Abstract
The two one-sided t-tests (TOST) procedure has been used to evaluate average bioequivalence (BE). As a regulatory standard, it is crucial that TOST distinguish BE from not-BE (NBE) when BE data are not lognormal. TOST was compared with a Bayesian procedure (BEST by Kruschke) in simulated datasets of test/reference ratios (T/R) which were BE and NBE, wherein (1) log(T/R) or T-R were normally distributed, (2) sample sizes ranged 10–50, and (3) extreme log(T/R) or T-R values were randomly included in datasets. The 90% “credible interval” (CrI) from BEST is a Bayesian alternative of the 90% confidence interval (CI) of TOST and it can be derived from a posterior distribution that is more reflective of the observed mean log(T/R) distribution that often deviates from normality. In the absence of extreme T/R values, both methods agreed BE when observed T/R were lognormal. BEST more accurately concluded BE or NBE, while requiring fewer subjects, when observed log(T/R) or T-R were normal in the presence of extreme values. Overall, TOST and BEST perform comparably on lognormal T/R, while BEST is more accurate, requiring fewer subjects when datasets are normal for T-R or contain extreme values. Of note, the normally distributed datasets only rarely contain extreme values. Our results imply that when BEST and TOST yield different BE assessment results from bioequivalent products, TOST may disadvantage applicants when T/R are not lognormal and/or include extreme T/R values. Application of BEST can address the situation when T/R are not lognormal or include extreme data values. Application of BEST to BE data can be considered a useful alternative to TOST for evaluation of BE and for efficient development of BE formulations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Demonstrating average bioequivalence (BE) in small, two-treatment two-way crossover bioavailability (BA) studies has long relied upon the two one-sided t-tests (TOSTs) (1), applied to log-transformed geometric mean ratios (GMR) of test (T) (the test product is usually a generic version of the reference product) and reference (R) drug area-under-drug concentration-time curve (AUC) and maximum drug concentration (Cmax) values. TOST can generate 90% confidence intervals (CIs) of the GMRs from BE datasets and then BE is concluded if the confidence intervals (after exponentiation) of GMR for AUC and Cmax fall within BE acceptance regions, 80 to 125%.
Peck and Campbell (2) raised concerns of TOST for BE assessment, which were related to the following factors for valid inference of BE using TOST: (a) normality of mean log-transformed T/R of AUC and Cmax, (b) insufficiency of sample sizes to rely on the central limit theorem, and (c) robustness to extreme values of T/R. Robustness of TOST to deviations from normality of log-transformed T/R or presence of extreme T/R values has not been thoroughly investigated, although several reports have shown concern for the appropriateness of TOST in actual practice given the above-listed factors (3,4,5,6). Fitting log(T/R) data with a normal distribution does not easily accommodate data with heavier tails. The t-distribution is well-suited for such deviations from normality and can flexibly accommodate normally distributed log(T/R) data.
While other Bayesian approaches have been proposed for bioequivalence (7,8,9,10) and biosimilarity (11), a Bayesian procedure for BE, adapted from Kruschke—BEST (12)—is employed herein to address the above concerns. BEST enables a Bayesian BE test using the 90% “credible interval” (CrI) (13, 14) of the mean log(T/R), under the assumption of an underlying t-distribution for log(T/R). Non-influential prior distributions are employed in Bayesian estimation of t-distribution parameters (mean, standard distribution, and shape (degrees of freedom)) that can flexibly represent either normal or t-distributions with heavier tails. BEST assumes that log(T/R) follows a t-distribution with minimally informative priors, whereas TOST assumes log(T/R) is normally distributed and uses the t-test for inference about the mean. Here, the 90% CrI using the highest density interval (HDI) is a Bayesian alternative of the 90% CI of TOST, which is similar to the 90% most likely values for the true average log(T/R). Accordingly, the BE limits for the BEST are set as 80 to 125% (15).
To investigate the validity of inference of BE or not-BE (NBE) by TOST and BEST with respect to the above-listed concerns of TOST, we compared TOST with BEST in simulated BE datasets mimicking 200 scenarios, as described in the “Models for BE Simulation” section in the “Methods” section.
Methods
To compare performances of TOST and BEST in BE assessments, simulations were conducted to represent typical BE scenarios comprising lognormal T/R and normal T-R distributions, varying mean, different sample sizes, and presence of extreme outliers.
