Abstract
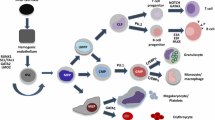
AML1/Runx1, originally identified as a gene located at the breakpoint of the t(8;21) translocation, encodes a transcription factor that is widely expressed in multiple hematopoietic lineages and that regulates the expression of a variety of hemato-poietic genes. Numerous studies have shown that AML1 is a critical regulator of hematopoietic development. In addition, AML1 is a frequent target for chromosomal translocation in human leukemia.The activity of AML1 can be modulated by various types of posttranslational modification, including phosphorylation and acetylation. Phosphorylation by extracellular signal-regulated kinase (ERK) is one of the mechanisms that dictate whether AML1 acts as either a transcriptional repressor or an activator of gene expression. Recently, a physiological role for AML1 in adult hematopoiesis was revealed by conditional gene targeting in mice. Remarkably, adult hematopoietic progenitors are maintained even in the absence of AML1, in stark contrast to the total disruption of definitive hematopoiesis during embryogenesis. AML1 is, however, critical for megakaryopoiesis and plays an important role in T-cell and B-cell development in adult mice. Recent analyses engineered to recreate hematopoiesis in vitro revealed that the transcriptional activity of AML1 is closely related with the potential of AML1 to generate hematopoietic cells and support thymocyte development.
Similar content being viewed by others
References
Okuda T, Nishimura M, Nakao M, Fujita Y. RUNX1/AML1: a central player in hematopoiesis. Int J Hematol. 2001;74:252–257.
Yamagata T, Maki K, Mitani K. Runx1/AML1 in normal and abnormal hematopoiesis. Int J Hematol. 2005;82:1–8.
Miyoshi H, Shimizu K, Kozu T, Maseki N, Kaneko Y, Ohki M. t(8;21) breakpoints on chromosome 21 in acute myeloid leukemia are clustered within a limited region of a single gene, AML1. Proc Natl Acad Sci U S A. 1991;88:10431–10434.
Taniuchi I, Osato M, Egawa T, et al. Differential requirements for Runx proteins in CD4 repression and epigenetic silencing during T lymphocyte development. Cell. 2002;111:621–633.
Bertrand JY, Giroux S, Golub R, et al. Characterization of purified intraembryonic hematopoietic stem cells as a tool to define their site of origin. Proc Natl Acad Sci U S A. 2005;102:134–139.
Cumano A, Dieterlen-Lievre F, Godin I. Lymphoid potential, probed before circulation in mouse, is restricted to caudal intraembryonic splanchnopleura. Cell. 1996;86:907–916.
Okuda T, van Deursen J, Hiebert SW, Grosveld G, Downing JR. AML1, the target of multiple chromosomal translocations in human leukemia, is essential for normal fetal liver hematopoiesis. Cell. 1996;84:321–330.
Cai Z, de Bruijn M, Ma X, et al. Haploinsufficiency of AML1 affects the temporal and spatial generation of hematopoietic stem cells in the mouse embryo. Immunity. 2000;13:423–431.
Wang Q, Stacy T, Miller JD, et al. The CBFβ subunit is essential for CBFα2 (AML1) function in vivo. Cell. 1996;87:697–708.
North TE, de Bruijn MF, Stacy T, et al. Runx1 expression marks long-term repopulating hematopoietic stem cells in the midgestation mouse embryo. Immunity. 2002;16:661–672.
Takakura N,Watanabe T,Suenobu S, et al. A role for hematopoietic stem cells in promoting angiogenesis. Cell. 2000;102:199–209.
Burns CE, DeBlasio T, Zhou Y, Zhang J, Zon L, Nimer SD. Isolation and characterization of runxa and runxb, zebrafish members of the runt family of transcriptional regulators. Exp Hematol. 2002;30:1381–1389.
Kalev-Zylinska ML, Horsfield JA, Flores MV, et al. Runx1 is required for zebrafish blood and vessel development and expression of a human RUNX1-CBF2T1 transgene advances a model for studies of leukemogenesis. Development. 2002;129:2015–2030.
Liu P,Tarle SA, Hajra A, et al. Fusion between transcription factor CBFβ/PEBP2β and a myosin heavy chain in acute myeloid leukemia. Science. 1993;261:1041–1044.
Golub TR, Barker GF, Bohlander SK, et al. Fusion of the TEL gene on 12p13 to the AML1 gene on 21q22 in acute lymphoblastic leukemia. Proc Natl Acad Sci U S A. 1995;92:4917–4921.
Mitani K, Ogawa S, Tanaka T, et al. Generation of the AML1-EVI-1 fusion gene in the t(3;21)(q26;q22) causes blastic crisis in chronic myelocytic leukemia. EMBO J. 1994;13:504–510.
Hiebert SW, Sun W, Davis JN, et al. The t(12;21) translocation converts AML-1B from an activator to a repressor of transcription. Mol Cell Biol. 1996;16:1349–1355.
Tanaka T, Mitani K, Kurokawa M, et al. Dual functions of the AML1/Evi-1 chimeric protein in the mechanism of leukemogenesis in t(3;21) leukemias. Mol Cell Biol. 1995;15:2383–2392.
Okuda T, Cai Z, Yang S, et al. Expression of a knocked-in AML1-ETO leukemia gene inhibits the establishment of normal definitive hematopoiesis and directly generates dysplastic hematopoietic progenitors. Blood. 1998;91:3134–3143.
