. Daniel, D.L. Wood, Identifying degradation mechanisms in lithium-ion batteries with coating defects at the cathode. Appl. Energy 231, 446 (2018)
J. Ren, M. Ledwaba, N.M. Musyoka, H.W. Langmi, M. Mathe, S. Liao, W. Pang, Structural defects in metal–organic frameworks (MOFs): Formation, detection and control towards practices of interests. Coord. Chem. Rev. 349, 169 (2017)
U. Ruett, J. Almer, P. Kenesei, J.-S. Park, R. Osborn, Y. Ren, D. Robinson, M. Krogstad, S. Rosenkranz, X. Zhang, M. Li, K. Wiaderek, APS: High-energy x-rays expediting applied and fundamental research. Synchrotron Radiat. News 33, 44 (2020). https://doi.org/10.1080/08940886.2020.1841498
N. Heidenreich, S. Waitschat, H. Reinsch, Investigation of the kinetic stabilization of a Ce4+-based MOF by in-situ powder x-ray diffraction. Z. Anorg. Allg. Chem. 644, 1826 (2018)
M. Holt, R. Harder, R. Winarski, V. Rose, Nanoscale hard x-ray microscopy methods for materials studies. Annu. Rev. Mater. Res. 43, 183 (2013)
F. Rovaris, M.H. Zoellner, P. Zaumseil, M.A. Schubert, A. Marzegalli, L. Di Gaspare, M. De Seta, T. Schroeder, P. Storck, G. Schwalb, C. Richter, T.U. Schulli, G. Capellini, F. Montalenti, Misfit-dislocation distributions in heteroepitaxy: From mesoscale measurements to individual defects and back. Phys. Rev. Appl. 10, 054067 (2018)
M. Kodur, R.E. Kumar, Y. Luo, D.N. Cakan, X. Li, M. Stuckelberger, D.P. Fenning, X-ray microscopy of halide perovskites: Techniques, applications, and prospects. Adv. Energy Mater. 10, 1903170 (2020)
M. Meduna, F. Isa, A. Jung, A. Marzegalli, M. Albani, G. Isella, K. Zweiacker, L. Miglio, H. von Kanel, Lattice tilt and strain mapped by x-ray scanning nanodiffraction in compositionally graded SiGe/Si microcrystals. J. Appl. Crystallogr. 51, 368 (2018)
Y. Takahashi, A. Suzuki, S. Furutaku, K. Yamauchi, Y. Kohmura, T. Ishikawa, Bragg x-ray ptychography of a silicon crystal: Visualization of the dislocation strain field and the production of a vortex beam. Phys. Rev. B 87, 121201 (2013)
V.L.R. Jacques, S. Ravy, D. Le Bolloc’h, E. Pinsolle, M. Sauvage-Simkin, F. Livet, Bulk dislocation core dissociation probed by coherent x rays in silicon. Phys. Rev. Lett. 106, 065502 (2011)
J. Miao, Coherent diffraction imaging. Microsc. Microanal. 20, 368 (2014)
M.J. Cherukara, R. Pokharel, T.S. O’Leary, J.K. Baldwin, E. Maxey, W. Cha, J. Maser, R.J. Harder, S.J. Fensin, R.L. Sandberg, Three-dimensional x-ray diffraction imaging of dislocations in polycrystalline metals under tensile loading. Nat. Commun. 9, 3776 (2018)
A. Ulvestad, Y. Nashed, G. Beutier, M. Verdier, S.O. Hruszkewycz, M. Dupraz, Identifying defects with guided algorithms in Bragg coherent diffractive imaging. Sci. Rep. 7, 9920 (2017)
J.R. Fienup, Phase retrieval algorithms: A comparison. Appl. Opt. 21, 2758 (1982)
V. Elser, Phase retrieval by iterated projections. J. Opt. Soc. Am. A 20, 40 (2003)
M.J. Cherukara, Y.S.G. Nashed, R.J. Harder, Real-time coherent diffraction inversion using deep generative networks. Sci. Rep. 8(1), 16520 (2018)
L. Wu, P. Juhas, S. Yoo, I. Robinson, Complex imaging of phase domains by deep neural networks. IUCrJ 8, 12 (2021)
