Abstract
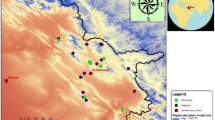
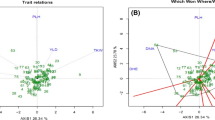
The knowledge about genetic diversity in the wild relatives of wheat provides useful information for breeding programs and gene pool management. In the present study, an assessment of agro-morphological diversity and molecular variability among 70 accessions of Triticum, belonging to T. boeoticum, T. urartu, T. durum and T. aestivum species, collected from different regions of Iran was made. According to phenotypic analysis, all traits except peduncle length, stem diameter and the number of seeds per spike indicated a high level of diversity among studied accessions. Also, principal component analysis identified six components that explained 87.53% of the total variation in agro-morphological traits. In molecular analysis, 15 start codon targeted (SCoT) polymorphism primers produced 166 bands, out of which, 162 (97.59%) were polymorphic. Analysis of molecular variance (AMOVA) indicated the 63% of the variation resided among populations. The maximum value of polymorphism information content (PIC), the observed (Na) and effective (Ne) number of alleles, Nie’s gene diversity (He) and Shannon’s information index (I) was detected for T. boeoticum than the other species. The SCoT-based tree revealed three different groups corresponding to the genomic constitution in Triticum germplasm, which was in part confirmed by STRUCTURE and principal coordinate (PCoA) analyses. Our results indicated a remarkable level of genetic diversity among studied Iranian Triticum species, especially T. boeoticum, which can be of interest for future breeding and other analyses associated with future studies of the wild relatives of wheat. More importantly, our results revealed that SCoT markers could be used to accurate evaluate genetic diversity and phylogenetic relationships among different Triticum species.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Alsaleh, A., Shehzad Baloch, F., Nachit, M., Ozkan, H. 2016. Phenotypic and genotypic intra-diversity among Anatolian durum wheat “Kunduru” landraces. Biochem. Syst. Ecol. 65:9–16.
Bhattacharyya, P., Kumaria, S., Kumar, S., Tandon, P. 2013. Start codon targeted (SCoT) marker reveals genetic diversity of Dendrobium nobile Lindl., an endangered medicinal orchid species. Gene. 529:21–26.
Börner, A., Plaschke, J., Korzun, V., Worland, A. 1996. The relationships between the dwarfing genes of wheat and rye. Euphytica 89:69–75.
Collard, B.C.Y., Mackill, D.J. 2009. Start codon targeted (SCoT) polymorphism: a simple, novel DNA marker technique for generating gene-targeted markers in plants. Plant Mol. Biol. Rep. 27:86–93.
Doyle, J.J., Doyle, J.L. 1987. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical. Bulletin 19:11–15.
Dumolin-Lapegue, S., Demesure, B., Fineschi, S., Le Corre, V., Petit, R.J. 1997. Phylogeographic structure of white oaks throughout the European continent. Genetics 146:1475–1487.
Earl, D.A., von Holdt, B.M. 2012. STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 4:359–361.
Eticha, F., Bekele, E., Belay, G., Börner, A. 2005. Phenotypic diversity in durum wheat collected from Bale and Wello regions of Ethiopia. Plant. Genet. Resour. 3:35–43.
Etminan, A., Pour-Aboughadareh, A., Mohammadi, R., Ahmadi-Rad, A., Noori, A., Mahdavian, Z., Moradi, Z. 2016. Applicability of start codon targeted (SCoT) and inter-simple sequence repeat (ISSR) markers for genetic diversity analysis in durum wheat genotypes. Biotechnol. Biotec. Eq. 30:1075–1081.
Evanno, G., Regnaut, S., Goudet, J. 2005. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecol. 14:2611–2620.
Gorji, A.M., Poczai, P., Polgar, Z., Taller, J. 2011. Efficiency of arbitrarily amplifed dominant markers (SCoT, ISSR and RAPD) for diagnostic fingerprinting in tetraploid potato. Am. J. Potato. Res. 88:226–237.
Hamidi, H., Talebi, R., Keshavarz, F. 2014. Comparative efficiency of functional gene-based markers, start codon targeted polymorphism (SCoT) and conserved DNA-derived Polymorphism (CDDP) with ISSR markers for diagnostic fingerprinting in wheat (Triticum aestivum L.). Cereal. Res. Commun. 42:558–567.
Hamrick, J.L., Godt, M.J.W., Murawski, D.A., Loveless, M.D. 1991. Correlation between species traits and allozyme diversity: implications for conservation biology. In: Falk, D.A., Holsinger, K.E. (eds), Genetic and Conservation of Rare Plants. Oxford University Press. New York, USA. pp. 75–86.
Hurd, E.A. 1971. Techniques for measuring plant drought stress. In: Larson, K.L., Rachter, J.D. (eds), Drought Injury and Resistance in Crops. CSSA Special Publication II, Crop Science Society of America. Cambridge University Press. Madison, WI, USA. pp. 77–88.
Hutchenson, K. 1970. A test for comparing diversities based on the Shannon formula. J. Theor. Biol. 29:151– 154.
IBPGR 1985. Revised Descriptor List for Wheat (Triticum spp.). International Board for Plant Genetic Resources. Rome, Italy.
Moghaddam, M., Ehdaie, B., Waines, J.G. 1997. Genetic variation and interrelationships of agronomic characters in landraces of bread wheat from southeastern Iran. Euphytica 95:361–369.
