Abstract
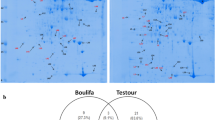
Hordeum spontaneum (wild barley) is a good gene source to improve salt tolerance in barley because it rapidly hybridizes and recombines with barley cultivars. Proteomics can assist in identifying proteins associated with a certain environmental or developmental signal. We employed a proteomic approach to understand the mechanisms of plant responses to salinity in a salt tolerant accession of H. spontaneum. At the 4-leaf stage, wild barley plants were exposed to 0 (control treatment) or 300 mM NaCl (salt treatment). The salt treatment lasted 3 weeks. Total proteins of leaf 4 were extracted and separated by two-dimensional gel electrophoresis. More than 500 protein spots were reproducibly detected. Of these, 29 spots showed significant differences between salt treatment and control. Using MALDI-TOF-TOF MS, we identified 29 cellular proteins, which represented 16 different proteins. These were classified into six categories and a group with unknown biological function. The proteins identified were involved in many different cellular functions. Three spots were identified as unknown proteins; searching in the NCBI database revealed that there was a 71% match with clathrin assembly protein putative [Ricinus communis], a 67% match with actin binding protein [Zea mays], and a 66% match with phosphatidylinositol kinase [Arabidopsis thaliana]. Other proteins identified included ribulose-1,5-bisphosphate carboxylase/oxygenase (Rubisco), oxygen-evolving enhancer protein (OEE), photosystem II reaction centerWprotein (Psbw), ribosomal proteins, chloroplast RNA binding protein (ChRBP), superoxide dismutase (SOD), malate dehydrogenase (MDH), thioredoxin h (Trx), nucleoside diphosphate kinase (NDPK), profilin, translationally-controlled tumor protein (TCTP), polyamine oxidase (PAO) and universal stress protein family (USP).
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Askari, H., Edqvist, J., Hajheidari, M., Kafi, M., Salekdeh, G.H. 2006. Effects of salinity levels on proteome of Suaeda aegyptiaca leaves. Proteomics 6:2542–2554.
Bradford, M.M. 1976. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-Dye binding. Anal. Biochem. 72:248–254.
Cooley, T., Walters, D.R., 2002. Polyamine metabolism in barley reacting hypersensitively to the powdery mildew fungus Blumeria graminis f. sp. Hordei. Plant Cell Environ. 25:461–468.
Dadashi Dooki, A., Mayer-Posne, F., Askari, H., Zaiee, A., Salekdeh, G.H. 2006. Proteomic responses of rice young panicles to salinity. Proteomics 6:6498–6507.
Damerval, C., de Vienne, D., Zivy, M., Thiellement, H. 1986. Technical improvements in two-dimensional electrophoresis increase the level of genetic variation detected in wheat-seedling proteins. Electrophoresis 7:52–54.
Escobar Galvis, M.L., Marttila, S., Hakansson, G., Forsberg, J., Knorpp, C. 2001. Heat stress response in pea involves interaction of mitochondrial nucleoside diphosphate kinase with a novel 86-kilodalton protein. Plant Physiol. 126:69–77.
Görg, A., Postel, W., Günther, S. 1988. The current state of two-dimensional electrophoresis with immobilized pH gradients. Electrophoresis 9:531–546.
Groppa, M.D., Benavides, M.P. 2008. Polyamines and abiotic stress: recent advances. Amino Acids 34:35–45.
Hajduch, M., Rakwal, R., Agrawal, G.K., Yonekura, M., Pretova, A. 2001. High-resolution two-dimensional electrophoresis separation of proteins from metal-stressed rice (Oryza sativa L.) leaves: Drastic reductions/fragmentation of ribulose-1,5-bisphosphate carboxylase/oxygenase and induction of stress-related proteins. Electrophoresis 22:2824–2831.
Helena, S., Ake, S. 2004. A Pisum sativum glyoxysomal malate dehydrogenase induced by cadmium exposure. J. of DNA Sequencing and Mapping 15:206–208.
Janicka-Russak, M., Kabala, K., Mlodzinska, E., Klobus, G. 2010. The role of polyamines in the regulation of the plasma membrane and the tonoplast proton pumps under salt stress. J. Plant Physiol. 167:261–269.
Laloi, C., Mestres-Ortega, D., Marco, Y., Meyer, Y., Reichheld, J.P. 2004. The Arabidopsis cytosolic thioredoxin h5 gene induction by oxidative stress and its W-box-mediated response to pathogen elicitor. Plant Physiol. 134:1006–1016.
Leshem, Y., Seri, L., Levine, A. 2007. Induction of phosphatidylinositol 3-kinase-mediated endocytosis by salt stress leads to intracellular production of reactive oxygen species and salt tolerance. The Plant Journal 51:185–197.