H. Chan, Y.S.G. Nashed, S. Kandel, S.O. Hruszkewycz, S.K.R.S. Sankaranarayanan, R.J. Harder, M.J. Cherukara, Rapid 3D nanoscale coherent imaging via physics-aware deep learning. Appl. Phys. Rev. 8, 021407 (2021). https://doi.org/10.1063/5.0031486
A. Scheinker, R. Pokharel, Adaptive 3D convolutional neural network-based reconstruction method for 3D coherent diffraction imaging. J. Appl. Phys. 128, 184901 (2020). https://doi.org/10.1063/5.0014725
L. Wu, S. Yoo, A.F. Suzana, T. A. Assefa, R.J. Harder, W. Cha, I.K. Robinson, 3D coherent x-ray imaging via deep convolutional neural networks. arXiv:abs/2103.00001 (2021)
B. Lim, E. Bellec, M. Dupraz, S. Leake, A. Resta, A. Coati, M. Sprung, E. Almog, E. Rabkin, T. Schulli, M.I. Richard, A convolutional neural network for defect classification in Bragg coherent x-ray diffraction. NPJ Comput. Mater. 7, 115 (2021)
O. Okwuashi, C.E. Ndehedehe, Deep support vector machine for hyperspectral image classification. Pattern Recognit. 103, 107298 (2020)
M. Sheykhmousa, M. Mahdianpari, H. Ghanbari, F. Mohammadimanesh, P. Ghamisi, S. Homayouni, Support vector machine versus random forest for remote sensing image classification: A meta-analysis and systematic review. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 13, 6308 (2020)
S.B. Kotsiantis, I.D. Zaharakis, P.E. Pintelas, Machine learning: A review of classification and combining techniques. Artif. Intell. Rev. 26, 159 (2006)
T.J. Brinker, A. Hekler, A.H. Enk, C. Berking, S. Haferkamp, A. Hauschild, M. Weichenthal, J. Klode, D. Schadendorf, T. Holland-Letz, C. von Kalle, S. Frohling, B. Schilling, J.S. Utikal, Deep neural networks are superior to dermatologists in melanoma image classification. Eur. J. Cancer 119, 11 (2019)
P.K. Mallick, S.H. Ryu, S.K. Satapathy, S. Mishra, G.N. Nguyen, P. Tiwari, Brain MRI image classification for cancer detection using deep wavelet autoencoder-based deep neural network. IEEE Access 7, 46278 (2019)
K. He, X. Zhang, S. Ren, J. Sun, “Deep Residual Learning for Image Recognition,” in Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition (2016), p. 770
S. Yu, S. Jia, C. Xu, Convolutional neural networks for hyperspectral image classification. Neurocomputing 219, 88 (2017)
K. Weiss, T.M. Khoshgoftaar, D.D. Wang, A survey of transfer learning. J. Big Data 3, 9 (2016)
K. Simonyan, A. Zisserman, “Very Deep Convolutional Networks for Large-Scale Image Recognition,” in 3rd International Conference on Learning Representations, ICLR 2015 - Conference Track Proceedings, vol. 1 (2015), arXiv:abs/1409.1556
G. Zeng, Y. He, Z. Yu, X. Yang, R. Yang, L. Zhang, Preparation of novel high copper ions removal membranes by embedding organosilane-functionalized multiwalled carbon nanotube. J. Chem. Technol. Biotechnol. 91, 2322 (2016)
K. Jing, J. Xu, H.X. Zugeng, “NASABN: A Neural Architecture Search Framework for Attention-Based Networks,” 2020 International Joint Conference on Neural Networks (IJCNN) (2020), p. 1
L. Guilin, Z. Xing, W. Zitong, L. Zhenguo, Z. Tong, Stacnas: Towards stable and consistent optimization for differentiable neural architecture search, arXiv, 1 (2019), arXiv:1909.11926v4
H. Jin, Q. Song, X. Hu, “Auto-Keras: An Efficient Neural Architecture Search System,” in Proceedings of the 25th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining (Association for Computing Machinery, 2019), p. 1946
