Abstract
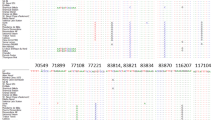
By employing in silico tools, we devised new chloroplast microsatellite primers for inferring phyloge-netic relationships within Brassicaceae. Microsatellite repeats were scanned in 12 chloroplast genomes of Brassicaceae, regions flanking these repeats were aligned and 19 universal primers were designed. Fifteen of these primer pairs are predicted to yield polymorphic amplicons, that are more or less evenly distributed throughout the chloroplast genomes. Finally, using PCR, we have validated three primer pairs on a limited ‘test set’ of plants, different from those used in computational analysis.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Angioi, S. A., Desiderio, F., Rail, D., Bitocchi, E., Attene, G., Papa, R. (2009) Development and use of chloroplast microsatellites in Phaseolus spp. and other legumes. Plant Biol (Stuttg) 11, 598–612.
Beilstein, M. A., Al-Shehbaz, I. A., Kellogg, E. A. (2006) Brassicaceae: Phylogeny and trichome evolution. Am. J. Botany 93, 607–619.
Benson, D. A., Karsch-Mizrachi, I., Lipman, D. J., Ostell, J., Wheeler, D. L. (2006) GenBank. Nucleic Acids Res. 34, D16-20.
Carver, T., Thomson, N., Bleasby, A., Berriman, M., Parkhill, J. (2009) DNAPlotter: circular and linear interactive genome visualization. Bioinformatics 25, 119–120.
Cheng, Y., de Vicente, M. C., Meng, H., Guo, W., Tao, N., Deng, X. (2005) A set of primers for analyzing chloroplast DNA diversity in Citrus and related genera. Tree Physiol. 25, 661–672.
Clegg, M. T., Gaut, B. S., Learn, G. H., Jr., Morton, B. R. (1994) Rates and patterns of chloroplast DNA evolution. Proc. Natl Acad Sci. USA, 91, 6795–6801.
Couvreur, T. L., Franzke, A., Al-Shehbaz, I. A., Bakker, F. T., Koch, M. A., Mummenhoff K. (2010) Molecular phylogenetics, temporal diversification, and principles of evolution in the mustard family (Brassicaceae). Mol. Biol. Evol. 27, 55–71.
Grant, J. R., Stothard, P. (2008) The CGView Server: a comparative genomics tool for circular genomes. Nucleic Acids Res. 36, Wl 81–184.
He, G., Meng, R., Newman, M., Gao, G., Pittman, R. N., Prakash, C. S. (2003) Microsatellites as DNA markers in cultivated peanut (Arachis hypogaea L.). BMC Plant Biol. 3, 3.
Hong, C. P., Piao, Z. Y., Kang, T. W., Batley, J., Yang, T. J., Hur, Y K., Bhak, J., Park, B. S., Edwards, D., Lim, Y P. (2007) Genomic distribution of simple sequence repeats in Brassica rapa. Mol. Cells 23, 349–356.
Katti, M. V., Sami-Subbu, R., Ranjekar, P. K., Gupta, V. S. (2000) Amino acid repeat patterns in protein sequences: their diversity and structural-functional implications. Protein Sci. 9, 1203–1209.
Koch, M. A., Mummenhoff, K. (2006) Editorial: Evolution and phylogeny of the Brassicaceae. Plant Syst. Evol. 259, 81–83.
Li, Y C., Korol, A. B., Fahima, T., Beiles, A., Nevo, E. (2002) Microsatellites: genomic distribution, putative functions and mutational mechanisms: a review. Mol. Ecol. 11, 2453–2465.
Powell, W., Morgante, M., McDevitt, R., Vendramin, G. G., Rafalski, J. A. (1995) Polymorphic simple sequence repeat regions in chloroplast genomes: applications to the population genetics of Pines. Proc. Natl Acad. Sci. USA, 92, 7759–7763.
Provan, J., Biss, P. M., McMeel, D., Mathews, S. (2004). Universal primers for the amplification of chloroplast microsatellites in grasses (Poaceae). Mol. Ecol. Notes 4, 262–264.
Rajendrakumar, P., Biswal, A. K., Balachandran, S. M., Srinivasarao, K., Sundaram, R. M. (2007) Simple sequence repeats in organellar genomes of rice: frequency and distribution in genie and inter-genic regions. Bioinformatics 23, 1–4.
Rajendrakumar, P., Biswal, A. K., Balachandran, S. M., Sundaram, R. M. (2008) In silico analysis of microsatellites in organellar genomes of major cereals for understanding their phylogenetic relationships. In Silico Biol. 8, 87–104.
Rozen, S., Skaletsky, H. (2000) Primer3 on the WWW for general users and for biologist programmers. Methods Mol. Biol. 132, 365–386.
Scheen, A. C., Elven, R., Brochmann, C. (2002) A molecular-morphological approach solves taxo-nomic controversy in arctic Draba (Brassicaceae). Canadian Journal of Botany 80, 59–71.
Stothard, P., Wishart, D. S. (2005) Circular genome visualization and exploration using CGView. Bioinformatics 21, 537–539.
Vendramin, G. G., Lelli, L., Rossi, P., Morgante, M. (1996) A set of primers for the amplification of 20 chloroplast microsatellites in Pinaceae. Mol. Ecol. 5, 595–598.
Weising, K., Gardner, R. C. (1999) A set of conserved PCR primers for the analysis of simple sequence repeat polymorphisms in chloroplast genomes of dicotyledonous angiosperms. Genome 42. 9–19.
Zardoya, R., Vollmer, D. M., Craddock, C., Streelman, J. T., Karl, S., Meyer, A. (1996) Evolutionary conservation of microsatellite flanking regions and their use in resolving the phylogeny of cichlid fishes (Pisces: Perciformes). Proc. Biol. Sci. 263, 1589–1598.
Acknowledgement
Financial assistance provided by institutional CSIR grants in deeply acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Awasthi, P., Ahmad, I., Gandhi, S.G. et al. Development of Chloroplast Microsatellite Markers for Phylogenetic Analysis in Brassicaceae. BIOLOGIA FUTURA 63, 463–473 (2012). https://doi.org/10.1556/ABiol.63.2012.4.5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1556/ABiol.63.2012.4.5