Abstract
Background
Methylation changes within tumor suppressor (TS) genes or polycomb group target (PcG) genes alter cell fates. Chromatin associated with PcG targets is bivalent in stem cells, while TS genes are not normally bivalent. PcG target methylation changes have been identified in tumor stem cells, and abnormal methylation is found in TS genes in cancers. If the epigenetic states of genes influence DNA methylation, then methylation of PcG targets and TS genes may evolve differently during cancer development. More importantly, methylation changes may be part of a sequence in tumorigenesis.
Methods
Chromatin and methylation states of 4 PcG targets and 2 TS genes were determined in colon cancer cells. The methylation states were also detected in 100 pairs of colon cancer samples. Principle component analysis (PCA) was used to reveal whether TS methylation or PcG methylation was the main methylation change associated with colon cancers.
Results
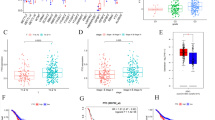
Chromatin and methylation states differ in colon cancer cell lines. The methylation states within PcG targets clustered independently from the methylation states in TS genes, a finding we previously reported in liver cancers. PCA in colon cancers revealed the strongest association with methylation changes in 2 TS genes, HIC1 and RassF1A. Loss of HIC1 methylation correlated with decreased tumor migration.
Conclusions
PcG and TS methylation states cluster independently from each other. The deduced principle component correlated better with TS methylation than PcG methylation in colon cancer. Abnormal methylation changes may represent a sequential biomarker profile to identify developing colon cancer.




Similar content being viewed by others
References
Allan RS, Zueva E, Cammas F, Schreiber HA, Masson V, Belz GT, et al. An epigenetic silencing pathway controlling T helper 2 cell lineage commitment. Nature. 2012;487:249–53.
Bibikova M, Chudin E, Wu B, Zhou L, Garcia EW, Liu Y, et al. Human embryonic stem cells have a unique epigenetic signature. Genome Res. 2006;16:1075–83.
Youn JI, Kumar V, Collazo M, Nefedova Y, Condamine T, Cheng P, et al. Epigenetic silencing of retinoblastoma gene regulates pathologic differentiation of myeloid cells in cancer. Nat Immunol. 2013;14:211–20.
Yusa A, Miyazaki K, Kimura N, Izawa M, Kannagi R. Epigenetic silencing of the sulfate transporter gene DTDST induces sialyl Lewisx expression and accelerates proliferation of colon cancer cells. Cancer Res. 2010;70:4064–73.
Baylin SB, Ohm JE. Epigenetic gene silencing in cancer: a mechanism for early oncogenic pathway addiction? Nat Rev Cancer. 2006;6:107–16.
Hsiao SH, Huang TH, Leu YW. Excavating relics of DNA methylation changes during the development of neoplasia. Semin Cancer Biol. 2009;19:198–208.
Leu YW, Huang TH, Hsiao SH. Epigenetic reprogramming of mesenchymal stem cells. Adv Exp Med Biol. 2013;754:195–211.
Baylin SB. Stem cells, cancer, and epigenetics. In: StemBook. Cambridge: Harvard Stem Cell Institute; 2008.
Hsiao SH, Lee KD, Hsu CC, Tseng MJ, Jin VX, Sun WS, et al. DNA methylation of the Trip10 promoter accelerates mesenchymal stem cell lineage determination. Biochem Biophys Res Commun. 2010;400:305–12.
Laurent L, Wong E, Li G, Huynh T, Tsirigos A, Ong CT, et al. Dynamic changes in the human methylome during differentiation. Genome Res. 2010;20:320–31.
Wamstad JA, Alexander JM, Truty RM, Shrikumar A, Li F, Eilertson KE, et al. Dynamic and coordinated epigenetic regulation of developmental transitions in the cardiac lineage. Cell. 2012;151:206–20.
Sauvageau M, Sauvageau G. Polycomb group proteins: multi-faceted regulators of somatic stem cells and cancer. Cell Stem Cell. 2010;7:299–313.
