Abstract
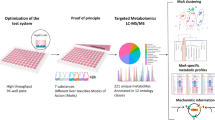
Selective chemical inhibitors are critical for reaction phenotyping to identify drug-metabolizing enzymes that are involved in the elimination of drug candidates. Although relatively selective inhibitors are available for the major cytochrome P450 enzymes (CYP), they are quite limited for the less common CYPs and non-CYPs. To address this gap, we developed a multiplexed high throughput screening (HTS) assay using 20 substrate reactions of multiple enzymes to simultaneously monitor the inhibition of enzymes in a 384-well format. Four 384-well assay plates can be run at the same time to maximize throughput. This is the first multiplexed HTS assay for drug-metabolizing enzymes reported. The HTS assay is technologically enabled with state-of-the-art robotic systems and highly sensitive modern LC-MS/MS instrumentation. Virtual screening is utilized to identify inhibitors for HTS based on known inhibitors and enzyme structures. Screening of ~4600 compounds generated many hits for many drug-metabolizing enzymes including the two time-dependent and selective aldehyde oxidase inhibitors, erlotinib and dibenzothiophene. The hit rate is much higher than that for the traditional HTS for biological targets due to the promiscuous nature of the drug-metabolizing enzymes and the biased compound selection process. Future efforts will focus on using this method to identify selective inhibitors for enzymes that do not currently have quality hits and thoroughly characterizing the newly identified selective inhibitors from our screen. We encourage colleagues from other organizations to explore their proprietary libraries using a similar approach to identify better inhibitors that can be used across the industry.
Graphical abstract









Similar content being viewed by others
Abbreviations
- ABT:
-
1-Aminobenzotriazole
- ADME:
-
Absorption, distribution, metabolism, and excretion
- AO:
-
Aldehyde oxidase
- CBR:
-
Carbonyl reductase
- CE:
-
Collision energy
- CES:
-
Carboxylesterase
- CO2 :
-
Carbon dioxide
- CXP:
-
Collision cell exit potential
- CYP:
-
Cytochrome P450
- DDI:
-
Drug-drug interaction
- DMSO:
-
Dimethyl sulfoxide
- DP:
-
Declustering potential
- EPHX:
-
Epoxide hydrolase
- f m :
-
Fraction metabolized
- FMO:
-
Flavin-containing monooxygenase
- HEPES:
-
4-(2-Hydroxyethyl)-1-piperazineethanesulfonic acid
- HHEP:
-
Human hepatocytes
- HLM:
-
Human liver microsomes
- HPLC:
-
High-performance liquid chromatography
- HTS:
-
High throughput screening
- IC50 :
-
Half-maximal inhibitory concentration
- IVIVE:
-
In vitro-to-in vivo extrapolation
- K I :
-
Inactivation rate constant
- k inact :
-
The maximum rate of enzyme inactivation
- K i,u :
-
Unbound inhibition constant
- K m :
-
Michaelis constant
- LC-MS/MS:
-
Liquid chromatography with tandem mass spectrometry
- MAO:
-
Monoamine oxidase
- MRM:
-
Multiple-reaction monitoring
- 4-MU:
-
4-Methylumbelliferone
- M/Z:
-
Mass-to-charge ratio
- Na2CO3 :
-
Sodium carbonate
- NADPH:
-
Nicotinamide adenine dinucleotide phosphate, reduced form
- NAT:
-
N-acetyltransferase
- PK:
-
Pharmacokinetics
- Q1:
-
First quadrupole mass filter
- Q3:
-
Second quadrupole mass filter
- TDI:
-
Time-dependent inhibition
- UGT:
-
Uridine 5′-diphospho-glucuronosyltransferase
- WEM:
-
Williams E medium
References
Sun D, Gao W, Hu H, Zhou S. Why 90% of clinical drug development fails and how to improve it? Acta Pharm Sin B. 2022;12(7):3049–62. https://doi.org/10.1016/j.apsb.2022.02.002.
Dickson M, Gagnon JP. Key factors in the rising cost of new drug discovery and development. Nat Rev Drug Discov. 2004;3(5):417–29. https://doi.org/10.1038/nrd1382.
Hughes JP, Rees S, Kalindjian SB, Philpott KL. Principles of early drug discovery. Br J Pharmacol. 2011;162(6):1239–49. https://doi.org/10.1111/j.1476-5381.2010.01127.x.
