Abstract
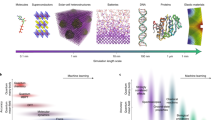
Machine learning (ML) has revolutionized disciplines within materials science that have been able to generate sufficiently large datasets to utilize algorithms based on statistical inference, but for many important classes of materials the datasets remain small. However, a rapidly growing number of approaches to embedding domain knowledge of materials systems are reducing data requirements and allowing broader applications of ML. Furthermore, these hybrid approaches improve the interpretability of the predictions, allowing for greater physical insights into the factors that determine material properties. This review introduces a number of these strategies, providing examples of how they were implemented in ML algorithms and discussing the materials systems to which they were applied.











Similar content being viewed by others
References
C. Kittel: Physical theory of ferromagnetic domains. Rev. Mod. Phys. 21, 541 (1949).
P.J. Flory: Molecular theory of rubber elasticity. Polym. J. 17, 1 (1985).
J.J. Stickel and R.L. Powell: Fluid mechanics and rheology of dense suspensions. Annu. Rev. Fluid Mech. 37, 129 (2005).
B.L. DeCost, T. Francis, and E.A. Holm: Exploring the microstructure manifold: image texture representations applied to ultrahigh carbon steel microstructures. Acta Mater 133, 30 (2017).
K. Saravanan, J.R. Kitchin, O.A. von Lilienfeld, and J.A. Keith: Alchemical predictions for computational catalysis: potential and limitations. J. Phys. Chem. Lett. 8, 5002 (2017).
R. Ramprasad, R. Batra, G. Pilania, A. Mannodi-Kanakkithodi, and C. Kim: Machine learning in materials informatics: recent applications and prospects. NPJ Comput. Mater. 3, 54 (2017).
A. Jain, S.P. Ong, G. Hautier, W. Chen, W.D. Richards, S. Dacek, S. Cholia, D. Gunter, D. Skinner, G. Ceder, and K.A. Persson: Commentary: The Materials Project: a materials genome approach to accelerating materials innovation. APL Mater. 1, 011002 (2013).
D.L. McDowell and S.R. Kalidindi: The materials innovation ecosystem: a key enabler for the Materials Genome Initiative. MRS Bull. 41, 326 (2016).
M. Qin, Z. Lin, Z. Wei, B. Zhu, J. Yuan, I. Takeuchi, and K. Jin: High-throughput research on superconductivity. Chinese Phys. B 27, 127402 (2018).
T.Z.H. Gani and H.J. Kulik: Understanding and breaking scaling relations in single-site catalysis: Methane to methanol conversion by Fe IV O. ACS Catal. 8, 975 (2018).
S. Ramakrishna, T.Y. Zhang, W.-C. Lu, Q. Qian, J.S.C. Low, J.H.R. Yune, D.Z.L. Tan, S. Bressan, S. Sanvito, and S.R. Kalidindi: Materials informatics. J. Intell. Manuf (2018). https://doi.org/10.1007/s10845-018-1392-0
M. McBride, N. Persson, E. Reichmanis, M. Grover, M. McBride, N. Persson, E. Reichmanis, and M.A. Grover: Solving materials’ small data problem with dynamic experimental databases. Processes 6, 79 (2018).
R. Kuhne, R.-U. Ebert, and G. Schuurmann: Model selection based on structural similarity-method description and application to water solubility prediction. J. Chem. Inf. Model. 46, 636 (2006).
L.D. Hughes, D.S. Palmer, F. Nigsch, and J.B.O. Mitchell: Why are some properties more difficult to predict than others? A study of QSPR models of solubility, melting point, and log P. J. Chem. Inf. Model. 48, 220 (2008).
B. Sanchez-Lengeling, L.M. Roch, J.D. Perea, S. Langner, C.J. Brabec, and A. Aspuru-Guzik: A Bayesian approach to predict solubility parameters. Adv. Theory Simul 2, 1 (2019).
B. Meredig, A. Agrawal, S. Kirklin, J.E. Saal, J.W. Doak, A. Thompson, K. Zhang, A. Choudhary, and C. Wolverton: Combinatorial screening for new materials in unconstrained composition space with machine learning. Phys. Rev. B 89, 094104 (2014).
K. Hansen, F. Biegler, R. Ramakrishnan, W. Pronobis, O.A. von Lilienfeld, K.-R. Müller, and A. Tkatchenko: Machine learning predictions of molecular properties: accurate many-body potentials and nonlocality in chemical space. J. Phys. Chem. Lett. 6, 2326 (2015).
