Abstract
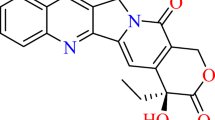
The studies of novel inhibitors of DNA topoisomerase I (Topo I) have already become very promising in cancer chemotherapy. Identifying the new drug-binding residues is playing an important role in the design and optimization of Topo I inhibitors. The designed compounds may have novel scaffolds, thus will be helpful to overcome the toxicities of current camptothecin (CPT) drugs and may provide a solution to cross resistance with these drugs. Multiple sequence alignments were performed on eukaryotic DNA topoisomerase I superfamily and thus the evolutionary tree was constructed. The Evolutionary Trace method was applied to identify functionally important residues of human Topo I. It has been demonstrated that class-specific hydrophobic residues Ala351, Met428, Pro431 are located around the 7,9-position of CPT, indicating suitable substitution of hydrophobic group on CPT will increase antitumor activity. The conservative residue Lys436 in the superfamily is of particular interest and new CPT derivatives designed based on this residue may greatly increase water solubility of such drugs. It has also been demonstrated that the residues Asn352 and Arg364 were conservative in the superfamily, whose mutation will render CPT resistance. As our molecular docking studies demonstrated they did not make any direct interaction with CPT, they are important drug-binding site residues for future design of novel non-camptothecin lead compounds. This work provided a strong basis for the design and synthesis of novel highly potent CPT derivatives and virtual screening for novel lead compounds.
Similar content being viewed by others
References
Zhang, Q., Ye, Y. Z., Ding, D. F., Structural basis of functional specificity, Acta Biophysica Sinica, 2001, 17: 687–694.
Lichtarge, O., Bourne, H. R., Cohen, F. E., Evolutionarily conserved Gαβγ binding surfaces support a model of the G protein-receptor complex, Proc. Natl. Acad. Sci. USA, 1996, 93: 7507–7511.
Lichtarge, O., Bourne, H. R., Cohen, F. E., An evolutionary trace method defines binding surfaces common to protein families, J. Mol. Biol., 1996, 257: 342–358.
Madabushi, S., Yao, H., Marsh, M. et al., Structural clusters of evolutionary trace residues are statistically significant and common in proteins, J. Mol. Biol., 2002, 316: 139–154.
Song, Y. L., Zhang, W. N., Ji, H. T. et al., Flexible molecular docking studies of antineoplastic camptothecins derivatives on DNA-topoisomerase complex, Acta Chimica Sinica, 2003, 61: 1860–1866.
Redinbo, M. R., Stewart, L., Kuhn, P. et al., Crystal structures of human topoisomerase I in covalent and noncovalent complexes with DNA, Science, 1998, 279: 1504–1513.
Fujimori, A., Harker, W. G., Kohlhagen, G. et al., Mutation at the catalytic site of topoisomerase I in CEM/C2, a human leukemia cell line resistant to camptothecin, Cancer Res., 1995, 55: 1339–1346.
Tamura, H., Kohchi, C., Yamada, R. et al., Molecular cloning of a cDNA of a camptothecin resistant human DNA topoisomerase I and identification of mutation sites, Nucleic Acids Res., 1991, 19: 69–75.
Rubin, E., Pantazis, P., Bharti, A. et al., Identification of a mutant human topoisomerase I with intact catalytic activity and resistance to 9-nitro-camptothecin, J. Biol. Chem., 1994, 269: 2433–2439.
Benedetti, P., Fiorani, P., Capuani, L. et al., Camptothecin resistance from a single mutation changing glycine 363 of human DNA topoisomerase I to cysteine, Cancer Res., 1993, 53: 4343–4348.
Pond, C. D., Li, X. G., Rubin, E. H. et al., Effects of mutations in the F361 to R364 region of topoisomerase I (Topo I), in the presence and absence of 9-aminocamptothecin, on the Topo I-DNA interaction, Anticancer Drugs, 1999, 10: 647–653.
Urasaki, Y., Laco, G. S., Pourquier, P. et al., Characterization of a novel topoisomerase I mutation from a camptothecin-resistant human prostate cancer cell line, Cancer Res., 2001, 61: 1964–1969.
Laco, G. S., Collins, J. R., Luke, B. T. et al., Human topoisomerase I inhibition: Docking camptothecin and derivatives into a structure-based active site model, Biochemistry, 2002, 41: 1428–1435.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Song, Y., Qi, Y., Zhang, W. et al. Evolutionary trace analysis of eukaryotic DNA topoisomerase I superfamily: Identification of novel antitumor drug binding site. Sci. China Ser. C.-Life Sci. 48, 375–384 (2005). https://doi.org/10.1360/062004-20
Received:
Issue Date:
DOI: https://doi.org/10.1360/062004-20