Abstract
Background
Alterations in DNA methylation have been demonstrated in a variety of malignancies, including papillary thyroid cancer (PTC). The full extent of dysregulation in PTC and the downstream affected pathways remains unclear. Here we report a genome-wide analysis of PTC methylation, the dysregulation of various canonical pathways, and assess its potential as a diagnostic test.
Methods
A discovery set utilized 49 PTCs and matched normal controls from The Cancer Genome Atlas. Another set of 16 PTCs and 13 normal controls were used as a replication set. Genome-wide methylation analysis was done using Illumina 450 K methylation chips. Differentially methylated loci (DML) were identified by comparing PTC and matched normal tissues. DML were defined as false-discovery rate p < 0.05 and absolute Δβ ≥ 0.2. DML were then analyzed for pathway and disease commonalities using Qiagen Ingenuity Pathway Analysis.
Results
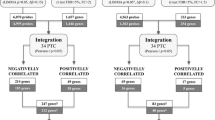
Of 485,577 CpG sites analyzed, 1226 DML were identified in our discovery and replication sets, and 1061 (86.5 %) DML showed hypomethylation when comparing tumor with normal tissue. Support vector machine classification was able to differentiate benign from malignant tissue in 107 (94.7 %) of 113 tested samples, including 15 (83.3 %) of 18 samples lacking a clearly deleterious mutation. Statistically significant associations with multiple canonical pathways, diseases, and biofunctions were observed including PI3K, PTEN, wnt/β-catenin, and p53.
Conclusions
Epigenetic dysregulation of multiple canonical pathways are associated with the development of PTC. This methylation signature shows promise as a future adjunctive screening test for thyroid nodules.

Similar content being viewed by others
References
Baylin SB, Herman JG, Graff JR, Vertino PM, Issa JP. Alterations in DNA methylation: a fundamental aspect of neoplasia. Adv Cancer Res. 1998;72:141–96.
Baylin SB, Herman JG. DNA hypermethylation in tumorigenesis: epigenetics joins genetics. Trends Genet. 2000;16:168–74.
Clark SJ, Melki J. DNA methylation and gene silencing in cancer: which is the guilty party? Oncogene. 2002;21:5380–7.
Hou P, Liu D, Xing M. Genome-wide alterations in gene methylation by the BRAF V600E mutation in papillary thyroid cancer cells. Endocr Relat Cancer. 2011;18:687–97.
Xing M. Gene methylation in thyroid tumorigenesis. Endocrinology. 2007;148:948–53.
Faam B, Ghaffari MA, Ghadiri A, Azizi F. Epigenetic modifications in human thyroid cancer. Biomed Rep. 2015;3:3–8.
Rodriguez-Rodero S, Fernandez AF, Fernandez-Morera JL, Castro-Santos P, Bayon GF, Ferrero C, et al. DNA methylation signatures identify biologically distinct thyroid cancer subtypes. J Clin Endocrinol Metab. 2013;98:2811–21.
Ellis RJ, Wang Y, Stevenson HS, Boufraqech M, Patel D, Nilubol N, et al. Genome-wide methylation patterns in papillary thyroid cancer are distinct based on histological subtype and tumor genotype. J Clin Endocrinol Metab. 2014;99:E329–37.
Mancikova V, Buj R, Castelblanco E, Inglada-Perez L, Diez A, de Cubas AA, et al. DNA methylation profiling of well-differentiated thyroid cancer uncovers markers of recurrence free survival. Int J Cancer. 2014;135:598–610.
Cancer Genome Atlas Research Network. Integrated genomic characterization of papillary thyroid carcinoma. Cell. 2014;159:676–90.
Hsiao SJ, Nikiforov YE. Molecular approaches to thyroid cancer diagnosis. Endocr Relat Cancer. 2014;21:T301–13.
Medici M, Kwong N, Angell TE, Marqusee E, Kim MI, Frates MC, et al. The variable phenotype and low-risk nature of RAS-positive thyroid nodules. BMC Med. 2015;13:184.
Schalkwyk LC PR, Wong CC, Touleimat wfcbN, Defrance M, Teschendorff A, Maksimovic J. wateRmelon: Illumina 450K methylation array normalization and metrics. R package version 1.5.1. 2013.
R Development Core Team. R: a language and environment for statistical computing. Vienna: R Foundation for Statistical Computing; 2014.
Alvarez-Nnez F, Bussaglia E, Mauricio D, Ybarra J, Vilar M, Lerma E, et al. PTEN promoter methylation in sporadic thyroid carcinomas. Thyroid. 2006;16:17–23.
Hou P, Ji M, Xing M. Association of PTEN gene methylation with genetic alterations in the phosphatidylinositol 3-kinase/AKT signaling pathway in thyroid tumors. Cancer. 2008;113:2440–7.
Rampal R, Alkalin A, Madzo J, Vasanthakumar A, Pronier E, Patel J, et al. DNA hydroxymethylation profiling reveals that WT1 mutations result in loss of TET2 function in acute myeloid leukemia. Cell Rep. 2014;9:1841–55.
Acknowledgment
R.H.G. was supported by award K12CA139160 from the National Cancer Institute (NCI). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NCI or the U.S. National Institutes of Health. We thank Dr. Larissa Furtado for her assistance and expertise in characterizing the somatic mutation profiles of these samples. We thank Dr. Nicole Cipriani for her assistance and expertise in endocrine pathology evaluations. The authors used data generated by TCGA managed by the NCI and National Human Genome Research Institute; information about TCGA can be found online at http://cancergenome.nih.gov.
Disclosure
Michael G. White, Sapna Nagar, Briseis Aschebrook-Kilfoy, Farzana Jasmine, Muhammad G. Kibriya, Habibul Ahsan, Peter Angelos, Edwin L. Kaplan, and Raymon H. Grogan declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
White, M.G., Nagar, S., Aschebrook-Kilfoy, B. et al. Epigenetic Alterations and Canonical Pathway Disruption in Papillary Thyroid Cancer: A Genome-wide Methylation Analysis. Ann Surg Oncol 23, 2302–2309 (2016). https://doi.org/10.1245/s10434-016-5185-4
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-016-5185-4