Y. Liu, T. Zhao, W. Ju, and S. Shi: Materials discovery and design using machine learning. J. Mater. 3, 159 (2017).
R.C. Rowe and E.A. Colbourn: Neural computing in product formulation. Chem. Educ. 8, 1 (2003).
M. Tanco, E. Viles, L. Ilzarbe, and M.J. Alvarez: Implementation of design of experiments projects in industry. Appl. Stoch. Model. Bus. Ind. 25, 478 (2009).
D.C. Montgomery: Design and Analysis of Experiments. 8th ed. (Wiley, New York, 2012).
M.I. Jordan and T.M. Mitchell: Machine learning: trends, perspectives, and prospects. Science 349, 255 (2015).
H.A. Haenssle, C. Fink, R. Schneiderbauer, F. Toberer, T. Buhl, A. Blum, A. Kalloo, A. Ben Hadj Hassen, L. Thomas, A. Enk, L. Uhlmann, and m.A. Holger Haenssle: Man against machine: diagnostic performance of a deep learning convolutional neural network for dermoscopic melanoma recognition in comparison to 58 dermatologists. Ann. Oncol. 29, 1836 (2018).
T.L. Griffiths, E.R. Baraff, and J.B. Tenenbaum: Using physical theories to infer hidden causal structure. Proc. Annu. Meet. Cogn. Sci. Soc. 26, 500 (2004).
R.S. Michalski: Toward a Unified Theory of Learning: An Outline of Basic Ideas. In First World Conference on the Fundamentals of Artificial Intelligence (Paris), (1991).
J.G. Carbonell, R.S. Michalski, and T.M. Mitchell: An overview of machine learning. In Machine Learning: An Artificial Intelligence Approach, edited by R.S. Michalski, J.G. Carbonell and T.M. Mitchell (Springer-Verlag, Berlin, 1983).
J.B. Tenenbaum, T.L. Griffiths, and C. Kemp: Theory-based Bayesian models of inductive learning and reasoning. Trends Cogn. Sci. 10, 309 (2006).
B.M. Lake, R. Salakhutdinov, and J.B. Tenenbaum: Human-level concept learning through probabilistic program induction. Science 350, 1332 (2015).
W.J. Frawley and G. Piatetsky-Shapior: Knowedge Discovery in Databases. 1st ed. (The MIT Press, Cambridge, 1991).
D. Sacha, M. Sedlmair, L. Zhang, J.A. Lee, J. Peltonen, D. Weiskopf, S.C. North, and D.A. Keim: What you see is what you can change: human-centered machine learning by interactive visualization. Neurocomputing 268, 164 (2017).
A. Jain, G. Hautier, S. Ping Ong, and K. Persson: New opportunities for materials informatics: resources and data mining techniques for uncovering hidden relationships. J. Mater. Res. 31, 977 (2016).
Q. Wu, P. Suetens, and A. Oosterlinck: Integration of heuristic and Bayesian approaches in a pattern-classification system. In Knowledge Discovery Databases, 1st ed, edited by G. Piatetsky-Shapiro, and W.J. Frawley (The MIT Press, Cambridge, 1991), pp. 249–260.
R. Tibshirani: Regression shrinkage and selection via the Lasso. J. R. Stat. Soc. Ser. B 58, 267 (1996).
J.B.O. Mitchell: Machine learning methods in chemoinformatics. Wiley Interdiscip. Rev. Comput. Mol. Sci. 4, 468 (2014).
C.Z. Mooney and R.D. Duval: Bootstrapping A Nonparametric Approach to Statistical Inference (Sage Publications, Inc, Newbury Park, CA, 1993).
V. Svetnik, A. Liaw, C. Tong, J.C. Culberson, R.P. Sheridan, and B.P. Feuston: Random forest: a classification and regression tool for compound classification and QSAR modeling. J. Chem. Inf. Comput. Sci. 43, 1947 (2003).
M. Xu, P. Watanachaturaporn, P.K. Varshney, and M.K. Arora: Decision tree regression for soft classification of remote sensing data. Remote Sens. Environ. 97, 322 (2005).
A. Liaw and M. Wiener: Classification and regression by RandomForest. R News 2/3, 18 (2002).
C.E. Rasmussen: Gaussian processes in machine learning. In Adv. Lect. Mach. Learn. edited by O. Bousquet, U. von Luxburg and G. Rätsch (Springer-Verlag, Berlin, 2003), pp. 63–71.