Valk-Lingbeek ME, Bruggeman SW, van Lohuizen M. Stem cells and cancer; the polycomb connection. Cell. 2004;118:409–18.
Bernstein BE, Mikkelsen TS, Xie X, Kamal M, Huebert DJ, Cuff J, et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell. 2006;125:315–26.
Surface LE, Thornton SR, Boyer LA. Polycomb group proteins set the stage for early lineage commitment. Cell Stem Cell. 2010;7:288–98.
Agger K, Cloos PA, Christensen J, Pasini D, Rose S, Rappsilber J, et al. UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature. 2007;449:731–4.
Lan F, Bayliss PE, Rinn JL, Whetstine JR, Wang JK, Chen S, et al. A histone H3 lysine 27 demethylase regulates animal posterior development. Nature. 2007;449:689–94.
Vastenhouw NL, Zhang Y, Woods IG, Imam F, Regev A, Liu XS, et al. Chromatin signature of embryonic pluripotency is established during genome activation. Nature. 2010;464:922–6.
Margueron R, Reinberg D. The Polycomb complex PRC2 and its mark in life. Nature. 2011;469:343–9.
Balch C, Nephew KP, Huang TH, Bapat SA. Epigenetic “bivalently marked” process of cancer stem cell-driven tumorigenesis. Bioessays. 2007;29:842–5.
Ohm JE, McGarvey KM, Yu X, Cheng L, Schuebel KE, Cope L, et al. A stem cell-like chromatin pattern may predispose tumor suppressor genes to DNA hypermethylation and heritable silencing. Nat Genet. 2007;39:237–42.
Irizarry RA, Ladd-Acosta C, Wen B, Wu Z, Montano C, Onyango P, et al. The human colon cancer methylome shows similar hypo- and hypermethylation at conserved tissue-specific CpG island shores. Nat Genet. 2009;41:178–86.
Gebhard C, Schwarzfischer L, Pham TH, Schilling E, Klug M, Andreesen R, Rehli M. Genome-wide profiling of CpG methylation identifies novel targets of aberrant hypermethylation in myeloid leukemia. Cancer Res. 2006;66:6118–28.
Yegnasubramanian S, Haffner MC, Zhang Y, Gurel B, Cornish TC, Wu Z, et al. DNA hypomethylation arises later in prostate cancer progression than CpG island hypermethylation and contributes to metastatic tumor heterogeneity. Cancer Res. 2008;68:8954–67.
Gaudet F, Hodgson JG, Eden A, Jackson-Grusby L, Dausman J, Gray JW, et al. Induction of tumors in mice by genomic hypomethylation. Science. 2003;300:489–92.
Teng IW, Hou PC, Lee KD, Chu PY, Yeh KT, Jin VX, et al. Targeted methylation of two tumor suppressor genes is sufficient to transform mesenchymal stem cells into cancer stem/initiating cells. Cancer Res. 2011;71:4653–63.
Eggers H, Steffens S, Grosshennig A, Becker JU, Hennenlotter J, Stenzl A, et al. Prognostic and diagnostic relevance of hypermethylated in cancer 1 (HIC1) CpG island methylation in renal cell carcinoma. Int J Oncol. 2012;40:1650–8.
Pehlivan S, Artac M, Sever T, Bozcuk H, Kilincarslan C, Pehlivan M. Gene methylation of SFRP2, P16, DAPK1, HIC1, and MGMT and KRAS mutations in sporadic colorectal cancer. Cancer Genet Cytogenet. 2010;201:128–32.
Pan J, Chen J, Zhang B, Chen X, Huang B, Zhuang J, et al. Association between RASSF1A promoter methylation and prostate cancer: a systematic review and meta-analysis. PLoS One. 2013;8:e75283.
Hesson LB, Cooper WN, Latif F. The role of RASSF1A methylation in cancer. Dis Mark. 2007;23:73–87.
Chen SP, Wu CC, Huang SY, Kang JC, Chiu SC, Yang KL, Pang CY. beta-catenin and K-ras mutations and RASSF1A promoter methylation in Taiwanese colorectal cancer patients. Genet Test Mol Biomark. 2012;16:1277–81.
