Abstract
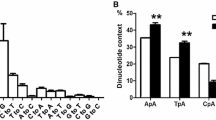
Proviral DNA was extracted from donkey leukocyte infected with Chinese donkey leukocyte attenuated equine infectious anemia virus (DLA-EIAV), and peripheral blood lymphocytes (PBL) from a horse infected with the virulent EIAV strain Liaoning (EIAV L). The entire proviral DNA from both viruses was cloned and sequenced. The lengths of complete genomic sequences of DLA-EIAV and EIAV L provirus were 8266 bp and 8235 bp, respectively. Sequence comparison indicated that DLA-EIAV shares 97.0% and 97.5% in sequence homology with EIAV L and donkey-adapted EIAV (DA-EIAV), respectively. Lots of variations occurred in long terminal repeat (LTR, consisting of U3, R, U5), ORF S2, and env regions between DLA-EIAV and EIAV L. The nucleotide sequence differences of the two viruses in U3, R, U5, ORF S2, and env are 13.2%, 7.5%, 5.1%, 3.9%, and 2.7%, respectively, and predicted amino acid sequence differences in env and S2 coding regions are 4.4% and 8.8%, respectively. Six conserved regions are characterized in Gp90. There is a cis-activating GATA motif in ENH of DLA-EIAV and EIAV L. Two N-linked glycosylation sites disappeared in DLA-EIAV Gp90 in comparison with that of EIAV L. A bHLH transcription factor binding consensus sequence was found in LTR of DLA-EIAV but not in EIAV L. Furthermore, there is a mutation in the stem of DLA-EIAV TAR resulting in formation of a uridine tuber. Further study is needed to uncover the relationship between sequence changes and their biological functions of DLA-EIAV and L.
Similar content being viewed by others
References
Crawford, T. B., Wardrop, K. J., Tornquist, S. J. et al., A primary production deficit in the thrombocytopenia of equine infectious anemia, J. Virol., 1996, 70: 7842–7850.
Coggins, L., Carriers of equine infectious anemia virus, J. Am. Vet. Med. Assoc., 1984, 184: 279–281.
Wang, L., Yu, L., Rong, J. G. et al., Integration dynamics of proviral DNA in fetal donkey dermal cell infected by attenuated strain of equine infectious anemia virus, Chinese Journal of Virology, 1997, 13(3): 235–239.
Wang, L., Yu, L., Zhang, S. J. et al., Cloning and sequence analysis of Chinese attenuated equine infectious anemia virus (EIAV) long terminal repeat, Chinese Journal of Veterinary Science, 1998, 18(1): 1–5.
Shen, R. X., Xu, Z. D., He, Y. S. et al., Immune study of equine infectious anemia virus, Scientia Agricultura Sinica, 1979, (4): 1–15.
Sambrook, J., Fritsch, E. F., Maniatis, T., Molecular Cloning: A Laboratory Manual, 2nd ed., New York: Cold Spring Harbor Laboratory Press, 1989.
Kawakami, T., Sherman, L., Dahlbery, J. et al., Nucleotide sequence analysis of equine infectious anemia virus proviral DNA, Virology, 1987, 158: 300–312.
Payne, S. L., Rausch, J., Rushlow, K. et al., Characterization of infectious molecular clones of equine infectious anaemia virus, J. Gene Virol., 1994, 75: 425–429.
Payne, S. L., Qi, X. M., Shao, H. et al., Disease induction by virus derived from molecular clones of equine infectious anemia virus, J. Virol., 1998, 72: 483–487.
Jacks, T., Power, M. D., Masiarz, F. R. et al., Characterization of ribosomal frameshifting in HIV-1 gag-pol expression, Nature, 1988, 331: 280–283.
Zhang, W., David, B. A., Oaks, J. L. et al., Natural variation of equine infectious anemia Gag protein cytotoxic T lymphocyte epitopes, Virology, 1999, 261: 242–252.
Lonning, S. M., Zhang, W., Leib, S. R. et al., Detection and induction of equine infectious anemia virus specific cytotoxic T lymphocyte responses by use of recombinant retroviral vectors, J. Virol., 1999, 73: 2762–2769.
Lu, J. L., He, Z. Z., Lin, X. Y. et al., Cell-mediated immunity of equine infectious anemia Virus, Scientia Agricultura Sinica, 1981, 5: 83–88.
Chackerian, B. L., Rudensey, M., Overbaugh, J., Specific N-linked and O-linked glycosylation modification by neutralizing antibodies, J. Virol., 1997, 71: 7719–7727.
Katz, J. M., Naeve, C. W., Webster, R. G., Host-cell mediated variation in H3N2 influenza virus, Virology, 1987, 156: 386–395.
Ito, E., Toki, T., Ishihara, H. et al., Erythroid transcription factor GATA-lis abundantly transcribed in mouse testis, Nature, 1993, 362: 466–468.
Narayan, R. R., Henderson, L. E., Sowder, R. C. et al., Biology and pathogenesis of equine infectious anemia virus, J. Gen. Virol., 1989, 70: 1617–1639.
Nielsen, A. L., Nørby, P. L., Pedersen, F. S. et al., Various modes of basic helix-loop-helix protein-mediated regulation of murine leukemia virus transcription in lymohoid cell lines, J. Virol., 1996, 70: 5892–5901.
Carroll, R., Martarano, L., Derse, D., Identification of lentivirus Tat functional domains through generation of equine infectious anemia virus/human immunodeficiency virus type 1 Tat gene chimeras, J. Virol., 1991, 65: 3460–3467.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, L., Tong, G., Liu, H. et al. Proviral genomic sequence analysis of Chinese donkey leukocyte attenuated equine infectious anemia virus vaccine and its parental virus strain Liaoning. Sci. China Ser. C.-Life Sci. 45, 57–67 (2002). https://doi.org/10.1360/02yc9007
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1360/02yc9007