Doytchinova I. Drug design-Past, present, future. Molecules (Basel, Switzerland). 2022;27(5). doi: https://doi.org/10.3390/molecules27051496.
Waring MJ, Arrowsmith J, Leach AR, Leeson PD, Mandrell S, Owen RM, et al. An analysis of the attrition of drug candidates from four major pharmaceutical companies. Nat Rev Drug Discov. 2015;14(7):475–86. https://doi.org/10.1038/nrd4609.
Bocci G, Carosati E, Vayer P, Arrault A, Lozano S, Cruciani G. ADME-Space: A new tool for medicinal chemists to explore ADME properties. Sci Rep. 2017;7(1):6359. https://doi.org/10.1038/s41598-017-06692-0.
Di L. Reaction phenotyping to assess victim drug-drug interaction risks. Expert Opin Drug Discov. 2017;12(11):1105–15. https://doi.org/10.1080/17460441.2017.1367280.
Zientek MA, Youdim K. Reaction phenotyping: Advances in the experimental strategies used to characterize the contribution of drug-metabolizing enzymes. Drug Metab Dispos. 2015;43(1):163–81. https://doi.org/10.1124/dmd.114.058750.
Bohnert T, Patel A, Templeton I, Chen Y, Lu C, Lai G, et al. Evaluation of a new molecular entity as a victim of metabolic drug-drug interactions-An industry perspective. Drug Metab Dispos. 2016;44(8):1399–423. https://doi.org/10.1124/dmd.115.069096.
Strelevitz TJ, Orozco CC, Obach RS. Hydralazine as a selective probe inactivator of aldehyde oxidase in human hepatocytes: Estimation of the contribution of aldehyde oxidase to metabolic clearance. Drug Metab Dispos. 2012;40(7):1441–8. https://doi.org/10.1124/dmd.112.045195.
Yang X, Johnson N, Di L. Evaluation of cytochrome P450 selectivity for hydralazine as an aldehyde oxidase inhibitor for reaction phenotyping. J Pharm Sci. 2019;108(4):1627–30. https://doi.org/10.1016/j.xphs.2018.11.007.
Yang X, Atkinson K, Di L. Novel cytochrome P450 reaction phenotyping for low-clearance compounds using the hepatocyte relay method. Drug Metab Dispos. 2016;44(3):460–5.
Tess D, Chang GC, Keefer C, Carlo A, Jones R, Di L. In vitro-in vivo extrapolation and scaling factors for clearance of human and preclinical species with liver microsomes and hepatocytes. AAPS J. 2023;25(3):40. https://doi.org/10.1208/s12248-023-00800-x.
Ramsden D, Smith D, Arenas R, Frederick K, Cerny MA. Identification and characterization of a selective human carbonyl reductase 1 substrate. Drug Metab Dispos. 2018;46(10):1434–40. https://doi.org/10.1124/dmd.118.082487.
Lee CA, Jones JP 3rd, Katayama J, Kaspera R, Jiang Y, Freiwald S, et al. Identifying a selective substrate and inhibitor pair for the evaluation of CYP2J2 activity. Drug Metab Dispos. 2012;40(5):943–51. https://doi.org/10.1124/dmd.111.043505.
Li X, Jeso V, Heyward S, Walker GS, Sharma R, Micalizio GC, Cameron MD. Characterization of T-5 N-oxide formation as the first highly selective measure of CYP3A5 activity. Drug Metab Dispos. 2014;42(3):334–42. https://doi.org/10.1124/dmd.113.054726.
Kung P-P, Bingham P, Brooun A, Collins M, Deng Y-L, Dinh D, et al. Optimization of orally bioavailable enhancer of zeste homolog 2 (EZH2) inhibitors using ligand and property-based design strategies: Identification of development candidate (R)-5,8-dichloro-7-(methoxy (oxetan-3-yl)methyl)-2-((4-methoxy-6-methyl-2-oxo-1,2-dihydropyridin-3-yl)methyl)-3,4-dihydroisoquinolin-1(2H)-one (PF-06821497). J Med Chem. 2018;61(3):650–65. https://doi.org/10.1021/acs.jmedchem.7b01375.
Ghosal A. Chapter 8 - Evaluation of the clearance mechanism of non-CYP-mediated drug metabolism and DDI as a victim drug. In: Ma S, Chowdhury SK, editors. Identification and Quantification of Drugs, Metabolites, Drug Metabolizing Enzymes, and Transporters. Second ed. Amsterdam: Elsevier; 2020. p. 237–71.