Ma, Y., Cheng, Z., Wang, W., Sun, Y. 2007. Proteomic analysis of high yield rice variety mutated from spaceflight. Adv. Space Res. 40:535–539.
Mikami, K., Katagiri, T., Iuchi, S., Yamaguchi-Shinozaki, K., Shinozaki, K. 1998. A gene encoding phosphatidylinositol-4-phosphate 5-kinase is induced by water stress and abscisic acid in Arabidopsis thaliana. The Plant Journal 15:563–568.
Moon, H., Lee, B., Choi, G., Shin, D., Prasad, D.T., Lee, O., Kwak, S.S., Kim, D.H., Nam, J., Bahk, J., Hong, J.C., Lee, S.Y., Cho, M.J., Lim, C.O., Yun, D.J. 2003. NDP kinase 2 interacts with two oxidative stress-activated MAPKs to regulate cellular redox state and enhances multiple stress tolerance in transgenic plants. Proc. Natl. Acad. Sci. USA. 100:358–363.
Munns, R., Tester, M. 2008. Mechanisms of salinity tolerance. Ann. Rev. Plant Biol. 59:651–681.
Nevo, E., Krugman, T., Beiles, A. 1993. Genetic resources for salt tolerance in the wild progenitors of wheat (Triticum dicoccoides) and barley (Hordeum spontaneum) in Israel. Plant Breed. 110:338–341.
O’Toole, R., Williams, H.D. 2003. Universal stress proteins and Mycobacterium tuberculosis. Res. Microbiol. 154:387–392.
Salekdeh, G.H., Siopongco, J., Wade, L.J., Ghareyazie, B., Bennett, J. 2002. Proteomic analysis of rice leaves during drought stress and recovery. Proteomics 2:1131–1145.
Shavrukov, Y., Gupta, N., Miyazaki, J., Baho, M., Chalmers, K., Tester, M., Langridge, P., Collins, N. 2010. HvNax3 — a locus controlling shoot sodium exclusion derived from wild barley (Hordeum vulgare ssp. spontaneum). Funct. Integr. Genomics 10:277–291.
Sobhanian, H., Razavizadeh, R., Nanjo, Y., Ehsanpour, A., Rastgar Jazii, F., Motamed, N., Komatsu, S. 2010. Proteome analysis of soybean leaves, hypocotyls and roots under salt stress. Proteome Sci. 8:19–33.
Sottosanto, J.B., Gelli, A., Blumwald, E. 2004. DNA array analyses of Arabidopsis thaliana lacking a vacuolar Na+/H+ antiporter: Impact of AtNHX1 on gene expression. The Plant Journal 40:752–771.
Sreenivasulua, N., Grimma, B., Wobusa, U., Weschkea, W. 2000. Differential response of antioxidant compounds to salinity stress in salt-tolerant and salt-sensitive seedlings of foxtail millet (Setaria italica). Physiological Plantarum 109:435–442.
Sugihara, K., Hanagata, N., Dubinsky, Z., Baba, S., Karube, I. 2000. Molecular characterization of cDNA encoding oxygen evolving enhancer protein 1 increased by salt treatment in the mangrove Bruguiera gymnorrhiza. Plant Cell Physiol. 41:1279–1285.
Unanue, E.R., Ungewickell, E., Branton, D. 1981. The binding of clathrin triskelions to membranes from coated vesicles. Cell 26:439–446.
Uribe, R., Jay, D. 2009. A review of actin binding proteins: New perspectives. Mol. Biol. Rep. 36:21–125.
Visa, N. 2005. Actin in transcription. EMBO Rep. 6:218–219.
Witzel, K., Weidneri, A., Surabhi, G., Varsheney, R., Kunze, G.H., Bck-sorlin, G., Borneri, A., Mock, H. 2010. Comparative analysis of the grain proteome fraction in barley genotypes with contrasting salinity tolerance during germination. Plant Cell Environ. 33:211–222.
Zorb, C., Schmitt, S., Neeb, A., Karl, S., Linder, M., Schubert, S. 2004. The biochemical reaction of maize (Zea mays L.) to salt stress is characterized by a mitigation of symptoms and not by a specific adaptation. Plant Sci. 167:91–100.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Pécsváradi
Electronic Supplementary Material (ESM)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Fatehi, F., Hosseinzadeh, A., Alizadeh, H. et al. The Proteome Response of Hordeum spontaneum to Salinity Stress. CEREAL RESEARCH COMMUNICATIONS 41, 78–87 (2013). https://doi.org/10.1556/CRC.2012.0017
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1556/CRC.2012.0017