Abstract
Background
Identification of molecular markers for sensitive detection of hepatocellular carcinoma (HCC) is required to achieve efficacious personalized therapy.
Methods
We focused here on SAM domain, SH3 domain, and nuclear localization signals 1 (SAMSN1) and investigated expression and methylation status of SAMSN1 in HCC cell lines and 144 pairs of surgical specimens.
Results
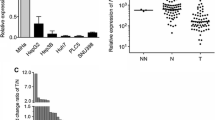
SAMSN1 was expressed at significantly lower levels in tumor tissue compared with the corresponding noncancerous tissues of patients with HCC. Analysis of HCC cell lines revealed that hypermethylation of the SAMSN1 promoter correlated with decreased expression of SAMSN1 mRNA. Furthermore, treating cells with a DNA-demethylating drug increased SAMSN1 transcription. The levels of SAMSN1 mRNA in noncancerous liver were not affected by background liver inflammation or fibrosis. Moreover, the levels of SAMSN1 mRNA in HCC tissues inversely correlated with tumor size and preoperative levels of proteins induced by vitamin K absence. The clinical significance of SAMSN1 was further indicated by the correlation between its decreased expression in patients with HCC and their shorter overall and recurrence-free survival as well as recurrence following initial resection. Moreover, multivariate analysis identified SAMSN1 as an independent prognostic factor of HCC progression. The expression pattern of SAMSN1 correlated significantly with that of SAMSN1 mRNA, making it possible to use PCR techniques to readily quantitate SAMSN1 expression in tumors.
Conclusions
Our findings indicate that inhibition of SAMSN1 transcription through DNA hypermethylation may influence the progression of HCC and thus represent a novel biomarker of the phenotype of HCC cells.



Similar content being viewed by others
References
GLOBOCAN 2012. Estimated Cancer incidence, mortality and prevalence worldwide in 2012, stomach cancer. Available at http://globocan.iarc.fr.
Flores A, Marrero JA. Emerging trends in hepatocellular carcinoma: focus on diagnosis and therapeutics. Clin Med Insights Oncol. 2014;8:71–6.
Vivarelli M, Montalti R, Risaliti A. Multimodal treatment of hepatocellular carcinoma on cirrhosis: an update. World J Gastroenterol. 2013;19:7316–26.
Kanda M, Nomoto S, Nishikawa Y, Sugimoto H, Kanazumi N, Takeda S, Nakao A. Correlations of the expression of vascular endothelial growth factor B and its isoforms in hepatocellular carcinoma with clinico-pathological parameters. J Surg Oncol. 2008;98:190–6.
Llovet JM, Burroughs A, Bruix J. Hepatocellular carcinoma. Lancet. 2003;362:1907–17.
Ramakrishna G, Rastogi A, Trehanpati N, Sen B, Khosla R, Sarin SK. From cirrhosis to hepatocellular carcinoma: new molecular insights on inflammation and cellular senescence. Liver Cancer. 2013;2:367–83.
Yang JD, Roberts LR. Hepatocellular carcinoma: A global view. Nat Rev Gastroenterol Hepatol. 2010;7:448–58.
Kanda M, Nomoto S, Oya H, et al. Downregulation of DENND2D by promoter hypermethylation is associated with early recurrence of hepatocellular carcinoma. Int J Oncol. 2014;44:44–52.
Giannelli G, Rani B, Dituri F, Cao Y, Palasciano G. Moving towards personalised therapy in patients with hepatocellular carcinoma: the role of the microenvironment. Gut. 2014;63:1668–76.
Galuppo R, Ramaiah D, Ponte OM, Gedaly R. Molecular therapies in hepatocellular carcinoma: what can we target? Dig Dis Sci. 2014;59:1688–97.
Llovet JM, Ricci S, Mazzaferro V, et al. Sorafenib in advanced hepatocellular carcinoma. N Engl J Med. 2008;359:378–90.