To demonstrate the features of BE analysis via BEST, two real cases of anonymized Abbreviated New Drug Applications (ANDA) BE datasets, with and without approximately normally distributed log(T/R), were analyzed to generate key relevant diagnostic plots and posterior distributions of the log-transformed mean T/R ratios.
Models for BE Simulation
BA datasets were simulated according to the FDA-recommended linear mixed-effects model for BE analysis (15) of pharmacokinetic (PK) parameters AUC or Cmax or their logs (Yijk):
where μ is the parameter average; Pj is the period effect, j=1, 2; F(j, k) is the formulation effect for period j and sequence k=1,2; Sik is the random effect for subject i=1, …, nk in sequence k; and eijk is the residual error. In this model, fixed effects are the overall mean μ, P, and F; random effects S and e are independent normally distributed with mean zero and between and within subject variances (i.e., \({\sigma}_b^2\) and \({\sigma}_w^2\), respectively). Period and sequence carryover effects are set to zero in the simulations. The test and reference drug are assumed to have equal variances, but this can be easily generalized to unequal ones.
AUC and Cmax datasets representing two-treatment crossover studies were simulated to be either lognormally for T/R or normally distributed for T-R according to Eq. (1). Datasets were randomly generated, employing prespecified mean values, within-subject variability (\({\sigma}_w^2\Big)\) (20% CV (16, 17)), and sample sizes 10–50, reflecting typical BE studies. The overall mean (μ) of the simulated data was set to an arbitrary value (e.g., 100 ng/mL for Cmax and 1000 ng·h/mL for AUC) and between-subject variability (\({\sigma}_b^2\)) set equal to the within-subject variability. The selected values of μ and \({\sigma}_b^2\) do not affect the BE analysis in these crossover study simulations because both the fixed mean μ and random between-subject effect cancel out with the difference calculated from a crossover study.
Given the pre-specified and derived parameters, following Eq. (1), ( Yi11, Yi21) for i=1,…, n1 and ( Yi22, Yi12 ) for i=n1+1,…, n1+n2 were generated from the FDA-recommended linear mixed-effects model with mean (μ + FT, μ + FR) for formulation effects for the test FT and the reference FR = −FT and variance \(\upomega =\left[\begin{array}{cc}{\sigma}_w^2+{\sigma}_b^2& {\sigma}_b^2\\ {}{\sigma}_b^2& {\sigma}_w^2+{\sigma}_b^2\end{array}\right]\) (assuming no period effect).
For subject i, YTi denotes the PK parameter for the test product and YRi the reference product, with estimated least square means \({\overline{\mathit{\mathsf{Y}}}}_{\mathit{\mathsf{T}}}\) and \({\overline{\mathit{\mathsf{Y}}}}_{\mathit{\mathsf{R}}}.\)
To generate BE data that follow a lognormal distribution, normal data were simulated and the PK parameters were transformed back to the original unlogged scale by exponentiation.
To prevent occurrence of negative values when simulating normally distributed data (<1% incidence conditioned on \({\sigma}_b^2\) of 20%), negative values were replaced with a small positive value of 5% of the overall mean.
BE Analysis Methods: TOST vs BEST
In TOST (1), the equivalence test comprises two one-sided alpha = 0.05 tests, yielding two p-values. The null hypotheses are rejected (i.e., concluding bioequivalence) if both p-values are less than 0.05.
BEST employs non-influential priors for t-distribution parameters of AUC or Cmax distributions and the Markov Chain Monte Carlo procedure to compute posterior distributions (18) for these parameters. Analogous to TOST, BEST applied to log-transformed T/R generates the 90% CrI from the log-transformed GMR, consistent with the average BE approach. When lower and upper bounds of the CrI fall within the log-transformed acceptance region such as [log(0.80), log(1.25)], the BEST procedure supports BE.
For the Bayesian inference, the prior parameter distributions for the t-distribution are set to be minimally informative with insignificant influence on the estimates of the parameters. Specifically, the mean is assumed to have a prior normal distribution with mean and standard deviation that is 1000 times that of the sampled data; the prior on the standard deviation parameter is assumed to have a uniform distribution with a low limit of 1/1000 and a high limit of 1000 times of the standard deviation of the sampled data; the prior on the shape (degrees of freedom) parameter (i.e., as a measure of height of tails in a t distribution) is assumed to have a shifted exponential distribution as explained in Kruschke (12).
