Abstract
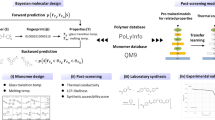
In this perspective, the authors challenge the status quo of polymer innovation. The authors first explore how research in polymer design is conducted today, which is both time consuming and unable to capture the multi-scale complexities of polymers. The authors discuss strategies that could be employed in bringing together machine learning, data curation, high-throughput experimentation, and simulations, to build a system that can accurately predict polymer properties from their descriptors and enable inverse design that is capable of designing polymers based on desired properties.




Similar content being viewed by others
References
A. Gregory and M.H. Stenzel: Complex polymer architectures via RAFT polymerization: From fundamental process to extending the scope using click chemistry and nature’s building blocks. Prog. Polym. Sci. 37, 38 (2012).
S.J. Garcia: Effect of polymer architecture on the intrinsic self-healing character of polymers. Eur. Polym. J. 53, 118 (2014).
A.C. Rinkenauer, S. Schubert, A. Traeger and U.S. Schubert: The influence of polymer architecture on in vitro pDNA transfection. J. Mater. Chem. B 3, 7477 (2015).
A. Dag, M. Callari, H. Lu and M.H. Stenzel: Modulating the cellular uptake of platinum drugs with glycopolymers. Polymer Chemistry 7, 1031 (2016).
D. Paramelle, S. Gorelik, Y. Liu and J. Kumar: Photothermally responsive gold nanoparticle conjugated polymer-grafted porous hollow silica nanocapsules. Chem. Commun. 52, 9897 (2016).
J. Kumar, A. Bousquet and M.H. Stenzel: Thiol-alkyne Chemistry for the Preparation of Micelles with Glycopolymer Corona: Dendritic Surfaces versus Linear Glycopolymer in Their Ability to Bind to Lectins. Macromol. Rapid Commun. 32, 1620 (2011).
J. Kumar, L. McDowall, G. Chen and M.H. Stenzel: Synthesis of thermoresponsive glycopolymersviacopper catalysed azide-alkyne ‘click’ chemistry for inhibition of ricin: the effect of spacer between polymer backbone and galactose. Polymer Chemistry 2, 1879 (2011).
J.-P. Correa-Baena, K. Hippalgaonkar, J. van Duren, S. Jaffer, V.R. Chandrasekhar, V. Stevanovic, C. Wadia, S. Guha and T. Buonassisi: Accelerating Materials Development via Automation, Machine Learning, and High-Performance Computing. Joule 2, 1410 (2018).
J. Bicerano: Prediction of Polymer Properties, (Taylor & Francis Inc, Bosa Roca, United States, 2002).
M.A.C. Stuart, W.T.S. Huck, J. Genzer, M. Muller, C. Ober, M. Stamm, G. B. Sukhorukov, I. Szleifer, V.V. Tsukruk, M. Urban, F. Winnik, S. Zauscher, I. Luzinov and S. Minko: Emerging applications of stimuliresponsive polymer materials. Nat. Mater. 9, 101 (2010).
R. Jiang, Q. Jin, B. Li, D. Ding and A.-C. Shi: Phase Diagram of Poly(ethylene oxide) and Poly(propylene oxide) Triblock Copolymers in Aqueous Solutions. Macromolecules 39, 5891 (2006).
H.S. Ashbaugh and M.E. Paulaitis: Monomer Hydrophobicity as a Mechanism for the LCST Behavior of Poly(ethylene oxide) in Water. Ind. Eng. Chem. Res. 45, 5531 (2006).
A. Halperin, M. Kröger and F.M. Winnik: Poly(N-isopropylacrylamide) Phase Diagrams: Fifty Years of Research. Angew. Chem. Int. Ed. 54, 15342 (2015).
R. Hoogenboom, H.M.L. Thijs, M.J.H.C. Jochems, B.M. van Lankvelt, M. W.M. Fijten and U.S. Schubert: Tuning the LCST of poly(2-oxazoline)s by varying composition and molecular weight: alternatives to poly(N-isopropylacrylamide)? Chem. Commun. 0, 5758 (2008).
G. Odian: Principles of Polymerization, Fourth Edition ed. (John Wiley & Sons, New York, United States, 2004).
J.S. Smith, O. Isayev and A.E. Roitberg: ANI-1: an extensible neural network potential with DFT accuracy at force field computational cost. Chem. Sci 8, 3192 (2017).
T.W. Anderson: An Introduction To Multivariate Statistical Analysis, (Wiley, New York, 1958).
G.E.P. Box and G.C. Tiao: Bayesian Inference in Statistical Analysis, (John Wiley & Sons, New York, United States, 2011).
C. Cortes and V. Vapnik: Support-Vector Networks. Machin. Learn. 20, 273 (1995).
L. Rokach and O. Maimon: Data Mining With Decision Trees: Theory and Applications, (World Scientific Publishing Co., Inc.2014).
Y. LeCun, Y. Bengio and G. Hinton: Deep learning. Nature 521, 436 (2015).
J.H. Friedman: Greedy Function Approximation: A Gradient Boosting Machine. Ann. Stat. 29, 1189 (2001).
V. Aseyev, H. Tenhu and F.M. Winnik: Non-ionic Thermoresponsive Polymers in Water, in Self Organized Nanostructures of Amphiphilic Block Copolymers II, edited by A. H. E. Müller and O. Borisov (Springer Berlin Heidelberg, Berlin, Heidelberg, 2011), pp. 29.