Cheng AL, Kang YK, Chen Z, et al. Efficacy and safety of sorafenib in patients in the Asia-Pacific region with advanced hepatocellular carcinoma: a phase III randomised, double-blind, placebo-controlled trial. Lancet Oncol. 2009;10:25–34.
Bruix J, Gores GJ, Mazzaferro V. Hepatocellular carcinoma: clinical frontiers and perspectives. Gut. 2014;63:844–55.
Zhao YJ, Ju Q, Li GC. Tumor markers for hepatocellular carcinoma. Mol Clin Oncol. 2013;1:593–8.
Kanda M, Nomoto S, Okamura Y, et al. Detection of metallothionein 1G as a methylated tumor suppressor gene in human hepatocellular carcinoma using a novel method of double combination array analysis. Int J Oncol. 2009;35:477–83.
Minguez B, Lachenmayer A. Diagnostic and prognostic molecular markers in hepatocellular carcinoma. Dis Markers. 2011;31:181–90.
Miki D, Ochi H, Hayes CN, Aikata H, Chayama K. Hepatocellular carcinoma: towards personalized medicine. Cancer Sci. 2012;103:846–50.
Overdevest JB, Theodorescu D, Lee JK. Utilizing the molecular gateway: the path to personalized cancer management. Clin Chem. 2009;55:684–97.
Zhu YX, Benn S, Li ZH, et al. The SH3-SAM adaptor HACS1 is up-regulated in B cell activation signaling cascades. J Exp Med. 2004;200:737–47.
Noll JE, Hewett DR, Williams SA, Vandyke K, Kok C, To LB, Zannettino AC. SAMSN1 is a tumor suppressor gene in multiple myeloma. Neoplasia. 2014;16:572–85.
Wang D, Stewart AK, Zhuang L, et al. Enhanced adaptive immunity in mice lacking the immunoinhibitory adaptor Hacs1. FASEB J. 2010;24:947–56.
Yan Y, Zhang L, Xu T, et al. SAMSN1 is highly expressed and associated with a poor survival in glioblastoma multiforme. PLoS One. 2013;8:e81905.
Yamada H, Yanagisawa K, Tokumaru S, et al. Detailed characterization of a homozygously deleted region corresponding to a candidate tumor suppressor locus at 21q11-21 in human lung cancer. Genes Chromosomes Cancer. 2008;47:810–8.
Kanda M, Nomoto S, Okamura Y, et al. Promoter hypermethylation of fibulin 1 gene is associated with tumor progression in hepatocellular carcinoma. Mol Carcinog. 2011;50:571–9.
Nomoto S, Kanda M, Okamura Y, et al. Epidermal growth factor-containing fibulin-like extracellular matrix protein 1, EFEMP1, a novel tumor-suppressor gene detected in hepatocellular carcinoma using double combination array analysis. Ann Surg Oncol. 2010;17:923–32.
Kanda M, Sugimoto H, Nomoto S, et al. Bcell translocation gene 1 serves as a novel prognostic indicator of hepatocellular carcinoma. Int J Oncol. 2015;46:641–8.
Takami H, Kanda M, Oya H, et al. Evaluation of MAGE-D4 expression in hepatocellular carcinoma in Japanese patients. J Surg Oncol. 2013;108:557–62.
Shimizu D, Kanda M, Nomoto S, et al. Identification of intragenic methylation in the TUSC1 gene as a novel prognostic marker of hepatocellular carcinoma. Oncol Rep. 2014;31:1305–13.
Kanda M, Shimizu D, Nomoto S, et al. Clinical significance of expression and epigenetic profiling of TUSC1 in gastric cancer. J Surg Oncol. 2014;110:136–44.
Oya H, Kanda M, Sugimoto H, et al. Dihydropyrimidinase-like 3 is a putative hepatocellular carcinoma tumor suppressor. J Gastroenterol. 2014. doi:10.1007/s00535-014-0993-4.
Kanda M, Nomoto S, Oya H, et al. Decreased expression of prenyl diphosphate synthase subunit 2 correlates with reduced survival of patients with gastric cancer. J Exp Clin Cancer Res. 2014;33:88.