Zientek M, Youdim K. Simultaneous determination of multiple CYP inhibition constants using a cocktail-probe approach. Methods in molecular biology (Clifton, NJ). 2013;987:11–23. https://doi.org/10.1007/978-1-62703-321-3_2.
Zientek M, Miller H, Smith D, Dunklee MB, Heinle L, Thurston A, et al. Development of an in vitro drug-drug interaction assay to simultaneously monitor five cytochrome P450 isoforms and performance assessment using drug library compounds. J Pharmacol Toxicol Methods. 2008;58(3):206–14. https://doi.org/10.1016/j.vascn.2008.05.131.
Yang X, Ryu S, Di L. Dasotraline as a selective cytochrome 2B6 inhibitor for reaction phenotyping. Biopharm Drug Dispos. 2019;40(9):358–61. https://doi.org/10.1002/bdd.2207.
Derringer G, Suich R. Simultaneous optimization of several response variables. J Qual Technol. 1980;12(4):214–9. https://doi.org/10.1080/00224065.1980.11980968.
Graczyk PP. Gini Coefficient: A new way to express selectivity of kinase inhibitors against a family of kinases. J Med Chem 2007;50(23):5773-5779. doi: https://doi.org/10.1021/jm070562u.
Margaillan G, Rouleau M, Klein K, Fallon JK, Caron P, Villeneuve L, et al. Multiplexed targeted quantitative proteomics predicts hepatic glucuronidation potential. Drug Metab Dispos. 2015;43(9):1331–5. https://doi.org/10.1124/dmd.115.065391.
Dalvie D, Di L. Aldehyde oxidase and its role as a drug metabolizing enzyme. Pharmacol Ther. 2019;201:137–80. https://doi.org/10.1016/j.pharmthera.2019.05.011.
Tan WK, Tan ARY, Sivanandam P, Goh EJH, Yap ZP, Saburulla NF, et al. In vitro inhibition of human aldehyde oxidase activity by clinically relevant concentrations of gefitinib and erlotinib: Comparison with select metabolites, molecular docking analysis, and impact on hepatic metabolism of zaleplon and methotrexate. J Pharmacol Exp Ther. 2020;374(2):295–307. https://doi.org/10.1124/jpet.120.265249.
Chen S, Austin-Muttitt K, Zhang LH, Mullins JGL, Lau AJ. In vitro and in silico analyses of the inhibition of human aldehyde oxidase by bazedoxifene, lasofoxifene, and structural analogues. J Pharmacol Exp Ther. 2019;371(1):75–86. https://doi.org/10.1124/jpet.119.259267.
Tunek A, Svensson LA. Bambuterol, a carbamate ester prodrug of terbutaline, as inhibitor of cholinesterases in human blood. Drug Metab Dispos. 1988;16(5):759–64.
Pistolozzi M, Du H, Wei H, Tan W. Stereoselective inhibition of human butyrylcholinesterase by the enantiomers of bambuterol and their intermediates. Drug Metab Dispos. 2015;43(3):344–52. https://doi.org/10.1124/dmd.114.060251.
Acknowledgements
Authors greatly appreciate the help of Sophia M. Shi in editing the manuscript and many colleagues for their helpful discussion.
Author information
Authors and Affiliations
Contributions
Substantial contributions to the conception or design of the work; or the acquisition, analysis, or interpretation of data for the work: JL, DV, GB, AC, WB, SJ, LT, JY, YC, GC, MT, and LD. Drafting the work or revising it critically for important intellectual content: JL, DV, WB, SJ, LT, and LD. Final approval of the version to be published: JL, DV, GB, AC, WB, SJ, LT, JY, YC, GC, MT, and LD. Agreement to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved: JL, DV, GB, AC, WB, SJ, LT, JY, YC, GC, MT, and LD.
Corresponding author
Ethics declarations
Conflict of Interest
Authors are employees of Pfizer Inc., New York, NY, USA, and may hold Pfizer stocks.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, J., Vernikovskaya, D., Bora, G. et al. Novel Multiplexed High Throughput Screening of Selective Inhibitors for Drug-Metabolizing Enzymes Using Human Hepatocytes. AAPS J 26, 36 (2024). https://doi.org/10.1208/s12248-024-00908-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12248-024-00908-8