C.E. Rasmussen and C.K.I. Williams: Gaussian Processes for Machine Learning, 2nd ed. (MIT Press, Cambridge, 2006).
H. Li, C. Collins, M. Tanha, G.J. Gordon, and D.J. Yaron: A density functional tight binding layer for deep learning of chemical hamiltonians. J. Chem. Theory Comput. 14, 5764 (2018).
Y. Li, H. Li, F.C. Pickard, B. Narayanan, F.G. Sen, M.K.Y. Chan, S.K.R.S. Sankaranarayanan, B.R. Brooks, and B. Roux: Machine learning force field parameters from ab initio data. J. Chem. Theory Comput 13, 4492 (2017).
K.T. Schütt, H. Glawe, F. Brockherde, A. Sanna, K.R. Müller, and E.K.U. Gross: How to represent crystal structures for machine learning: towards fast prediction of electronic properties. Phys. Rev. B 89, 205118 (2014).
L. Hu, X. Wang, L. Wong, and G. Chen: Combined first-principles calculation and neural-network correction approach for heat of formation. J. Chem. Phys. 119, 11501 (2003).
O.A. von Lilienfeld: Quantum machine learning in chemical compound space. Angew. Chemie Int. Ed. 57, 4164 (2018).
R.L. Gardas and J.A.P. Coutinho: A group contribution method for viscosity estimation of ionic liquids. Fluid Phase Equilib. 266, 195 (2008).
K. Paduszynski and U. Domanska: Viscosity of ionic liquids: an extensive database and a new group contribution model based on a feed-forward artificial neural network. J. Chem. Inf. Model. 54, 1311 (2014).
A. Mehrkesh and A.T. Karunanithi: New quantum chemistry-based descriptors for better prediction of melting point and viscosity of ionic liquids. Fluid Phase Equilib. 427, 498 (2016).
U. Preiss, S. Bulut, and I. Krossing: In silico prediction of the melting points of ionic liquids from thermodynamic considerations. A case study on 67 salts with a melting point range of 337 °C. J. Phys. Chem. B 114, 11133 (2010).
M.-R. Fatehi, S. Raeissi, and D. Mowla: Estimation of viscosities of pure ionic liquids using an artificial neural network based on only structural characteristics. J. Mol. Liq. 227, 309 (2017).
S.R. Kalidindi and M. De Graef: Materials data science: current status and future outlook. Annu. Rev. Mater. Res. 45, 171 (2015).
C.N. Magnan and P. Baldi: SSpro/ACCpro 5: almost perfect prediction of protein secondary structure and relative solvent accessibility using profiles, machine learning and structural similarity. Bioinformatics 30, 2592 (2014).
G. Pilania, C. Wang, X. Jiang, S. Rajasekaran, and R. Ramprasad: Accelerating materials property predictions using machine learning. Sci. Rep. 3, 2810 (2013).
H.J. Vandenburg, A.A. Clifford, K.D. Bartle, R.E. Carlson, J. Carroll, and I.D. Newton: A simple solvent selection method for accelerated solvent extraction of additives from polymers. Analyst 124, 1707 (1999).
C. Hansen: Hansen Solubility Parameters - A User’s Handbook (CRC Press, Boca Raton, 1999).
T. Lindvig, M.L. Michelsen, and G.M. Kontogeorgis: A Flory–Huggins model based on the Hansen solubility parameters. Fluid Phase Equilib. 203, 247 (2002).
T.A. Albahri: Accurate prediction of the solubility parameter of pure compounds from their molecular structures. Fluid Phase Equilib. 379, 96 (2014).
E. Stefanis and C. Panayiotou: Prediction of Hansen solubility parameters with a new group-contribution method. Int. J. Thermophys. 29, 568 (2008).
Y. Gal and Z. Ghahramani: Proceeding of 33rd International Conference on Machine Learning (New York), (2016).
L. Cao, C. Li, and T. Mueller: The use of cluster expansions to predict the structures and properties of surfaces and nanostructured materials. J. Chem. Inf. Model. 58, 2401 (2018).
T. Mueller and G. Ceder: Bayesian approach to cluster expansions. Phys. Rev. B 80, 024103 (2009).
K.T. Butler, D.W. Davies, H. Cartwright, O. Isayev, and A. Walsh: Machine learning for molecular and materials science. Nature 559, 547 (2018).
J. Ling, R. Jones, and J. Templeton: Machine learning strategies for systems with invariance properties. J. Comput. Phys. 318, 22 (2016).
W. E and P. Ming: Cauchy–Born rule and the stability of crystalline solids: static problems. Arch. Ration. Mech. Anal 183, 241 (2007).