Lee KD, Pai MY, Hsu CC, Chen CC, Chen YL, Chu PY, et al. Targeted Casp8AP2 methylation increases drug resistance in mesenchymal stem cells and cancer cells. Biochem Biophys Res Commun. 2012;422:578–85.
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D. Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet. 2006;38:904–9.
Yeung KY, Ruzzo WL. Principal component analysis for clustering gene expression data. Bioinformatics. 2001;17:763–74.
Chuang JC, Yoo CB, Kwan JM, Li TW, Liang G, Yang AS, Jones PA. Comparison of biological effects of non-nucleoside DNA methylation inhibitors versus 5-aza-2′-deoxycytidine. Mol Cancer Ther. 2005;4:1515–20.
Leu YW, Rahmatpanah F, Shi H, Wei SH, Liu JC, Yan PS, Huang TH. Double RNA interference of DNMT3b and DNMT1 enhances DNA demethylation and gene reactivation. Cancer Res. 2003;63:6110–5.
Yan PS, Venkataramu C, Ibrahim A, Liu JC, Shen RZ, Diaz NM, et al. Mapping geographic zones of cancer risk with epigenetic biomarkers in normal breast tissue. Clin Cancer Res. 2006;12:6626–36.
Eisen MB, Spellman PT, Brown PO, Botstein D. Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA. 1998;95:14863–8.
Taylor JM. Kendall’s and Spearman’s correlation coefficients in the presence of a blocking variable. Biometrics. 1987;43:409–16.
Zurita M, Lara PC, del Moral R, Torres B, Linares-Fernández JL, Arrabal SR, et al. Hypermethylated 14-3-3-sigma and ESR1 gene promoters in serum as candidate biomarkers for the diagnosis and treatment efficacy of breast cancer metastasis. BMC Cancer. 2010;10:217.
Pedersen PA, Kristensen FB. [The Danish Medical Statistics and Danish practical research]. Ugeskr Laeger. 1990;152:828–9.
Klein JP. Survival analysis methods in cancer studies. Cancer Treat Res. 2002;113:37–57.
Malinge S, Chlon T, Dore LC, Ketterling RP, Tallman MS, Paietta E, et al. Development of acute megakaryoblastic leukemia in Down syndrome is associated with sequential epigenetic changes. Blood. 2013;122:e33–43.
Sandhu R, Roll JD, Rivenbark AG, Coleman WB. Dysregulation of the epigenome in human breast cancer: contributions of gene-specific DNA hypermethylation to breast cancer pathobiology and targeting the breast cancer methylome for improved therapy. Am J Pathol. 2015;185:282–92.
Zhang X, Wallace AD, Du P, Kibbe WA, Jafari N, Xie H, et al. DNA methylation alterations in response to pesticide exposure in vitro. Environ Mol Mutagen. 2012;53:542–9.
Acknowledgment
Y.W.L., S.H.H., P.Y.L., and Y.L.C. were supported by Changhua Christian Hospital, Taiwan (100-CCH-KMU-007). Y.W.L., S.H.H., and P.Y.C. were supported by NSC Taiwan (100-2320-B-194-001, 101-2320-B-194-001, and 100-2321-B-750-001). Y.W.L. and K.D.L. were supported by MoST, Taiwan (103-2314-B-182A-090). Y.W.L. and S.H.H. were supported by MoST (NSC 102-2320-B-194-003-MY3) and NHRI (NHRI-EX102-10259NI), Taiwan.
Disclosures
The authors declare that they have no commercial interests in the subject of this study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Hong-Chang Chen and Hsuan-Yuan Huang have contributed equally to this paper.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, HC., Huang, HY., Chen, YL. et al. Methylation of the Tumor Suppressor Genes HIC1 and RassF1A Clusters Independently From the Methylation of Polycomb Target Genes in Colon Cancer. Ann Surg Oncol 24, 578–585 (2017). https://doi.org/10.1245/s10434-015-5024-z
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-015-5024-z