Abstract
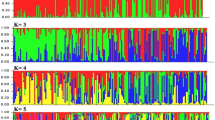
The microsatellites, as one of the most robust markers for identification of wheat varieties, were used for assessment of genetic diversity and population structure to promote effective use of genetic resources. In this study, the set of 284 wheat varieties were genotyped using 30 microsatellite markers. The chosen SSR markers were located among almost all linkage groups and covered all three genomes. The genotypes used originate from 24 different breeding centers worldwide and are included in an extensive core collection of the Institute of Field and Vegetable Crops in Novi Sad, Serbia. The total number of detected alleles was 349 at all analyzed loci. The average number of detected allelic variant per locus was 11.5. The mean value of polymorphic information content was 0.68. According to the probability of data obtained by program Structure, the results have shown presence of 6 subpopulations within the studied set of genotypes. The population structure positively correlated to some extent with geographic origin. The available pedigree data were included for additional explanation of population structure. The results of this study should provide valuable information for future association studies using the diverse wheat breeding material.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Borojevic, K., Borojevic, K. 2005. Historic role of the wheat variety Akakomughi in Southern and Central European wheat breeding programs. Breeding Sci. 55:253–256.
Chao, S.M., Zhang, W.J., Dubcovsky, J., Sorrels, M. 2007. Evaluation of genetic diversity and genome-wide likage disequilibrium among U.S. wheat (Triticum aestivum L.) germplasm representing different market classes. Crop Sci. 47:1018–1030.
Chen, X., Min, D., Yasir, T.A., Hu, Y.G. 2012. Genetic diversity, population structure and linkage disequilibrium in elite Chinese winter wheat investigated with SSR markers. PLoS one 7 (9):e44510. doi: https://doi.org/10.1371/journal.pone0044510
Chen, X.M., He, Z.H., Shi, J.R., Xia, L.Q., Ward, R., Zhou, Y., Jiang, G.L. 2003. Genetic diversity of high quality winter wheat varieties (lines) based on SSR marker. Acta Agronomica Sinica 29:13–19.
Couviour, F.L., Faure, S., Poupard, B., Flodrops, Y., Dubreuil, P., Praud, S. 2011. Analysis of genetic structure in a panel of elite wheat varieties and relevance for association mapping. Theor. Appl. Genet. 123:715–727.
Doyle, J.J., Doyle, J.L. 1990. Isolation of plant DNA from fresh tissue. Focus 12:13–15.
Flint-Garcia, S.A., Thornsberry, J.M., Buckler, E.S. 4th 2003. Structure of linkage disequilibrium in plants. Annu. Rev. Plant Biol. 54:357–374.
Huang, X.Q., Börner, A., Röder, M.S., Ganal, M.W. 2002. Assessing genetic diversity of wheat (Triticum aestivum L.) germplasm using microsatellite markers. Theor. Appl. Genet. 105:699–707.
Kobiljski, B., Dencic, S., Kondic-Spika, A., Lohwasser, U., Börner, A. 2009. Locating stable across environment QTL involved in the determination of agronomic characters in wheat. Cereal Res. Commun. 37:327–333.
Liu, K., Muse, S.V. 2005. PowerMarker: An integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129.
Neumann, K., Kobiljski, B., Dencic, S., Varshney, R.K., Börner, A. 2011. Genome-wide association mapping: A case study in bread wheat (Triticum aestivum L.). Mol. Breed. 27:37–58.
Pritchard, J.K., Stephens, M., Donnelly, P. 2000. Inference of population structure using multilocus genotype data. Genetics 155:945–959.
Randhawa, H.S., Muhammad, A., Pozniak, C., Clarke, J.M., Graf, R.J., Fox, S.L., Humphreys, D.G., Knox, R.E., Depauw, R.M., Singh, A.K., Cuthbert, R.D., Hucl, P., Spaner, D. 2013. Application of molecular markers to wheat breeding in Canada. Plant Breed. 132:458–471.
Röder, M.S., Korzun, V., Wendehake, K., Plaschke, J., Tixier, M-H., Leroz, P., Ganal, M.W. 1998. A microsatellite map of wheat. Genetics 194:2007–2023.
Röder, M.S., Wendehake, K., Korzun, V., Bredemeijer, G., Laborie, D., Bertrand, L., Isaac, P., Rendell, S., Jackson, J., Cooke, R.J., Vosman, B., Ganal, M. 2002. Construction and analysis of a microsatellite-based database of European wheat varieties. Theor. Appl. Genet. 106:67–73.
Stachel, M., Lelley, T., Grausgruber, H., Vollmann, J. 2000. Application of microsatellites in wheat (Triticum aestivum L.) for studying genetic differention caused by selection for adaptation and use. Theor. Appl. Genet. 100:242–248.
Varshney, R.K., Thudi, M., Aggarwal, R., Börner, A. 2007. Genic molecular markers in plants: Development and applications. In: Varshney, R.K., Tuberosa, R. (eds), Genomics-assisted Crop Improvement: Genomics Approaches and Platforms. Vol. 1. Springer, Dordrecht, The Netherlands, pp. 13–29.
Wang, Y., Wang, C., Zhang, H., Yue, Z., Liu, X., Ji, W. 2013. Genetic analysis of wheat (Triticum aestivum L.) and related species with SSR markers. Genetic Res. and Crop Evol. 60:1105–1117.
Yu, J., Buckler, E.S. 2006. Genetic association mapping and genome organization of maize. Current Opinion in Biotechnol. 17:155–160.
Zhang, D., Bai, G., Zhu, C., Yu, J., Carver, B.F. 2010. Genetic diversity, population structure, and linkage disequilibrium in U.S. elite winter wheat. The Plant Genome 3:117–127.
Zhang, L., Liu, D.C., Guo, X.L., Yang, W.L., Sun, J.Z., Wang, D.W., Sourdille, P., Zhang, A.M. 2011. Investigation of genetic diversity and population structure of common wheat cultivars in northern China using DArT markers. BMC Genetics 12:42.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Börner
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Brbaklić, L., Trkulja, D., Kondić-Špika, A. et al. Determination of Population Structure of Wheat Core Collection for Association Mapping. CEREAL RESEARCH COMMUNICATIONS 43, 22–28 (2015). https://doi.org/10.1556/CRC.2014.0027
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1556/CRC.2014.0027