Evaluating BE on the original unlogged scale, the (non-Bayesian) arithmetic mean ratio (AMR) procedure assumes the T-R data are normally distributed and generates the 90% CI of the T-R difference, dividing this CI interval by the estimated mean of BA of the reference formulation, and adding 1.0 (19). Using the CI of the difference between the test and reference products allows for cancellation of the effect from the same subject during crossover. The same acceptance region of 80–125% is used for AMR in this report. Because the FDA recommends that the TOST analysis be applied to log-transformed data, AMR-based BE assessments have not been submitted for BE evaluation in the past.
BEST can also provide a Bayesian BE test from 90% CrI of AMR, assuming the T-R differences have a t-distribution. The BEST procedure for AMR (BEST AMR) is described as the following steps:
-
1)
Zi = (YTi − YRi) / \({\overline{Y}}_R\).
-
2)
BEST AMR generated the 90% CrI of mean Zi’s per Bayesian posterior distribution.
-
3)
The calculated CrI (lower, upper) is then incremented by 1, i.e., CrIAMR = [CrIZ + 1].
If this CrI of AMR falls within the acceptance region (such as 80–125%), BEST AMR supports BE between T and R.
Simulations were also reported for another BEST AMR procedure called BEST AMRmu, where the observed average \({\overline{Y}}_R\) in (1) is replaced by the true reference mean μ + FR = μR, which is typically not known in practice.
Accuracy and Passing Rates of TOST and BEST on Simulated Datasets
Lognormally and normally distributed values were simulated as follows:
-
1.
Preset mean T/R values M ranged 0.80 to 1.25, where M is the exponentiated mean log-ratio for lognormal data and the ratio of the means for normal data.
-
2.
Sample sizes for each of T and R: 10, 20, 30, 40, and 50
-
3.
Each scenario was simulated 1000 times.
-
4.
TOST and BEST passing rates, which were the rates that meet the testing definition of BE, were calculated for lognormal and normal data, where mean values M=0.9, 1.0 and 1.11 are considered BE and 0.80 and 1.25 NBE.
Generation of Extreme Values
Extreme values occur in BE datasets that can significantly alter the data distribution, leading to deviations from lognormality or normality that challenge the validity of inference of BE. To evaluate the robustness of the TOST and the BEST in the presence of extreme values, BE datasets with extreme values were simulated in lognormal and normal data. Extreme values were randomly generated by multiplying the log(T)–log(R) and T-R differences by a factor of 10 with 5% probability. The TOST and BEST procedures were applied to each 1000 replicates, calculating passing rates for each BE scenario. While a greater number of simulation iterations than 1000 could be executed to derive highly precise BE passing rates to inform type I or type II errors, evaluations with 20,000 iterations on a few selected testing scenarios resulted in minimal changes in the simulation-derived BE passing rates. No differentiation of acceptance or rejection of BE with respect to T or R origin of extreme values in the reference or test datasets was reported, as might be the case for a regulatory agency.
Results
Features of BE Analysis via BEST
The BEST procedure generates diagnostic histograms that facilitate the evaluation of assumptions that are critical for valid statistical inference, which we illustrate here for two anonymized ANDA BE datasets (Fig. 1)—case 1 (panels A–C, approximately normally distributed log(T/R)) and case 2 (panels D–F, not normally distributed log(T/R), containing at least one extreme value). These two anonymous ANDA BE datasets comprised AUCinf data from two 2-way crossover PK studies with 16 (8 for each sequence) and 28 (14 for each sequence) subjects, respectively. Panels A and B (case 1) and D and E (case 2) depict posterior distributions of the mean and shape (degrees of freedom) parameters for the log-transformed T/R respectively for each case. Panels C and F, are posterior predictive checks or visual of goodness-of-fit plots, and show the estimated posterior predictive distribution for log(T/R) overlying simple histograms of the real data. Comparing the posterior distributions of the log-transformed mean T/R for cases 1 and 2 in panels A and D in Fig. 1, panel D shows wider spread than a normal distribution and appears to be skewed to the right. These plots illustrate a unique feature of using BEST, which reveal the underlying true distributions of the observed T/R ratio—a critical assumption for the TOST approach.
