Abstract
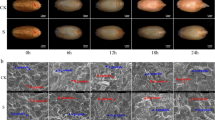
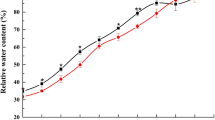
Seed germination is a new beginning for the crop life cycle, which is closely related to seed sprouting and subsequent plant growth and development, and ultimately affects grain yield and quality. Salt stress is one of the most important abiotic stress factors that restrict crop production. Therefore, it is highly important to improve crop salt tolerance and sufficient utilization of saline-alkali land. In this study, we identified the phosphorylated proteins involved in salt stress response by combining SEM, 2-DE, Pro-Q Diamond staining and tandem mass spectrometry. The results showed that salt stress significantly inhibited seed germination and starch degradation. In total, 14 phosphorylated protein spots (11 unique proteins) in the embryo and 6 phosphorylated protein spots (4 unique proteins) in the endosperm were identified, which mainly involved in stress/defense, protein metabolism and energy metabolism. The phosphorylation of some proteins such as cold regulated proteins, 27K protein, EF-1β and superoxide dismutase could play important roles in salt stress tolerance.
Similar content being viewed by others
References
Agrawal, G.K., Thelen, J.J. 2005. Development of as implified, economical polyacrylamide gel staining protocol for phosphoproteins. Proteomics 5:4684–4688.
Ahmad, P., Prasad, M.N.V. 2012. Abiotic stress responses in plants: metabolism, productivity and sustainability. Springer Science & Business Media Alcohol Alcoholism 30:153–161.
Andre, C., Froehlich, J.E., Moll, M.R., Benning, C. 2007. A heteromeric plastidic pyruvate kinase complex involved in seed oil biosynthesis in Arabidopsis. Plant Cell 19:2006–2022.
Beck, E., Ziegler, P. 1989. Biosynthesis and degradation of starch in higher plants. Annu. Rev. Plant Biol. 40:95–117.
Bewley, J.D., Black, M. 1994. Seeds physiology of development and germination. Plenum Press.
Blom, N., Gammeltoft S., Brunak S. 1999. Sequence and structure-based prediction of eukaryotic protein phosphorylation sites. J. Mol. Biol. 294:1351–1362.
Borsani, O., Valpuesta, V., Botella, M.A. 2001. Evidence for a role of salicylic acid in the oxidative damage generated by NaCl and osmotic stress in Arabidopsis seedlings. Plant Physiol. 126:1024–1030.
Carr-Schmid, A., Valente, L., Loik, V.I., Williams, T., Starita, L.M., Kinzy, T.G. 1999. Mutations in elongation factor 1β, a guanine nucleotide exchange factor, enhance translational fidelity. Mol. Cell. Biol. 19:5257–5266.
Cheng, X.X., Yu, M., Zhang, N., Zhou, Z.Q., Xu, Q.T., Mei, F.Z., Qu, L.H. 2016. Reactive oxygen species regulate programmed cell death progress of endosperm in winter wheat (Triticum aestivum L.) under waterlogging. Protoplasma 253:311–327.
Dong, K., Zhen, S., Cheng, Z., Cao, H., Ge, P., Yan Y. 2015. Proteomic analysis reveals key proteins and phosphoproteins upon seed germination of wheat (Triticum aestivum L.). Front. Plant Sci. 6:1017.
Fetzer, S., Lauber, J., Will, C.L., Lührmann, R. 1997. The [U4/U6.U5] tri-snRNP-specific 27K protein is a novel SR protein that can be phosphorylated by the snRNP-associated protein kinase. RNA 3:344.
Gu, A., Hao, P., Lv, D., Zhen, S., Bian, Y., Ma, C., Li, X., Zhang, W., Yan, Y. 2015. Integrated proteome analysis of the wheat embryo and endosperm reveals central metabolic changes involved in water deficit response during the grain development. J. Agric. Food Chem. 63:8478–8487.
Guo, G., Lv, D., Yan, X., Subburaj, S., Ge, P., Li, X., Hu, Y., Yan, Y. 2012. Proteome characterization of developing grains in bread wheat cultivars (Triticum aestivum L.). BMC Plant Biol. 12:147.
Han, C., Zhen, S., Zhu, G., Bian, Y., Yan, Y. 2017. Comparative metabolome analysis of wheat embryo and endosperm reveals the dynamic changes of metabolites during seed germination. Plant Physiol. Biochem. 115:20–327.
