Abstract
Background
Cholangiocarcinoma can be classified in intrahepatic cholangiocarcinoma (ICC) and perihilar cholangiocarcinoma (PCC). Moreover, PCC includes two different forms: extrahepatic (EH) PCC, which arises from the perihilar EH large ducts, and intrahepatic (IH) PCC, in which a significant liver mass invades the perihilar bile ducts. In this study, we investigated the molecular profile and molecular prognostic factors in EH-PCC, IH-PCC, and ICC submitted to curative surgery.
Methods
Ninety-one patients with cholangiocarcinoma (38 EH-PCC, 18 IH-PCC, and 35 ICC), who underwent curative surgery in a single tertiary hepatobiliary surgery referral center were assessed for mutational status in 56 cancer-related genes.
Results
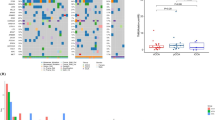
The most frequently mutated genes in EH-PCC were KRAS (47.4 %), TP53 (23.7 %) and ARID1A (15.8 %); in IH-PCC were KRAS (22.2 %), PBRM1 (16.7 %), and PIK3CA (16.7 %); and in ICC were IDH1 (17.1 %), NRAS (17.1 %), and BAP1 (14.3 %). The presence of mutations in ALK, IDH1, and TP53 genes was significantly associated with poor prognosis in patients with EH-PCC (p < 0.001, p = 0.043, and p = 0.019, respectively). Mutation of the TP53 gene was significantly associated with poor prognosis in patients with IH-PCC (p = 0.049). The presence of mutations in ARID1A, PIK3C2G, STK11, TGFBR2, and TP53 genes was significantly associated with poor prognosis in patients with ICC (p = 0.012, p = 0.030, p = 0.030, p = 0.011, and p = 0.011, respectively).
Conclusions
Mutational gene profiling identified different gene mutations in EH-PCC, IH-PCC, and ICC. Moreover, our study reported specific prognostic genes that can identify patients with poor prognosis after curative surgery who may benefit from traditional or target adjuvant treatments.


Similar content being viewed by others
References
de Groen PC, Gores GJ, LaRusso NF, Gunderson LL, Nagorney DM. Biliary tract cancers. N Engl J Med. 1999;341:1368–78.
Patel T. Cholangiocarcinoma—controversies and challenges. Nat Rev Gastroenterol Hepatol. 2011;8:189–200.
International Union Against Cancer (UICC). TNM classification of malignant tumours. 7th ed. Chichester: Wiley-Blackwell; 2010.
American Joint Committee on Cancer (AJCC). Cancer staging manual. 7th ed. New York: Springer; 2010.
Sano T, Shimada K, Sakamoto Y, Ojima H, Esaki M, Kosuge T. Prognosis of perihilar cholangiocarcinoma: hilar bile duct cancer versus intrahepatic cholangiocarcinoma involving the hepatic hilus. Ann Surg Oncol. 2008;15:590–9.
Ebata T, Kamiya J, Nishio H, Nagasaka T, Nimura Y, Nagino M. The concept of perihilar cholangiocarcinoma is valid. Br J Surg. 2009;96:926–34.
Nakeeb A, Pitt HA, Sohn TA, et al. Cholangiocarcinoma. A spectrum of intrahepatic, perihilar, and distal tumors. Ann Surg. 1996;224:463–73.
Miyazaki M, Ohtsuka M, Miyakawa S, et al. Classification of biliary tract cancers established by the Japanese Society of Hepato-Biliary-Pancreatic Surgery: 3rd English edition. J Hepatobiliary Pancreat Sci. 2015;22:181–96.
Jiao Y, Pawlik TM, Anders RA, et al. Exome sequencing identifies frequent inactivating mutations in BAP1, ARID1A and PBRM1 in intrahepatic cholangiocarcinomas. Nat Genet. 2013;45:1470–3.
Voss JS, Holtegaard LM, Kerr SE, et al. Molecular profiling of cholangiocarcinoma shows potential for targeted therapy treatment decisions. Hum Pathol. 2013;44:1216–22.
Borger DR, Zhu AX. IDH mutations: new genetic signatures in cholangiocarcinoma and therapeutic implications. Exp Rev Anticancer Ther. 2012;12:543–6.