Posterior distributions of estimated parameters (i.e., mean (a and d), and normality (shape (degrees of freedom) on the log10 scale) (b and e) of log ratios) and the posterior predictive check (c, f) for cases 1 (a–c) and 2 (d–f). Bayesian highest density credible intervals (HDI) are labeled for the mean and shape (degrees of freedom) parameters
Panels B and E show posterior distributions of the estimated shape (degrees of freedom) parameter for cases 1 and 2, in which values on the horizontal axis are displayed on a logarithmic scale, base 10. Along with the goodness-of-fit plots in panels C and F in these cases, the values of the modes of 1.44 and 0.627 (log base 10) correspond to the exponentiated shape (degrees of freedom) parameter values of 27.54 and 4.24, indicating approximate lognormality for case 1, since the t-distribution is well-approximated by a normal distribution when the shape (degrees of freedom) parameter approaches 30 (case 1) and above. Case 2 is an example of a heavy-tailed data distribution with an extreme value for which lognormality was rejected by the Shapiro-Wilk test but the t-distribution accommodated the extreme value and fit the log(T/R) data well.
BEST vs TOST Performance Evaluation via Simulated Data
Figure 2 shows passing rate comparisons for simulated lognormal T/R distributions, with average mean values of NBE (0.8, 1.25) and BE (0.9, 1.0, 1.11) and sample sizes ranging 10–50. Means of 1.25 and 1.11 are reciprocals of 0.80 and 0.9. Regardless of NBE and BE status and sample size, TOST (blue) and BEST (red) exhibit nearly identical passing rate-sample size curves. While TOST is optimal for lognormal data, BEST sacrifices imperceptibly less power. The passing rates for TOST and BEST are well-controlled (type I error rate of ≤ 0.05) for NBE scenarios.
Since the AMR procedure and BEST AMR did not appear to differ in passing rates, only BEST AMR (and BEST AMRmu) are presented in Figs. 2 and 3. In lognormally distributed datasets (Fig. 2), the performances of BEST AMR (gray) and BEST AMRmu (green) are inferior for mean M=0.9 and slightly inferior at M=1.0 (and superior for 1.11) and the type I error rates appear to be controlled at ≤ 0.05 for NBE for all four procedures at M=0.80 and 1.25 except BEST AMR at 1.25. While Figs. 4 and 5 show that BEST yielded higher passing rates than TOST for BE products, BEST and TOST appear to have comparable type I error rates. Of note, we observed a slight increase in the passing rate at M=0.80 or 1.25 for BEST but 20,000 iterations confirm type I errors less than 0.05 for n<50. When T/R data are lognormally distributed, sample sizes to achieve 80% power to pass BE at M=0.9, 1.0 and 1.11 are estimated to be ~35, ~17, and ~ 37, respectively, for either TOST or BEST approaches.
For simulated normally distributed T-R, Fig. 3 shows passing rate comparisons for NBE (M=0.8, 1.25) and BE (M=0.9, 1.0, 1.11) with sample sizes ranging 10–50. BEST AMR and BEST AMRmu are superior to TOST and BEST for BE mean values M of 0.9, 1.0, and 1.11, and BEST is superior to TOST for the same M values. All four methods appear to correctly conclude NBE with a passing rate of ≤ 0.05 at 0.80 and 1.25, except BEST AMR at 1.25.
BEST vs TOST Performance on Datasets with Extreme Values
Figures 4 and 5 show performances when extreme observed log(T/R) and T-R values are randomly included in simulated normal datasets. BEST yields markedly higher passing rates and lower sample size requirements than TOST in lognormally distributed T/R values: ~ 45, 20, and 45 subjects for M (0.9, 1.0, 1.11) are required for 80% passing rate by BEST vs >> 50 subjects for TOST (Fig. 4), and somewhat similarly with normally distributed T/R (> 50, 22, 45 for BEST >> 50 for TOST). Type I error rates of BEST and TOST in NBE scenarios (0.8 and 1.25) are controlled at less than 5% for normal data and 6% or less for lognormal data.
Discussion
FDA encourages analysis of BE data only on the log scale and discourages testing for lognormality. FDA’s rationale (15) for this policy is that tests of lognormality in typical small samples have insufficient power to reject the hypothesis of lognormality, and failure to reject the hypothesis does not confirm that the data are approximately lognormal. However, because power is low for alternative distributions in small sample sizes (20), when (log)normality (i.e., lognormality or normality) is rejected in a small dataset, it can signal a potentially gross deviation. As noted above, an important benefit of the BEST approach is employment of a rich set of diagnostic tools that permit investigations of the distributions of BE datasets and model parameters that enable visual confirmation (or not) of lognormality of T/R distribution, and thus, validity of inference.