Horton, P., Park, K.J., Obayashi, T., Nakai, K. 2006. Protein subcellular localization prediction with WOLF PSORT. Proceedings of the 4th Asia-Pacific Bioinformatics Conference 3:39–48.
Kazuoka, T., Oeda, K. 1992. Heat-stable COR (cold-regulated) proteins associated with freezing tolerance in spinach: Plant & Cell Physiol. 33:1107–1114.
Li, M., Yin, X., Sakata, K., Yang, P., Komatsu, S. 2015. Proteomic analysis of phosphoproteins in the rice nucleus during the early stage of seed germination. J. Proteome Res. 14:2884–2896.
Lv, D.W., Subburaj, S., Cao, M., Yan, X., Li, X., Appels, R., Sun, D.F., Ma, W., Yan, Y.M. 2014. Proteome and phosphoproteome characterization reveals new response and defense mechanisms of Brachypodium distachyon leaves under salt stress. Mol. Cell. Proteomics 13:632–652.
Lv, D.W., Zhu, G.R., Zhu, D., Bian, Y.W., Liang, X.N., Cheng, Z.W., Deng, X., Yan, Y.M. 2016. Proteomic and phosphoproteomic analysis reveals the response and defense mechanism in leaves of diploid wheat T. monococcum under salt stress and recovery. J. Proteomics 128:388–402.
Malcolm, E., Sumner, R.N. 1998. Sodic soils: distribution, properties, management, and environmental consequences. New York: Oxford University Press.
Silva-Sanchez, C., Li, H., Chen, S. 2015. Recent advances and challenges in plant phosphoproteomics. Proteomics 15:1127–1141.
Sirover, M.A. 2011. On the functional diversity of glyceraldehyde-3-phosphate dehydrogenase: biochemical mechanisms and regulatory control. Biochim. Biophys. Acta 1810:741–751.
Small, I., Peeters, N., Legeai, F., Lurin, C. 2004. Predotar: A tool for rapidly screening proteomes for N-terminal targeting sequences. Proteomics 4:1581–1590.
Tetlow, I.J., Wait, R., Lu, Z.X., Akkasaeng, R., Bowsher, C.G., Esposito, S., Kosar-Hashemi, B., Morell, M.K., Emes, M.J. 2004. Protein phosphorylation in amyloplasts regulates starch branching enzyme activity and protein-protein interactions. Plant Cell 16:694–708.
Vu, L.D., Verstraeten, I., Stes, E., Van Bel, M., Coppens, F., Gevaert, K., De Smet, I. 2017. Proteome profiling of wheat shoots from different cultivars. Front. Plant Sci. 8:626.
Wang, P., Kong, C.H., Hu, F., Xu, X.H. 2007. Allantoin involved in species interactions with rice and other organisms in paddy soil. Plant Soil 296:43.
Wu, J.Y., Liu, J.H., Li, Q., Fu, Z.J. 2009. Effect of salt stress on oat seed germination and seeding membrane permeability. J. Triticeae Crops 29:341–345.
Xing, S.J., Zhang, J.F. 2006. Land degradation mechanism and vegetation restoration technology in the Yellow River delta. Beijing: China Forestry Press.
Yang, X., Qiao, J. 2007. The production and trade of wheat in the world. Beijing: Life World. 9:22–25.
Zhang, J., Wang, G., Huang, S., Xuan, J., Jia, X., Guo, Z. 2015. Functions of alcohol dehydrogenase family in abiotic stress responses in plants. Chinese Agric. Sci. Bull. 31:246–250.
Zhang, M., Ma, C., Lv, D., Zhen, S., Li, X., Yan, Y. 2014. Comparative phosphoproteome analysis of the developing grains in bred wheat (Triticum aestivum L.) under well-watered and water-deficit conditions. J. Proteome Res. 13:4281–4297.
Zhu, J.K. 2003. Regulation of ion homeostasis under salt stress. Curr. Opin. Plant Biol. 6:441–445.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by I. Molnár
Rights and permissions
About this article
Cite this article
Luo, X., Han, C., Deng, X. et al. Identification of Phosphorylated Proteins in Response to Salt Stress in Wheat Embryo and Endosperm during Seed Germination. CEREAL RESEARCH COMMUNICATIONS 47, 53–66 (2019). https://doi.org/10.1556/0806.46.2018.061
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1556/0806.46.2018.061