Castilla LH, Wijmenga C, Wang Q, et al. Failure of embryonic hematopoiesis and lethal hemorrhages in mouse embryos heterozygous for a knocked-in leukemia gene CBFB-MYH11. Cell. 1996;87:687–696.
Perry C, Eldor A, Soreq H. Runx1/AML1 in leukemia: disrupted association with diverse protein partners. Leuk Res. 2002;26:221–228.
Kitabayashi I, Yokoyama A, Shimizu K, Ohki M. Interaction and functional cooperation of the leukemia-associated factors AML1 and p300 in myeloid cell differentiation. EMBO J. 1998;17:2994–3004.
Yamaguchi Y, Kurokawa M,Imai Y,et al. AML1 is functionally regulated through p300-mediated acetylation on specific lysine residues. J Biol Chem. 2004;279:15630–15638.
Imai Y, Kurokawa M,Tanaka K, et al. TLE, the human homolog of Groucho, interacts with AML1 and acts as a repressor of AML1- induced transactivation. Biochem Biophys Res Commun. 1998;252:582–589.
Lutterbach B, Westendorf JJ, Linggi B, Isaac S, Seto E, Hiebert SW. A mechanism of repression by acute myeloid leukemia-1, the target of multiple chromosomal translocations in acute leukemia. J Biol Chem. 2000;275:651–656.
Tanaka T, Kurokawa M, Ueki K, et al. The extracellular signal-regulated kinase pathway phosphorylates AML1, an acute myeloid leukemia gene product, and potentially regulates its transactivation ability. Mol Cell Biol. 1996;16:3967–3979.
Imai Y, Kurokawa M, Yamaguchi Y, et al. The corepressor mSin3A regulates phosphorylation-induced activation, intranuclear location, and stability of AML1. Mol Cell Biol. 2004;24:1033–1043.
Minucci S, Pelicci PG. Histone deacetylase inhibitors and the promise of epigenetic (and more) treatments for cancer. Nat Rev Cancer. 2006;6:38–51.
Hayashi K, Natsume W, Watanabe T, et al. Diminution of the AML1 transcription factor function causes differential effects on the fates of CD4 and CD8 single-positive T cells. J Immunol. 2000;165:6816–6824.
Hayashi K, Abe N, Watanabe T, et al. Overexpression of AML1 transcription factor drives thymocytes into the CD8 single-positive lineage. J Immunol. 2001;167:4957–4965.
Komine O, Hayashi K, Natsume W, et al. The Runx1 transcription factor inhibits the differentiation of naive CD4+ T cells into the Th2 lineage by repressing GATA3 expression. J Exp Med. 2003;198:51–61.
Ichikawa M, Asai T, Saito T, et al. AML-1 is required for megakary- ocytic maturation and lymphocytic differentiation, but not for maintenance of hematopoietic stem cells in adult hematopoiesis. Nat Med. 2004;10:299–304.
Growney JD, Shigematsu H, Li Z, et al. Loss of Runx1 perturbs adult hematopoiesis and is associated with a myeloproliferative phenotype. Blood. 2005;106:494–504.
Mukouyama Y, Hara T, Xu M, et al. In vitro expansion of murine multipotential hematopoietic progenitors from the embryonic aorta-gonad-mesonephros region. Immunity. 1998;8:105–114.
Okuda T, Takeda K, Fujita Y, et al. Biological characteristics of the leukemia-associated transcriptional factor AML1 disclosed by hematopoietic rescue of AML1-deficient embryonic stem cells by using a knock-in strategy. Mol Cell Biol. 2000;20:319–328.
Nakano T, Kodama H, Honjo T. In vitro development of primitive and definitive erythrocytes from different precursors. Science. 1996;272:722–724.
Goyama S, Yamaguchi Y, Imai Y, et al. The transcriptionally active form of AML1 is required for hematopoietic rescue of the AML1-deficient embryonic para-aortic splanchnopleural (P-Sp) region. Blood. 2004;104:3558–3564.
Nishimura M, Fukushima-Nakase Y, Fujita Y, et al. VWRPY motifdependent and -independent roles of AML1/Runx1 transcription factor in murine hematopoietic development. Blood. 2004;103:562–570.
Fukushima-Nakase Y, Naoe Y, Taniuchi I, Hosoi H, Sugimoto T, Okuda T. Shared and distinct roles mediated through C-terminal subdomains of acute myeloid leukemia/Runt-related transcription factor molecules in murine development. Blood. 2005;105:4298–4307.
Schmitt TM, Zuniga-Pflucker JC. Induction of T cell development from hematopoietic progenitor cells by Delta-like-1 in vitro. Immunity. 2002;17:749–756.
Kawazu M, Asai T, Ichikawa M, et al. Functional domains of Runx1 are differentially required for CD4 repression, TCRβ expression, and CD4/8 double-negative to CD4/8 double-positive transition in thymocyte development. J Immunol. 2005;174:3526–3533.
Sun W, Downing JR. Haploinsufficiency of AML1 results in a decrease in the number of LTR-HSCs while simultaneously inducing an increase in more mature progenitors. Blood. 2004;104:3565–3572.
Mikkola HK, Klintman J, Yang H, et al. Haematopoietic stem cells retain long-term repopulating activity and multipotency in the absence of stem-cell leukaemia SCL/tal-1 gene. Nature. 2003;421:547–551.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Kurokawa, M. AML1/Runx1 as a Versatile Regulator of Hematopoiesis: Regulation of Its Function and a Role in Adult Hematopoiesis. Int J Hematol 84, 136–142 (2006). https://doi.org/10.1532/IJH97.06070
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1532/IJH97.06070