Inference using TOST relies on the assumption that log-transformed T/R follow a normal distribution, an assumption that is crucial in small sample sizes. Several publications have critiqued the use of a normal theory-based test such as TOST when the sample sizes are small and the data are not normal (3, 5, 9). While some may recommend sample sizes of over 30 for distributions with no extreme values and little skewness in order to provide assurance of approximate normality of the mean difference and the log-transformed GMR for the sample mean, the convergence rate of the central limit theorem is more complicated and a convergence bound based on the Berry-Esseen theorem depends on the variance, the sample size, and third absolute moment (21). In the presence of skewed distributions or very extreme values, estimated BE intervals may not be well approximated using normal theory-based procedures such as TOST, even when sample sizes are larger than 30. Beneficially, the posterior t-distribution parameter distributions and the posterior predictive distributions via the BEST procedure can be used to assess (log)normality.
BEST is an appealing alternative to TOST, by (a) accommodating a few extreme values or a heavy-tailed t-distribution and (b) providing the 90% credible interval for the mean difference, analogous to the 90% confidence interval of TOST. Importantly, Bayesian BEST procedures provide diagnostic posterior distributions for the parameters (mean, standard deviation, shape (degrees of freedom)) of the t-distribution, as well as the posterior distribution of any calculated quantities, such as the probability that the mean is in the BE acceptance region. Additionally, a value of 10 or less for the mode of posterior distribution of the shape (degrees of freedom) parameter can indicate that underlying data are not normally distributed but heavier tailed. In real BE datasets with measurement error, small sample size, and extreme values, the means of log-transformed T/R may not be well approximated by normal distributions, and hence the BEST method can be more resilient to violations of normality assumptions than TOST.
A crucial issue concerns which distribution adequately describes the T/R data for valid inference. Our simulations show that when the T/R are lognormal, TOST (absent extreme values) and BEST perform equally well. When sample sizes in BE studies are small or contain extreme values, normal-based inference such as TOST may be questionable. One alternative when data do not appear lognormal or normal would be to search for a different distribution. For instance, a Box-Cox transformation could be considered to transform data to be normally distributed (22). Other alternatives would be non-parametric methods based on the signed rank statistic (23, 24) or the bootstrap (25).
A criticism of using BEST is its reliance on broad prior distributions using current data; in particular, the prior for the mean is centered at the pooled mean from the current data and prior for the standard deviation is 1000 times its pooled standard deviation. Although not absolutely non-informative, the amount of prior information can be estimated from the effective sample size (26), which is approximately 1/1,000,000 of one observation or, to say it another way, a single observation in a bioequivalence study is five or six orders of magnitude more informative than this prior. Depending on the purpose of use, Bayesian procedures such as BEST can employ variably influential prior information that generate useful CrIs: (i) BEST with aforementioned non-influential priors depending slightly on the actual data for regulatory purpose, and (ii) BEST with more informative priors from pilot studies or the reference drug for formulation development. Since the BEST approach generally requires fewer subjects, serial BE studies in support of formulation optimization for BE may be replaced by fewer, smaller studies, especially if priors are made more informative from observations in the previous pilot studies. For example, in formulation development, extant information on the BE parameter distribution of the formulation from pilot study(ies) could be used to modify the prior distribution for BEST as described in Kruschke (12), leading to efficient formulation selection with smaller sample sizes in new drug development.
Schuirmann et al critiqued the Bayesian estimation approach using BEST (27), questioning the value of estimating the entire GMR distribution. A case can be made for estimating the entire log(T/R) distribution to assess lognormality and the effect of extreme values before making the BE determination. Challenging FDA’s admonition against testing for normality, the diagnostic evaluations by BEST of the entire GMR distributions of two real ANDA cases (Fig. 1) show lognormality in one case and not in the other, calling into question the validity of relying upon TOST in the later case. FDA guidance discourages robust inference procedures for BE assessment despite the potential biasing effect of extreme values on inference when the normality assumption of TOST is violated. In this case, the BEST procedure enables valid inference without violating the assumption that BE GMR datasets are well characterized by the t-distribution. Schuirmann et al (27) also expressed concern about how BEST sets minimally informative priors from the data; the amount of such influence is minimal as explained above.
When the underlying data are normally distributed, BEST is superior to TOST, and both are inferior to BEST AMR. However, BEST AMR does not appear to adequately control the passing rate at M=1.25, although BEST AMRmu does. Interestingly, this does not seem to be an issue at M=0.80. The behavior of BEST AMR in Fig. 2 for M=1.11 showing superiority to TOST and BEST may be in part due to its failure to control the type I error at M=1.25, a failure due to the variability introduced by dividing by the observed reference mean rather than its theoretical mean since this increases the variability of the estimated ratio. For normal data, BEST AMR using an acceptance region of [0.80, 1.22] is superior to BEST and TOST and maintains the type I error control (simulations not shown); more research is needed for the situation in which the underlying data are normally distributed.