Moradkhani, H., Pour-Aboughadareh, A., Mehrabi, A.A., Etminan, A. 2012. Evaluation of genetic relationships of Triticum–Aegilops species possessing D genome in different ploidy levels using microsatellites. Int. J. Agri. Crop. Sci. 23:1746–1751.
Moradkhani, H., Mehrabi, A.A., Etminan, A., Pour-Aboughadareh, A. 2015. Molecular diversity and phylogeny of Triticum-Aegilops species possessing D genome revealed by SSR and ISSR markers. Plant. Breed. Seed. Sci. 71:82–95.
Naghavi, M., Malaki, M., Alizadeh, H., Pirseiedi, M., Mardi, M. 2010. An assessment of genetic diversity in wild diploid wheat Triticum boeoticum from west of Iran using RAPD, AFLP and SSR markers. J. Agr. Sci. Tech. 11:585–598.
Nei, M. 1978. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetic 89:583–590.
Pathak, N.N., Nema, D.P. 1985. Genetic advance in landrace of wheat. Indian. J. Agri. Sci. 55:478–479.
Peakall, R., Smouse, P.E. 2006. GENALEX 6: genetic analysis in excel. Population genetic software for teaching and research. Mol. Ecol. Notes. 6:288–295.
Perrier, X., Flori, A., Bonnot, F. 2003. Data analysis methods. In: Hamon, P., Seguin, M., Perrier, X., Glaszmann, J.C. (eds), Genetic Diversity of Cultivated Tropical Plants. London, Plymouth, UK. pp. 43–76.
Poczai, P., Varga, I., Laos, M., Cseh, A., Bell, N., Valkonen, J.P. Hyvonen, J. 2013. Advances in plant genetargeted and functional markers: a review. Plant. Methods 9:1–31.
Pour-Aboughadareh, A., Mohmoudi, M., Ahmadi, J., Moghaddam, M., Mehrabi, A.A., Alavikia, S.S. 2017. Agro-morphological and molecular variability in Triticum boeoticum accessions from Zagros Mountains, Iran. Genet. Resour. Crop. Evol. 64:545–556.
Powell, W., Morgante, M., Andre, C., Mm, Hanafey, Vogel, J., Tingey, S., Rafalski, A. 1996. The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol. Breed. 2:225–238.
Pritchard, J.K., Stephens, M., Donnelly, P. 2000. Inference of population structure using multilocus genotype data. Genetics 155:945–959.
Rajesh, M.K., Sabana, A.A., Rachana, K.E., Rahman, S., Jerard, B.A., Karun, A. 2015. Genetic relationship and diversity among coconut (Cocos nucifera L.) accessions revealed through SCoT analysis. 3 Bitech. 5:999–1006.
Rawashdeh, N.K., Haddad, N.I., Al-Ajlouni, M.M., Turk, M.A. 2007. Phenotypic diversity of durum wheat (Triticum durum Desf.) from Jordan. Genet. Resour. Crop. Evol. 54:129–138.
Rholf, F. 2000. Numerical Taxonomy and Multivariate Analysis System: version 2.10. Exeter Software, Setauket. New York, USA.
Salimi, A., Ebrahimzadeh, H., Taeb, M. 2005. Description of Iranian diploid wheat resources. Genet. Resour. Crop. Evol. 52:351–361.
van Slageren, M.W. 1994. Wild wheats: a monograph of Aegilops L. and Amblyopyrum (Jaub. & Spach) Eig (Poaceae). Wageningen Agricultural University Papers. Wageningen, The Netherlands.
Wolfinger, R.D., Federer, W.T., Cordero-Brana, O. 1997. Recovering information in augmented designs, using SAS PROC GLM and PROC MIXED. Agron. J. 89:856–859.
Wright, S. 1951. The genetical structure of populations. Ann. Eugen. 15:323–354.
Xiong, F., Zhong, R., Han, Z., Jiang, J., He, L., Zhuang, W., Tang, R. 2011. Start codon targeted polymorphism for evaluation of functional genetic variation and relationships in cultivated peanut (Arachis hypogaea L.) varieties. Mol. Biol. Rep. 38:3487–3494.
Yeh, F., Yang, R., Boyle, T. 1997. POPGENE, the User Friendly Shareware for Population Genetic Analysis. Molecular Biology and Biotechnology Center. University of Alberta. Edmonton, Canada.
Zhang, Z., Gao, J., Kong, D., Wang, A., Tang, S., Li, Y., Pang, X. 2015. Assessing genetic diversity in Ziziphus jujuba ‘Jinsixi-aozao’ using morphological and microsatellite (SSR) markers. Biochem. Syst. Ecol. 61:196–202.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Börner
Electronic Supplementary Material (ESM)
42976_2017_4504574_MOESM1_ESM.pdf
Assessment of Genetic Diversity among Iranian Triticum Germplasm using Agro-morphological Traits and Start Codon Targeted (SCoT) Markers
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Pour-Aboughadareh, A., Ahmadi, J., Mehrabi, A.A. et al. Assessment of Genetic Diversity among Iranian Triticum Germplasm Using Agro-morphological Traits and Start Codon Targeted (SCoT) Markers. CEREAL RESEARCH COMMUNICATIONS 45, 574–586 (2017). https://doi.org/10.1556/0806.45.2017.033
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1556/0806.45.2017.033