D.C. Ciresan, U. Meier, L.M. Gambardella, and J. Schmidhuber: Deep, big, simple neural nets for handwritten digit recognition. Neural Comput. 22, 3207 (2010).
N. Kambouchev, J. Fernandez, and R. Radovitzky: A polyconvex model for materials with cubic symmetry. Model. Simul. Mater. Sci. Eng. 15, 451 (2007).
A. Karpatne, G. Atluri, J.H. Faghmous, M. Steinbach, A. Banerjee, A. Ganguly, S. Shekhar, N. Samatova, and V. Kumar: Theory-guided data science: a new paradigm for scientific discovery from data. IEEE Trans. Knowl. Data Eng. 29, 2318 (2017).
H. Xiao, J.-L. Wu, J.-X. Wang, R. Sun, and C.J. Roy: Quantifying and reducing model-form uncertainties in Reynolds-averaged Navier–Stokes simulations: a data-driven, physics-informed Bayesian approach. J. Comput. Phys 324, 115 (2016).
J.-X. Wang, J.-L. Wu, and H. Xiao: Physics-informed machine learning approach for reconstructing Reynolds stress modeling discrepancies based on DNS data. Phys. Rev. Fluids 2, 34603 (2017).
L.M. Ghiringhelli, J. Vybiral, S.V. Levchenko, C. Draxl, and M. Scheffler: Big data of materials science: critical role of the descriptor. Phys. Rev. Lett. 114, 105503 (2015).
A. Menon, C. Gupta, K.M. Perkins, B.L. DeCost, N. Budwal, R.T. Rios, K. Zhang, B. Póczos, and N.R. Washburn: Elucidating multi-physics interactions in suspensions for the design of polymeric dispersants: a hierarchical machine learning approach. Mol. Syst. Des. Eng. 2, 263 (2017).
T. Hirata, J. Ye, P. Branicio, J. Zheng, A. Lange, J. Plank, and M. Sullivan: Adsorbed conformations of PCE superplasticizers in cement pore solution unraveled by molecular dynamics simulations. Sci. Rep. 7, 16599 (2017).
D. Marchon, P. Juilland, E. Gallucci, L. Frunz, and R.J. Flatt: Molecular and submolecular scale effects of comb-copolymers on tri-calcium silicate reactivity: toward molecular design. J. Am. Ceram. Soc. 100, 817 (2016).
J.-T. Ding and Z. Li: Effects of Metakaolin and silica fume on properties of concrete. ACI Mater. J. 99, 393 (2002).
N.R. Washburn, A. Menon, C.M. Childs, B. Poczos, and K.E. Kurtis: Machine learning approaches to admixture design for clay-based cements. In Calcined Clays for Sustainable Concrete, edited by F. Martirena, A. Favier and K. Scrivener (Springer, Dordrecht, 2017), pp. 488–493.
A. Menon, C.M. Childs, B. Poczós, N.R. Washburn, and K.E. Kurtis: Molecular engineering of superplasticizers for Metakaolin-Portland cement blends with hierarchical machine learning. Adv. Theory Simul 2, 1800164 (2018).
K. Yoshioka, E. Sakai, M. Daimon, and A. Kitahara: Role of steric hindrance in the performance of superplasticizers for concrete. J. Am. Ceram. Soc. 80, 2667 (1997).
M.L. Hutchinson, E. Antono, B.M. Gibbons, S. Paradiso, J. Ling, and B. Meredig: Overcoming data scarcity with transfer learning. In 31st Conference on Neural Information Processing Systems (NIPS 2017) (Long Beach, 2017), pp. 1–10.
M. Welborn, L. Cheng, and T.F. Miller: Transferability in machine learning for electronic structure via the molecular orbital basis. J. Chem. Theory Comput. 14, 4772 (2018).
A.P. Bartók, S. De, C. Poelking, N. Bernstein, J.R. Kermode, G. Csányi, and M. Ceriotti: Machine learning unifies the modeling of materials and molecules. Sci. Adv. 3, e1701816 (2017).
E.J. Parish and K. Duraisamy: A paradigm for data-driven predictive modeling using field inversion and machine learning. J. Comput. Phys. 305, 758 (2016).
Acknowledgments
Support from the National Science Foundation (CBET-1510600) is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Childs, C.M., Washburn, N.R. Embedding domain knowledge for machine learning of complex material systems. MRS Communications 9, 806–820 (2019). https://doi.org/10.1557/mrc.2019.90
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1557/mrc.2019.90