Robustness to Extreme T/R Values
Extreme values (“outlier data”) are values that are significantly discordant with data for that subject and/or deviate from the typical trajectory of concentration-time data of the subject in a BE study (28,29,30,31). In crossover BE studies, extreme T/R values often can be observed in one or a few subjects. Extreme values can indicate either product failure, measurement errors, inherently high variability, or subject-by-formulation interactions. Inspection of scatterplots of log(T) versus log(R) can facilitate identification of the source of the extreme values as from either test or reference product. From a regulatory perspective, extreme values may only be removed from the BE statistical analysis if there is real-time documentation demonstrating a protocol violation during the clinical and/or analytical/experimental phase of the BE study (32, 33).
Our simulations demonstrate that BEST procedures yield higher BE passing rates than TOST if extreme values occur in a log-transformed data set. An extreme value can cause a larger estimated standard deviation and hence wider intervals in logged and non-logged data but both BEST procedures dampen the effect of this extreme value.
Arguably, extreme values in the reference product as opposed to the test product on the untransformed or log-transformed scale should not disadvantage the abbreviated new drug applicant. Rather than deleting such extreme reference product values, application of BEST on the log scale can decrease their influence. In typical small BE datasets (n<50), BEST robustly characterizes log(T/R) from a t-distribution. Applicants submitting non-normally distributed BE datasets and/or datasets that include one or more extreme log(T/R) values may be disadvantaged by failing BE via TOST due to forced normality, while being BE via BEST.
TOST or BEST—Which to Employ for BE?
FDA has stated that TOST is a “size-alpha test, a valid statistical test for average BE,” and that “empirical experience supports the view that normal-theory inference methods will be valid, even with the small sample sizes of typical BE studies” (1). TOST is known to be a size-alpha test if the data are lognormal or the central limit theorem provides a reasonable normal approximation of the log(GMR) for the actual sample size. Notwithstanding the fact that the long-term frequency underpinning TOST for type I error control is not assured in small BE trials, type I error control and validity of BE are not guaranteed if the mean of log ratios is not approximately normally distributed. This could occur if the underlying data are not lognormal or the sample size is inadequate to guarantee that the central limit theorem will provide a reasonable normal approximation. Despite FDA discouragement of testing of the normality of log ratios (and differences) (32), clear deviations from normality of log-transformed ratios in some real BE datasets may call into question the validity of the unexamined use of TOST for BE. Evaluation of goodness-of-fit of the normal and t-distributions can provide additional valuable information on aptness of the statistical model for inference of BE. Far from adding unnecessary “regulatory burden,” employment of the correct statistical model is crucial for valid inference of BE. The BEST procedures enable the posterior diagnostics and employ data-informed simulations for type I error control, rather than relying upon the unattainable long-term frequency assumption.
Valid inference depends upon pre-specifying the analysis method in the Statistical Analysis Plan (SAP). BE data distributions in the real world may be approximately normal, lognormal, or neither and commonly include extreme values. In the SAP, an alternative BE evaluation method different from TOST should be based on sufficient scientific justification and communicated with the agency. Pivotal BE studies, preceded by small (underpowered for BE) pilot studies comparing formulations, offer an opportunity to select the “best candidate” distribution for powering the pivotal BE study. The use of BEST approach can also be pre-specified contingent on the scatterplot identification of extreme values as demonstrated above. When extreme values occur only or more frequently in the reference product data, adopting the BEST approach to gain more power can be scientifically justifiable.