Hibino S, Kanda M, Oya H, et al. Reduced expression of DENND2D through promoter hypermethylation is an adverse prognostic factor in squamous cell carcinoma of the esophagus. Oncol Rep. 2014;31:693–700.
Oya H, Kanda M, Takami H, et al. Overexpression of melanoma-associated antigen D4 is an independent prognostic factor in squamous cell carcinoma of the esophagus. Dis Esophagus. 2015;28:188–95.
Kanda M, Nomoto S, Oya H, et al. Dihydropyrimidinase-like 3 facilitates malignant behavior of gastric cancer. J Exp Clin Cancer Res. 2014;33:66.
Kanda M, Shimizu D, Nomoto S, et al. Prognostic impact of expression and methylation status of DENN/MADD domain-containing protein 2D in gastric cancer. Gastric Cancer. 2014. doi:10.1007/s10120-014-0372-0.
Claudio JO, Zhu YX, Benn SJ, et al. HACS1 encodes a novel SH3-SAM adaptor protein differentially expressed in normal and malignant hematopoietic cells. Oncogene. 2001;20:5373–7.
Lang PA, Recher M, Haussinger D, Lang KS. Genes determining the course of virus persistence in the liver: lessons from murine infection with lymphocytic choriomeningitis virus. Cell Physiol Biochem. 2010;26:263–72.
Stagg J, Galipeau J. Mechanisms of immune modulation by mesenchymal stromal cells and clinical translation. Curr Mol Med. 2013;13:856–67.
Vainer GW, Pikarsky E, Ben-Neriah Y. Contradictory functions of NF-kappaB in liver physiology and cancer. Cancer Lett. 2008;267:182–8.
Tokunaga F, Iwai K. Linear ubiquitination: a novel NF-kappaB regulatory mechanism for inflammatory and immune responses by the LUBAC ubiquitin ligase complex. Endocr J. 2012;59:641–52.
Ringelhan M, Reisinger F, Yuan D, Weber A, Heikenwalder M. Modeling human liver cancer heterogeneity: virally induced transgenic models and mouse genetic models of chronic liver inflammation. Curr Protoc Pharmacol. 2014;67:14.31.11–17.
El-Serag HB, Rudolph KL. Hepatocellular carcinoma: epidemiology and molecular carcinogenesis. Gastroenterology. 2007;132:2557–76.
Herath NI, Leggett BA, MacDonald GA. Review of genetic and epigenetic alterations in hepatocarcinogenesis. J Gastroenterol Hepatol. 2006;21:15–21.
Author information
Authors and Affiliations
Corresponding author
Additional information
Satoshi Sueoka and Mitsuro Kanda have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10434_2015_4524_MOESM1_ESM.tif
Supplementary material 1 RT-PCR analysis of the expression SAMSN1 expression in clinical specimens. (a) There were no significant differences in SAMSN1 mRNA levels among noncancerous tissues categorized by background uninvolved liver status. (b) SAMSN1 mRNA was expressed at lower levels in HCC tissues compared with those of the corresponding noncancerous tissues. (EPS 662 kb)
10434_2015_4524_MOESM2_ESM.tif
Supplementary material 2 (a) Correlation of SAMSN1 mRNA expression levels in HCC tissues with tumor size and (b) PIVKA II levels. (EPS 545 kb)
10434_2015_4524_MOESM3_ESM.tif
Supplementary material 3 SAMSN1 mRNA levels in HCC tissues categorized by the recurrence pattern at the initial (a) and second (b) recurrence. (EPS 17225 kb)
Rights and permissions
About this article
Cite this article
Sueoka, S., Kanda, M., Sugimoto, H. et al. Suppression of SAMSN1 Expression is Associated with the Malignant Phenotype of Hepatocellular Carcinoma. Ann Surg Oncol 22 (Suppl 3), 1453–1460 (2015). https://doi.org/10.1245/s10434-015-4524-1
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-015-4524-1