Chuang SC, Lee KT, Tsai KB, et al. Immunohistochemical study of DPC4 and p53 proteins in gallbladder and bile duct cancers. World J Surg. 2004;28:995–1000.
Yanagisawa N, Mikami T, Saegusa M, Okayasu I. More frequent beta-catenin exon 3 mutations in gallbladder adenomas than in carcinomas indicate different lineages. Cancer Res. 2001;61:19–22.
Chan-On W, Nairismagi ML, Ong CK, et al. Exome sequencing identifies distinct mutational patterns in liver fluke–related and non-infection-related bile duct cancers. Nat Genet. 2013;45:1474–8.
Churi CR, Shroff R, Wang Y, et al. Mutation profiling in cholangiocarcinoma: prognostic and therapeutic implications. PLoS One. 2014;9:e115383.
Simbolo M, Fassan M, Ruzzenente A, et al. Multigene mutational profiling of cholangiocarcinomas identifies actionable molecular subgroups. Oncotarget. 2014;5:2839–52.
Simbolo M, Gottardi M, Corbo V, et al. DNA qualification workflow for next generation sequencing of histopathological samples. PLoS One. 2013;8:e62692.
Zamo A, Bertolaso A, van Raaij AW, et al. Application of microfluidic technology to the BIOMED-2 protocol for detection of B-cell clonality. J Mol Diagn. 2012;14:30–37.
Sturm PD, Baas IO, Clement MJ, et al. Alterations of the p53 tumor-suppressor gene and K-ras oncogene in perihilar cholangiocarcinomas from a high-incidence area. Int J Cancer. 1998;78:695–8.
Borger DR, Tanabe KK, Fan KC, et al. Frequent mutation of isocitrate dehydrogenase (IDH) 1 and IDH2 in cholangiocarcinoma identified through broad-based tumor genotyping. Oncologist. 2012;17:72–9.
Zhu AX, Borger DR, Kim Y, et al. Genomic profiling of intrahepatic cholangiocarcinoma: refining prognosis and identifying therapeutic targets. Ann Surg Oncol. 2014;21:3827–34.
Xu RF, Sun JP, Zhang SR, et al. KRAS and PIK3CA but not BRAF genes are frequently mutated in Chinese cholangiocarcinoma patients. Biomed Pharmacother. 2011;65:22–6.
Park KW, Jung ES, Kim DG, et al. ERCC1 can be a prognostic factor in hilar cholangiocarcinoma and extrahepatic bile duct cancer, but not in intrahepatic cholangiocarcinoma. Cancer Res Treat. 2013;45:63–9.
Wang J, Wang X, Xie S, et al. p53 status and its prognostic role in extrahepatic bile duct cancer: a meta-analysis of published studies. Dig Dis Sci. 2011;56:655–62.
Robertson S, Hyder O, Dodson R, et al. The frequency of KRAS and BRAF mutations in intrahepatic cholangiocarcinomas and their correlation with clinical outcome. Hum Pathol. 2013;44:2768–73.
Wang P, Dong Q, Zhang C, et al. Mutations in isocitrate dehydrogenase 1 and 2 occur frequently in intrahepatic cholangiocarcinomas and share hypermethylation targets with glioblastomas. Oncogene. 2013;32:3091–100.
Acknowledgment
Supported in part by Associazione Italiana per la Ricerca sul Cancro (AIRC) Grants 12182 and 6421, FP7 EU project CAM-PaC 602783, the Italian Cancer Genome Project Grant from the Italian Ministry of Research (FIRB-RBAP10AHJB), and FIMP–Ministry of Health (CUP_J33G13000210001).
Disclosure
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Andrea Ruzzenente and Matteo Fassan shared first authorship. Calogero Iacono and Aldo Scarpa shared last authorship.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ruzzenente, A., Fassan, M., Conci, S. et al. Cholangiocarcinoma Heterogeneity Revealed by Multigene Mutational Profiling: Clinical and Prognostic Relevance in Surgically Resected Patients. Ann Surg Oncol 23, 1699–1707 (2016). https://doi.org/10.1245/s10434-015-5046-6
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-015-5046-6