Abstract
Background
Appendiceal mucinous neoplasm (AMN) with peritoneal metastasis is a rare but deadly disease with few prognostic or therapy-predictive biomarkers to guide treatment decisions. Here, we investigated the prognostic and biological attributes of gene expression-based AMN molecular subtypes.
Methods
AMN specimens (n = 138) derived from a population-based subseries of patients treated at our institution with cytoreductive surgery and hyperthermic intraperitoneal chemotherapy (CRS/HIPEC) between 05/2000 and 05/2013 were analyzed for gene expression using a custom-designed NanoString 148-gene panel. Signed non-negative matrix factorization (sNMF) was used to define a gene signature capable of delineating robustly-classified AMN molecular subtypes. The sNMF class assignments were evaluated by topology learning, reverse-graph embedding and cross-cohort performance analysis.
Results
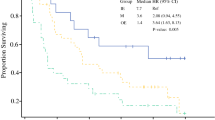
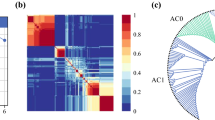
Three molecular subtypes of AMN were discerned by the expression patterns of 17 genes with roles in cancer progression or anti-tumor immunity. Tumor subtype assignments were confirmed by topology learning. AMN subtypes were termed immune-enriched (IE), oncogene-enriched (OE) and mixed (M) as evidenced by their gene expression patterns, and exhibited significantly different post-treatment survival outcomes. Genes with specialized immune functions, including markers of T-cells, natural killer cells, B-cells, and cytolytic activity showed increased expression in the low-risk IE subtype, while genes implicated in the promotion of cancer growth and progression were more highly expressed in the high-risk OE subtype. In multivariate analysis, the subtypes demonstrated independent prediction power for post-treatment survival.
Conclusions
Our findings suggest a greater role for the immune system in AMN than previously recognized. AMN subtypes may have clinical utility for predicting CRS/HIPEC treatment outcomes.



Similar content being viewed by others
References
Trivedi AN, Levine EA, Mishra G. Adenocarcinoma of the appendix is rarely detected by colonoscopy. J Gastrointest Surg. 2009;13(4):668–675.
Marmor S, Portschy PR, Tuttle TM, Virnig BA. The rise in appendiceal cancer incidence: 2000-2009. J Gastrointest Surg. 2015;19(4):743–750.
McCusker ME, Cote TR, Clegg LX, Sobin LH. Primary malignant neoplasms of the appendix: a population-based study from the surveillance, epidemiology and end-results program, 1973–1998. Cancer. 2002;94(12):3307–3312.
Stewart JHt, Shen P, Russell GB, et al. Appendiceal neoplasms with peritoneal dissemination: outcomes after cytoreductive surgery and intraperitoneal hyperthermic chemotherapy. Ann Surg Oncol. 2006;13(5):624–634.
Sugarbaker PH. New standard of care for appendiceal epithelial neoplasms and pseudomyxoma peritonei syndrome? Lancet Oncol. 2006;7(1):69-76.
Smith JW, Kemeny N, Caldwell C, Banner P, Sigurdson E, Huvos A. Pseudomyxoma peritonei of appendiceal origin. The Memorial Sloan-Kettering Cancer Center experience. Cancer. 1992;70(2):396–401.
Baratti D, Kusamura S, Nonaka D, et al. Pseudomyxoma peritonei: clinical pathological and biological prognostic factors in patients treated with cytoreductive surgery and hyperthermic intraperitoneal chemotherapy (HIPEC). Ann Surg Oncol. 2008;15(2):526–534.
Votanopoulos KI, Russell G, Randle RW, Shen P, Stewart JH, Levine EA. Peritoneal surface disease (PSD) from appendiceal cancer treated with cytoreductive surgery (CRS) and hyperthermic intraperitoneal chemotherapy (HIPEC): overview of 481 cases. Ann Surg Oncol. 2015;22(4):1274–1279.
Levine EA, Votanopoulos KI, Qasem SA, et al. Prognostic molecular subtypes of low-grade cancer of the appendix. J Am Coll Surg. 2016;222(4):493–503.
Levine EA, Blazer DG, 3rd, Kim MK, et al. Gene expression profiling of peritoneal metastases from appendiceal and colon cancer demonstrates unique biologic signatures and predicts patient outcomes. J Am Coll Surg. 2012;214(4):599–606. (discussion 606-597).
Bradley RF, Stewart JHt, Russell GB, Levine EA, Geisinger KR. Pseudomyxoma peritonei of appendiceal origin: a clinicopathologic analysis of 101 patients uniformly treated at a single institution, with literature review. Am J Surg Pathol. 2006;30(5):551–559.
Levine EA, Stewart JHt, Shen P, Russell GB, Loggie BL, Votanopoulos KI. Intraperitoneal chemotherapy for peritoneal surface malignancy: experience with 1000 patients. J Am Coll Surg. 2014;218(4):573–585.
Liu C, Su J, Yang F, Wei K, Ma J, Zhou X. Compound signature detection on LINCS L1000 big data. Mol Biosyst. 2015;11(3):714–722.
Benjamini Y, Hochberg Y. Controlling the false discovery rate—a practical and powerful approach to multiple testing. J Roy Stat Soc B Met. 1995;57(1):289–300.
Mao Q, Wang L, Goodison S, Sun Y. Dimensionality reduction via graph structure learning. In: Proceedings of the 21th ACM SIGKDD international conference on knowledge discovery and data mining; 2015; Sydney, NSW, Australia.
Mao Q, Wang L, Tsang IW, Sun Y. A novel regularized principal graph learning framework on explicit graph representation. arXiv preprint arXiv:1512.02752. 2015.
Qiu X, Hill A, Packer J, Lin D, Ma YA, Trapnell C. Single-cell mRNA quantification and differential analysis with Census. Nat Methods. 2017;14(3):309–315.
Kattan MW, Hess KR, Amin MB, et al. American Joint Committee on Cancer acceptance criteria for inclusion of risk models for individualized prognosis in the practice of precision medicine. CA Cancer J Clin. 2016;66(5):370–374.
Reilly BM, Evans AT. Translating clinical research into clinical practice: impact of using prediction rules to make decisions. Ann Intern Med. 2006;144(3):201–209.
Acknowledgements
The authors acknowledge the DEMON high performance computing cluster, the Greenplum massively parallel processing database and the Data Lake cloud storage and computing facility at Wake Forest University School of Medicine, the Texas Advanced Computing Center (TACC) at The University of Texas at Austin (http://www.tacc.utexas.edu), and the Extreme Science and Engineering Discovery Environment (XSEDE, which is supported by National Science Foundation grant number ACI-1548562), for providing high performance computing resources that have contributed to the research results reported within this paper.
Funding
This work was supported, in part, by the Orin Smith Family fund (to E.A.L.), pilot funds from the National Organization for Rare Disorders (to E.A.L. and L.D.M.) and the Wake Forest Baptist Compressive Cancer Center’s Shared Resources: Cancer Genomics (CGSR), Tumor Tissue & Pathology (TTPSR) and Bioinformatics (BISR) supported by the National Cancer Institute’s Cancer Center Support Grant award number P30CA012197. The content of this publication is solely the responsibility of the authors and does not necessarily represent the official views of the National Cancer Institute.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Su, J., Jin, G., Votanopoulos, K.I. et al. Prognostic Molecular Classification of Appendiceal Mucinous Neoplasms Treated with Cytoreductive Surgery and Hyperthermic Intraperitoneal Chemotherapy. Ann Surg Oncol 27, 1439–1447 (2020). https://doi.org/10.1245/s10434-020-08210-5
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-020-08210-5