Abstract
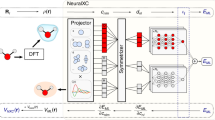
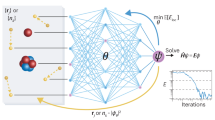
The authors developed a Behler–Parrinello-type neural network (NN) to improve the density-functional tight-binding (DFTB) energy and force prediction. The Δ-machine learning approach was adopted and the NN was designed to predict the energy differences between the density functional theory (DFT) quantum chemical potential and DFTB for a given molecular structure. Most notably, the DFTB-NN method is capable of improving the energetics of intramolecular hydrogen bonds and torsional potentials without modifying the framework of DFTB itself. This improvement enables considerably larger simulations of complex chemical systems that currently could not easily been accomplished using DFT or higher level ab initio quantum chemistry methods alone.




Similar content being viewed by others
References
W.M.C. Foulkes and R. Haydock: Tight-binding models and density-functional theory. Phys. Rev. B 39, 12520–12536 (1989).
D. Porezag, T. Frauenheim, T. Köhler, G. Seifert, and R. Kaschner: Construction of tight-binding-like potentials on the basis of density-functional theory: application to carbon. Phys. Rev. B 51, 12947–12957 (1995).
G. Seifert, D. Porezag, and T. Frauenheim: Calculations of molecules, clusters, and solids with a simplified LCAO-DFT-LDA scheme. Int. J. Quantum Chem. 58, 185–192 (1996).
M. Elstner, D. Porezag, G. Jungnickel, J. Elsner, M. Haugk, T. Frauenheim, S. Suhai, and G. Seifert: Self-consistent-charge density-functional tight-binding method for simulations of complex materials properties. Phys. Rev. B 58, 7260–7268 (1998).
B. Aradi, B. Hourahine, and T. Frauenheim: DFTB+, a sparse matrix-based implementation of the DFTB method. J. Phys. Chem. A 111, 5678–5684 (2007).
M. Elstner and G. Seifert: Density functional tight binding. Philos. Trans. R. Soc. A 372, 20120483–20120494 (2014).
A.S. Christensen, T. Kubar, Q. Cui and M. Elstner: Semiempirical quantum mechanical methods for noncovalent interactions for chemical and biochemical applications. Chem. Rev. 116, 5301–5337 (2016).
K.H. Lee, U. Schnupf, B.G. Sumpter, and S. Irle: Performance of density-functional tight-binding in comparison to ab initio and first-principles methods for isomer geometries and energies of glucose epimers in vacuo and solution. ACS Omega 3, 16899–16915 (2018).
M. Gaus, A. Goez, and M. Elstner: Parametrization and benchmark of DFTB3 for organic molecules. J. Chem. Theory Comput. 9, 338–354 (2013).
V.Q. Vuong, J. Akkarapattiakal Kuriappan, M. Kubillus, J.J. Kranz, T. Mast, T.A. Niehaus, S. Irle, and M. Elstner: Parametrization and benchmark of long-range corrected DFTB2 for organic molecules. J. Chem. Theory Comput. 14, 115–125 (2018).
J. Behler and M. Parrinello: Generalized neural-network representation of high-dimensional potential-energy surfaces. Phys. Rev. Lett. 98, 146401 (2007).
J. Behler: Atom-centered symmetry functions for constructing high-dimensional neural network potentials. J. Chem. Phys. 134, 074106 (2011).
J. Behler: Representing potential energy surfaces by high-dimensional neural network potentials. J. Phys.: Condens. Matter 26, 183001 (2014).
J. Behler: Constructing high-dimensional neural network potentials: a tutorial review. Int. J. Quantum Chem. 115, 1032–1050 (2015).
J.S. Smith, O. Isayev, and A.E. Roitberg: ANI-1: an extensible neural network potential with DFT accuracy at force field computational cost. Chem. Sci. 8, 3192–3203 (2017).
K.T. Schütt, F. Arbabzadah, S. Chmiela, K.R. Müller, and A. Tkatchenko: Quantum-chemical insights from deep tensor neural networks. Nat. Commun. 8, 13890 (2017).
R. Ramakrishnan, P.O. Dral, M. Rupp, and O.A. von Lilienfeld: Big data meets quantum chemistry approximations: the Δ-machine learning approach. J. Chem. Theory Comput. 11, 2087–2096 (2015).
T.T. Nguyen, E. Székely, G. Imbalzano, J. Behler, G. Csányi, M. Ceriotti, A.W. Götz, and F. Paesani: Comparison of permutationally invariant polynomials, neural networks, and Gaussian approximation potentials in representing water interactions through many-body expansions. J. Chem. Phys. 148, 241725 (2018).
L. Shen, J. Wu, and W. Yang: Multiscale quantum mechanics/molecular mechanics simulations with neural networks. J. Chem. Theory Comput. 12, 4934–4946 (2016).