J.N. Wei, D. Duvenaud and A. Aspuru-Guzik: Neural Networks for the Prediction of Organic Chemistry Reactions. ACS Cent. Sci 2, 725 (2016).
R. Gómez-Bombarelli, J. Aguilera-Iparraguirre, T.D. Hirzel, D. Duvenaud, D. Maclaurin, M.A. Blood-Forsythe, H.S. Chae, M. Einzinger, D.-G. Ha, T. Wu, G. Markopoulos, S. Jeon, H. Kang, H. Miyazaki, M. Numata, S. Kim, W. Huang, S.I. Hong, M. Baldo, R.P. Adams and A. Aspuru-Guzik: Design of efficient molecular organic light-emitting diodes by a high-throughput virtual screening and experimental approach. Nat. Mater. 15, 1120 (2016).
F. Häse, C. Kreisbeck and A. Aspuru-Guzik: Machine learning for quantum dynamics: deep learning of excitation energy transfer properties. Chem. Sci 8, 8419 (2017).
S.-L. Benjamin, O. Carlos, G. Gabriel L. and A.-G. Alan: Optimizing distributions over molecular space. An Objective-Reinforced Generative Adversarial Network for Inverse-design Chemistry (ORGANIC), (ChemRxiv, 2017), p. 10.26434/chemrxiv.5309668.v3.
D. Duvenaud, D. Maclaurin, J. Aguilera-Iparraguirre, R. Gomez-Bombarelli, T. Hirzel, A. Aspuru-Guzik and R.P. Adams: Convolutional networks on graphs for learning molecular fingerprints, in Proceedings of the 28th International Conference on Neural Information Processing Systems - Volume 2 (MIT Press, Montreal, Canada, 2015), pp. 2224.
R. Gómez-Bombarelli, J.N. Wei, D. Duvenaud, J.M. Hernández-Lobato, B. Sánchez-Lengeling, D. Sheberla, J. Aguilera-Iparraguirre, T.D. Hirzel, R.P. Adams and A. Aspuru-Guzik: Automatic Chemical Design Using a Data-Driven Continuous Representation of Molecules. ACS Cent. Sci 4, 268 (2018).
T.D. Huan, A. Mannodi-Kanakkithodi, C. Kim, V. Sharma, G. Pilania and R. Ramprasad: A polymer dataset for accelerated property prediction and design. Sci. Data 3, 160012 (2016).
A. Mannodi-Kanakkithodi, G. Pilania, T.D. Huan, T. Lookman and R. Ramprasad: Machine Learning Strategy for Accelerated Design of Polymer Dielectrics. Sci. Rep. 6, 20952 (2016).
C. Kim, A. Chandrasekaran, T.D. Huan, D. Das and R. Ramprasad: Polymer Genome: A Data-Powered Polymer Informatics Platform for Property Predictions. J. Phys. Chem. C 122, 17575 (2018).
M. Zeng, J.N. Kumar, Z. Zeng, S. Ramasamy, V.R. Chandrasekhar and K. Hippalgaonkar: Graph Convolutional Neural Networks for Polymers Property Prediction. arXiv, 1811.06231 (2018).
Q. Wei, R.G. Melko and J.Z.Y. Chen: Identifying polymer states by machine learning. Physical Review E 95, 032504 (2017).
J. Kumar, Q. Li, K.Y.T. Tang, T. Buonassisi, A.L. Gonzalez-Oyarce and J. Ye: Machine Learning Enables Polymer Cloud-Point Engineering via Inverse Design, (ChemRxiv, 2018), p. 10.26434/chemrxiv.7528343.v1.
M.G. Luca, V. Jan, A. Emre, O. Runhai, V.L. Sergey, D. Claudia and S. Matthias: Learning physical descriptors for materials science by compressed sensing. New Journal of Physics 19, 023017 (2017).
B. Dünweg and K. Kremer: Molecular dynamics simulation of a polymer chain in solution. The Journal of Chemical Physics 99, 6983 (1993).
R.D. Groot and P.B. Warren: Dissipative particle dynamics: Bridging the gap between atomistic and mesoscopic simulation. The Journal of Chemical Physics 107, 4423 (1997).
C. Kutzner, S. Páll, M. Fechner, A. Esztermann, B.L. de Groot and H. Grubmüller: Best bang for your buck: GPU nodes for GROMACS biomolecular simulations. Journal of Computational Chemistry 36, 1990 (2015).
S. Oliver, L. Zhao, A.J. Gormley, R. Chapman and C. Boyer: Living in the Fast Lane—High Throughput Controlled/Living Radical Polymerization. Macromolecules 52, 3 (2018).
P. Nikolaev, D. Hooper, F. Webber, R. Rao, K. Decker, M. Krein, J. Poleski, R. Barto and B. Maruyama: Autonomy in materials research: a case study in carbon nanotube growth. Npj Computational Materials 2, 16031 (2016).
Acknowledgments
J.N.K. and Q.L. are supported by the AME Programmatic Fund from the Agency for Science, Technology and Research under Grant No. A1898b0043. The concepts put forward in this paper were developed through discussions with Prof. Tonio Buonassisi, Dr. Kedar Hippalgaonkar, and Dr. Anibal L. Gonzalez-Oyarce.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kumar, J.N., Li, Q. & Jun, Y. Challenges and opportunities of polymer design with machine learning and high throughput experimentation. MRS Communications 9, 537–544 (2019). https://doi.org/10.1557/mrc.2019.54
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1557/mrc.2019.54