H. Qassim, A. Verma, D. Feinzimer, “Compressed Residual-VGG16 CNN Model for Big Data Places Image Recognition,” 2018 IEEE 8th Annual Computing and Communication Workshop and Conference (CCWC) (2018), p. 169
K.-S. Lee, S.-K. Jung, J.-J. Ryu, S.-W. Shin, J. Choi, Evaluation of transfer learning with deep convolutional neural networks for screening osteoporosis in dental panoramic radiographs. J. Clin. Med. 9, 392 (2020)
Z. Liu, T. Bicer, R. Kettimuthu, D. Gursoy, F. De Carlo, I. Foster, TomoGAN: Low-dose synchrotron x-ray tomography with generative adversarial networks: Discussion. J. Opt. Soc. Am. A 37, 422 (2020)
V. Dung, L.D. Anh, Autonomous concrete crack detection using deep fully convolutional neural network. Autom. Constr. 99, 52 (2019)
A. Krishnaswamy Rangarajan, R. Purushothaman, Disease classification in eggplant using pre-trained VGG16 and MSVM. Sci. Rep. 10(1), 2322 (2020)
X. Yu, W. Pang, Q. Xu, M. Liang, Mammographic image classification with deep fusion learning. Sci. Rep. 10, 14631 (2020)
S. Matsuda, T. Miyamoto, H. Yoshimura, T. Hasegawa, Personal identification with orthopantomography using simple convolutional neural networks: A preliminary study. Sci. Rep. 10, 13559 (2020)
M. Christopher, A. Belghith, C. Bowd, J.A. Proudfoot, M.H. Goldbaum, R.N. Weinreb, C.A. Girkin, J.M. Liebmann, L.M. Zangwill, Performance of deep learning architectures and transfer learning for detecting glaucomatous optic neuropathy in fundus photographs. Sci. Rep. 8(1), 16685 (2018)
Q. Yu, Y. Yang, F. Liu, Y.Z. Song, T. Xiang, T.M. Hospedales, Sketch-a-Net: A deep neural network that beats humans. Int. J. Comput. Vis. 122, 411 (2017)
P. Williams, “Demystifying Deep Convolutional Neural Networks for Sonar Image Classification,” in Proceedings of the 4th Underwater Acoustics Conference (2019), vol. 3, p. 513
R. Yan, F. Ren, Z. Wang, L. Wang, T. Zhang, Y. Liu, X. Rao, C. Zheng, F. Zhang, Breast cancer histopathological image classification using a hybrid deep neural network. Methods 173, 52 (2020)
L. Alzubaidi, J. Zhang, A.J. Humaidi, A. Al-Dujaili, Y. Duan, O. Al-Shamma, J. Santamaría, M.A. Fadhel, M. Al-Amidie, L. Farhan, Review of deep learning: Concepts, CNN architectures, challenges, applications, future directions. J. Big Data 8, 53 (2021)
M. Abadi, A. Agarwal, P. Barham, E. Brevdo, Z. Chen, C. Citro, G.S. Corrado, A. Davis, J. Dean, M. Devin, S. Ghemawat, I. Goodfellow, A. Harp, G. Irving, M. Isard, Y. Jia, R. Jozefowicz, L. Kaiser, M. Kudlur, J. Levenberg, D. Mane, R. Monga, S. Moore, D. Murray, C. Olah, M. Schuster, J. Shlens, B. Steiner, I. Sutskever, K. Talwar, P. Tucker, V. Vanhoucke, V. Vasudevan, F. Viegas, O. Vinyals, P. Warden, M. Wattenberg, M. Wicke, Y. Yu, X. Zheng, TensorFlow: Large-scale machine learning on heterogeneous systems (2015). http://tensorflow.org
P. Hirel, Atomsk: A tool for manipulating and converting atomic data files. Comput. Phys. Commun. 197, 212 (2015)
V. Favre-Nicolin, G. Girard, S. Leake, J. Carnis, Y. Chushkin, J. Kieffer, P. Paleo, M.-I. Richard, PyNX: High-performance computing toolkit for coherent x-ray imaging based on operators. J. Appl. Crystallogr. 53, 1404 (2020)