L. Shen and W. Yang: Molecular dynamics simulations with quantum mechanics/molecular mechanics and adaptive neural networks. J. Chem. Theory Comput. 14, 1442–1455 (2018).
M. Gaus, Q. Cui, and M. Elstner: DFTB3: extension of the self-consistent-charge density-functional tight-binding method (SCC-DFTB). J. Chem. Theory Comput. 7, 931–948 (2011).
Keras: Deep Learning for humans. https://github.com/keras-team/keras.
M. Abadi, P. Barham, J. Chen, Z. Chen, A. Davis, J. Dean, M. Devin, S. Ghemawat, G. Irving, M. Isard, M. Kudlur, J. Levenberg, R. Monga, S. Moore, D.G. Murray, B. Steiner, P. Tucker, V. Vasudevan, P. Warden, M. Wicke, Y. Yu, and X. Zheng: TensorFlow: a system for large-scale machine learning. In 12th USENIX Symposium on Operating Systems Design and Implementation (OSDI 16) (2016), pp. 265–283.
D. Nguyen and B. Widrow: Improving the learning speed of 2-layer neural networks by choosing initial values of the adaptive weights. In 1990 IJCNN International Joint Conference on Neural Networks, (1990) (1990), pp. 21–26.
A.D. Becke: Density-functional thermochemistry. III. The role of exact exchange. J. Chem. Phys. 98, 5648–5652 (1993).
A. Hjorth Larsen, J. Jørgen Mortensen, J. Blomqvist, I.E. Castelli, R. Christensen, M. Dulak, J. Friis, M.N. Groves, B. Hammer, C. Hargus, E.D. Hermes, P.C. Jennings, P. Bjerre Jensen, J. Kermode, J.R. Kitchin, E. Leonhard Kolsbjerg, J. Kubal, K. Kaasbjerg, S. Lysgaard, J. Bergmann Maronsson, T. Maxson, T. Olsen, L. Pastewka, A. Peterson, C. Rostgaard, J. Schiøtz, O. Schütt, M. Strange, K.S. Thygesen, T. Vegge, L. Vilhelmsen, M. Walter, Z. Zeng, and K.W. Jacobsen: The atomic simulation environment-a Python library for working with atoms. J. Phys.: Condens. Matter 29, 273002 (2017).
M. Valiev, E. Bylaska, N. Govind, K. Kowalski, T. Straatsma, H. Van Dam, D. Wang, J. Nieplocha, E. Apra, T. Windus, and W. de Jong: NWChem: a comprehensive and scalable open-source solution for large scale molecular simulations. Comput. Phys. Commun. 181, 1477–1489 (2010).
M.A. Addicoat, S. Fukuoka, A.J. Page, and S. Irle: Stochastic structure determination for conformationally flexible heterogenous molecular clusters: application to ionic liquids. J. Comput. Chem. 34, 2591–2600 (2013).
J. Rezác, K.E. Riley, and P. Hobza: S66: a well-balanced database of benchmark interaction energies relevant to biomolecular structures. J. Chem. Theory Comput. 7, 2427–2438 (2011).
J. Rezác: Empirical self-consistent correction for the description of hydrogen bonds in DFTB3. J. Chem. Theory Comput. 13, 4804–4817 (2017).
Acknowledgments
The authors thank Dr. Andreas Goetz (SDSC) and Dr. Thuong Nguyen (UCSD) for providing us their initial BPNN code for water clusters. We also thank Dr Jacek Jakowski, Dr. Panchapakesan Ganesh, and Dr. Maxim Ziatdinov (all ORNL) for helpful discussions. J.Z. acknowledges support under the Oak Ridge Science Semester (ORSS) grant from the Oak Ridge Institute for Science and Education (ORISE). V.Q.V. acknowledges support by an Energy Science and Engineering Fellowship of the Bredesen Center for Interdisciplinary Research and Graduate Education at the University of Tennessee, Knoxville. S.I. acknowledges support by the Fluid Interface Reactions, Structures and Transport (FIRST) Center, an Energy Frontier Research Center funded by the U.S. DOE Office of Science. Calculations were performed at the Center for Nanophase Materials Sciences, which is a U.S. Department of Energy Office of Science User Facility.
Author information
Authors and Affiliations
Corresponding author
Supporting information
43579_2019_9030867_MOESM1_ESM.pdf
Supporting information for: Artificial Neural Network Correction for Density-Functional Tight-Binding Molecular Dynamics Simulations
Supplementary material
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1557/mrc.2019.80.
Rights and permissions
About this article
Cite this article
Zhu, J., Vuong, V.Q., Sumpter, B.G. et al. Artificial neural network correction for density-functional tight-binding molecular dynamics simulations. MRS Communications 9, 867–873 (2019). https://doi.org/10.1557/mrc.2019.80
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1557/mrc.2019.80