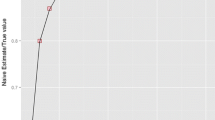
Two recent articles have used a Bayesian approach to bioequivalence on the log scale using the skew t-distribution, a four-parameter distribution which allows for non-symmetry (34, 35). Burger et al performed simulations for a crossover design with n=30 subjects and incorporated outliers by introducing contamination of 2.5 standard deviations of the lognormal with 1% probability (and 5% in the Supplemental Material). In the Supplemental Material, Burger et al reported 130 real datasets that showed outliers in 17 of the 130 data sets (13%). Among these, 17 datasets are 4 examples in which the Bayesian skew t estimator BayesT is outside the BE acceptance region [80%, 125%] but the two TOST-like estimators, Bayesian normal estimator (BayesN) and restricted maximum likelihood (REML), are not, and 5 examples in which REML and BayesN are outside but BayesT is inside. In 484 crossover studies submitted to FDA, we found that 28% rejected lognormality of T/R and 36% rejected normality of T-R by the Shapiro-Wilk test, mostly due to outliers. Additionally, in contrast to Burger et al, our study examined sample sizes from 10 to 50 and simulated outliers according to a 5% probability using a contamination standard deviation of 10 rather than 2.5, resulting in outliers more reflective of real-world data. We also examined the behavior in the realistic case in which the data are normal but not lognormal. An additional advantage is that BEST-BE provides easy access to diagnostic histograms for examining the posterior distribution of the parameters.
In this work, the extreme values were generated according to a symmetrical distribution around the underlying mean. Future work is warranted to assess methodology performance when data assume different distributions, with or without skewness after incorporating extreme values.
Conclusions
The 90% CrI from BEST is a Bayesian alternative for the 90% CI of TOST. The BEST 90% CrI can be derived from a distribution that is not normal and thus is more reflective of the observed log(T/R) distribution. In simulation studies, TOST and BEST demonstrate BE agreement when applied to lognormally distributed BE data. However, when T-R are normally distributed, then BEST demonstrates greater power than TOST and the BEST AMR demonstrates even greater power. In the presence of extreme values, BEST procedures significantly outperform TOST for underlying lognormal and normal BE distributions. Thus, TOST and BEST perform satisfactorily on lognormal T/R, while BEST is more accurate, requiring fewer subjects, when datasets contain extreme values or are normal for T-R. Application of BEST to BE data can be considered an informative alternative to TOST for evaluation of BE and for efficient development of BE formulations.
References
Schuirmann DJ. A comparison of the two one-sided tests procedure and the power approach for assessing the equivalence of average bioavailability. J Pharmacokinet Biopharm. 1987;15:657–80.
Peck CC, Campbell G. Bayesian approach to establish bioequivalence: why and how? Clin Pharmacol Ther. 2019;105:301–3.
Chow S, Liu M. Practical statistical issues in evaluation of average bioequivalence. J Biom Biostat. 2019;10:435.
Lacey LF, Keene ON, Pritchard JF, Bye A. Common noncompartmental pharmacokinetic variables: are they normally or log-normally distributed? J Biopharm Stat. 1997;7:171–8.
Patel HI. Dose-response in pharmacokinetics. Commun Stat Theory Methods. 1994;23:451–65.
Shen M, Russek-Cohen E, Slud EV. Checking distributional assumptions for pharmacokinetic summary statistics based on simulations with compartmental models. J Biopharm Stat. 2017;27:756–72.
Gelfand AE, Hills SE, Racine-Poon A, Smith AFM. Illustration of bayesian inference in normal data models using gibbs sampling. J Am Stat Assoc. 1990;85:972–85.
Ghosh P, Khattree R. Bayesian approach to average bioequivalence using Bayes' factor. J Biopharm Stat. 2003;13:719–34.
Ghosh P, Rosner GL. A semi-parametric bayesian approach to average bioequivalence. Stat Med. 2007;26:1224–36.
Selwyn MR, Dempster AP, Hall NR. A bayesian approach to bioequivalence for the 2 × 2 changeover design. Biometrics. 1981;37:11–21.
Weiss RE, Xia X, Zhang N, Wang H, Chi E. Bayesian methods for analysis of biosimilar phase iii trials. Stat Med. 2018;37:2938–53.
Kruschke JK. Bayesian estimation supersedes the t test. J Exp Psychol Gen. 2013;142:573–603.
Hespanhol L, Vallio CS, Costa LM, Saragiotto BT. Understanding and interpreting confidence and credible intervals around effect estimates. Braz J Phys Ther. 2019;23:290–301.
Kruschke JK, Liddell TM. The bayesian new statistics: hypothesis testing, estimation, meta-analysis, and power analysis from a bayesian perspective. Psychon Bull Rev. 2018;25:178–206.
U.S. Department of Health and Human Services FDA, Center for Drug Evaluation and Research (CDER). Statistical approaches to establishing bioequivalence 2001. https://www.fda.gov/media/70958/download.
Chung I, Oh J, Lee S, Jang IJ, Lee Y, Chung JY. A post hoc analysis of intra-subject coefficients of variation in pharmacokinetic measures to calculate optimal sample sizes for bioequivalence studies. Transl Clin Pharmacol. 2018;26:6–9.