V. Favre-Nicolin, J. Coraux, M.-I. Richard, H. Renevier, Fast computation of scattering maps of nanostructures using graphical processing units. J. Appl. Crystallogr. 44, 635 (2011)
O. Mandula, M. Elzo Aizarna, J. Eymery, M. Burghammer, V. Favre-Nicolin, PyNX.Ptycho: A computing library for x-ray coherent diffraction imaging of nanostructures. J. Appl. Crystallogr. 49, 1842 (2016)
S. Plimpton, Fast parallel algorithms for short-range molecular dynamics. J. Comput. Phys. 117(1), 1 (1995)
S.O. Hruszkewycz, M.V. Holt, M. Allain, V. Chamard, S.M. Polvino, C.E. Murray, P.H. Fuoss, Efficient modeling of Bragg coherent x-ray nanobeam diffraction. Opt. Lett. 40, 3241 (2015)
D.B. Williams, C.B. Carter, “The Transmission Electron Microscope,” in Transmission Electron Microscopy: A Textbook for Materials Science (Springer, Boston, 1996), p. 3
Y. Xie, D. Richmond, Pre-training on grayscale imagenet improves medical image classification, Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics) 11134 LNCS (2019), p. 476
M. Talo, U.B. Baloglu, O. Yıldırım, U. Rajendra Acharya, Application of deep transfer learning for automated brain abnormality classification using MR images. Cogn. Syst. Res. 54, 176 (2019)
J.W. Lee, W.B. Park, J.H. Lee, S.P. Singh, K.-S. Sohn, A deep-learning technique for phase identification in multiphase inorganic compounds using synthetic XRD powder patterns. Nat. Commun. 11, 86 (2020)
T. Holm-Jensen, T.M. Hansen, Linear waveform tomography inversion using machine learning algorithms. Math. Geosci. 52, 31 (2020)
M.J. Cherukara, T. Zhou, Y. Nashed, P. Enfedaque, A. Hexemer, R.J. Harder, M.V. Holt, AI-enabled high-resolution scanning coherent diffraction imaging. Appl. Phys. Lett. 117, 044103 (2020). https://doi.org/10.1063/5.0013065
O. Furat, M. Wang, M. Neumann, L. Petrich, M. Weber, C.E. Krill, V. Schmidt, Machine learning techniques for the segmentation of tomographic image data of functional materials. Front. Mater. 6, 145 (2019). https://doi.org/10.3389/fmats.2019.00145
Funding
We gratefully acknowledge the computing resources provided and operated by the Joint Laboratory for System Evaluation (JLSE) at Argonne National Laboratory. This work was performed at the Center for Nanoscale Materials and the Advanced Photon Source. The use of the Center for Nanoscale Materials and Advanced Photon Source, both Office of Science user facilities, was supported by the US Department of Energy (DOE), Office of Science, Office of Basic Energy Sciences, under Contract No. DE-AC02- 06CH11357. This work was also supported by Argonne LDRD 2018–019-N0: A.I C.D.I: Atomistically Informed Coherent Diffraction Imaging and by the US Department of Energy, Office of Science, Office of Basic Energy Sciences Data, Artificial Intelligence and Machine Learning at DOE Scientific User Facilities program under Award No. 34532.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Consent given by all authors.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Judge, W., Chan, H., Sankaranarayanan, S. et al. Defect identification in simulated Bragg coherent diffraction imaging by automated AI. MRS Bulletin 48, 124–133 (2023). https://doi.org/10.1557/s43577-022-00342-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1557/s43577-022-00342-1