Julious SA, Debarnot CA. Why are pharmacokinetic data summarized by arithmetic means? J Biopharm Stat. 2000;10:55–71.
Kruschke JK. Doing Bayesian data analysis: A tutorial with R and BUGS. Elsevier Academic Press; 2011. pp 143–91.
Chow S-C. Alternative approaches for assessing bioequivalence regarding normality assumptions. Drug Inf J. 1990;24(4):753–62. https://doi.org/10.1177/216847909002400411.
Razali NM, Wah YB. Power comparisons of shapiro-wilk, kolmogorov-smirnov, lilliefors and anderson-darling tests. J Stat Model Anal. 2011;2:21–33.
Lehmann EL. Elements of large sample theory. New York: Springer; 1999. p. 78.
Xiao W, Barron AM, Liu J. Robustness of bioequivalence procedures under box-cox alternatives. J Biopharm Stat. 1997;7:135–55.
Hauschke D, Steinijans VW, Diletti E. A distribution-free procedure for the statistical analysis of bioequivalence studies. Int J Clin Pharmacol Ther Toxicol. 1990;28:72–8.
Steinijans VW, Diletti E. Statistical analysis of bioavailability studies: parametric and nonparametric confidence intervals. Eur J Clin Pharmacol. 1983;24:127–36.
Bonate PL. Pharmacokinetic-pharmacodynamic modeling and simulation. New York: Springer; 2006. p 355–63.
Morita S, Thall PF, Muller P. Determining the effective sample size of a parametric prior. Biometrics. 2008;64:595–602.
Schuirmann DJ, Grosser S, Chattopadhyay S, Chow SC. On Bayesian analysis and hypothesis testing in the determination of bioequivalence. Clin Pharmacol Ther. 2019;105:304–6.
Chow, S.-C., & Liu, J.-P. (2008). Design and analysis of bioavailability and bioequivalence studies (3rd ed.). Chapman and Hall/CRC. https://doi.org/10.1201/9781420011678.
Chow SC, Tse SK. Outlier detection in bioavailability/bioequivalence studies. Stat Med. 1990;9:549–58.
Ki FY, Liu JP, Wang W, Chow SC. The impact of outlying subjects on decision of bioequivalence. J Biopharm Stat. 1995;5:71–94.
Wang W, Chow SC. Examining outlying subjects and outlying records in bioequivalence trials. J Biopharm Stat. 2003;13:43–56.
U.S. Department of Health and Human Services FDA, Center for Drug Evaluation and Research (CDER). Bioavailability studies submitted in ndas or inds - general considerations guidance for industry. April 2022. https://www.fda.gov/media/121311/download.
U.S. Department of Health and Human Services FDA, Center for Drug Evaluation and Research (CDER). Guidance for industry. Bioequivalence studies with pharmacokinetic endpoints for drugs submitted under an abbreviated new drug application. 2021. https://www.fda.gov/media/87219/download.
Burger DA, Schall R, van der Merwe S. A robust method for the assessment of average bioequivalence in the presence of outliers and skewness. Pharm Res. 2021;38(10):1697–709.
De Souza RM, Achcar JA, Martinez EZ, Mazucheli J. The use of asymmetric distributions in average bioequivalence. Stat Med. 2016;35(15):2525–42.
Acknowledgements
We want to give our special thanks to Drs. Jing Wang, Mingjiang Xu, and Somesh Chattopadhyay for their generous support in manuscript preparation, revision, and coordination. We also want to express our appreciation to the AAPS Journal reviewers for their constructive comments on our manuscript.
Author information
Authors and Affiliations
Contributions
C.P., G.C., and L.Z. conceptualized the study. C.P., G.C., L.Z., I.Y., M.H., and K.F. contributed to the data analysis plan, QC, and interpretation of the results and drafting the manuscript. I.Y., M.H., and K.F. collected and analyzed the data. I.Y., M.H., K.F., and L.Z. had full access to all the data in the study. All authors critically reviewed, edited, and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Disclaimer
This article reflects the views of the authors and should not be construed to represent FDA’s views or policies.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Peck, C., Campbell, G., Yoo, I. et al. Comparing a Bayesian Approach (BEST) with the Two One-Sided t-Tests (TOSTs) for Bioequivalence Studies. AAPS J 24, 97 (2022). https://doi.org/10.1208/s12248-022-00746-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12248-022-00746-6