Abstract
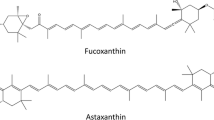
Fucoxanthin (FX) is a carotenoid with many pharmaceutical properties due to its antioxidant/anti-inflammatory and epigenetic effects. NFE2L2 is involved in the defense against oxidative stress/inflammation-mediated diseases, like anticancer effects elicited by phytochemicals including FX. However, the role of FX and NFE2L2 in metabolic rewiring, epigenomic reprogramming, and transcriptomic network in blocking pro-tumorigenic signaling and eliciting cancer-protective effects remains unknown. Herein, we utilized multi-omics approaches to evaluate the role of NFE2L2 and the impact of FX on tumor promoter TPA-induced skin cell transformation. FX blocked TPA-induced ROS and oxidized GSSG/reduced GSH in Nfe2l2wild-type(WT) but not Nfe2l2-knockdown (KD) cells. Both Nfe2l2 KD and TPA altered cellular metabolisms and metabolites which are tightly coupled to epigenetic machinery. The suppressive effects of FX on TPA-enhancedSAM/SAH was abrogated by Nfe2l2 KD indicating Nfe2l2 plays a critical role in FX-mediated metabolic rewiring and its potential consequences on epigenetic reprogramming. Epigenomic CpG methyl-seq revealed that FX attenuated TPA-induced differentially methylated regions (DMRs) of Uhrf1 and Dnmt1 genes. Transcriptomic RNA-seq showed that FX abrogated TPA-induced differentially expressed genes (DEGs) of Nfe2l2-related genes Nqo1, Ho1, and Keap1. Associative analysis of DEGs and DMRs identified that the mRNA expressions of Uhrf1 and Dnmt1 were correlated with the promoter CpG methylation status. Chromatin immunoprecipitation assay showed that FX restored Uhrf1 expression by regulating H3K27Me3 enrichment in the promoter region. In this context, FX/Nfe2l2’s redox signaling drives metabolic rewiring causing epigenetic and transcriptomic reprogramming potentially contributing to the protection of TPA-induced JB6 cellular transformation skin cancer model.

Graphical abstract






Similar content being viewed by others

References
McGuire S. World Cancer Report 2014. Geneva, Switzerland: World Health Organization, International Agency for Research on Cancer, WHO Press, 2015. Adv Nutr. 2016;7(2):418–9. https://doi.org/10.3945/an.116.012211.
Atalay S, Dobrzyńska I, Gęgotek A, Skrzydlewska E. Cannabidiol protects keratinocyte cell membranes following exposure to UVB and hydrogen peroxide. Redox Biology. 2020;36:101613. https://doi.org/10.1016/j.redox.2020.101613.
Wu WS, Tsai RK, Chang CH, Wang S, Wu JR, Chang YX. Reactive oxygen species mediated sustained activation of protein kinase C alpha and extracellular signal-regulated kinase for migration of human hepatoma cell Hepg2. Mol Cancer Res. 2006;4(10):747–58. https://doi.org/10.1158/1541-7786.MCR-06-0096.
Cross CE, Halliwell B, Borish ET, Pryor WA, Ames BN, Saul RL, et al. Oxygen radicals and human disease. Ann Intern Med. 1987;107(4):526–45. https://doi.org/10.7326/0003-4819-107-4-526.
Ames BN, Shigenaga MK, Hagen TM. Oxidants, antioxidants, and the degenerative diseases of aging. Proc Natl Acad Sci U S A. 1993;90(17):7915–22. https://doi.org/10.1073/pnas.90.17.7915.
Xian D, Lai R, Song J, Xiong X, Zhong J. Emerging perspective: role of increased ROS and redox imbalance in skin carcinogenesis. Oxid Med Cell Longev. 2019;2019:8127362. https://doi.org/10.1155/2019/8127362.
Zhang H, Tang Y, Zhang Y, Zhang S, Qu J, Wang X, Kong R, Han C, Liu Z. Fucoxanthin: a promising medicinal and nutritional ingredient. Evid Based Complement Alternat Med. 2015;2015:723515. https://doi.org/10.1155/2015/723515.
Muradian K, Vaiserman A, Min KJ, Fraifeld VE. Fucoxanthin and lipid metabolism: a minireview. Nutr Metab Cardiovasc Dis. 2015;25(10):891–7. https://doi.org/10.1016/j.numecd.2015.05.010.
Yang Y, Yang I, Cao M, Su ZY, Wu R, Guo Y, Fang M, Kong AN. Fucoxanthin elicits epigenetic modifications, Nrf2 activation and blocking transformation in mouse skin JB6 P+ cells. AAPS J. 2018;20(2):32. https://doi.org/10.1208/s12248-018-0197-6.
Datta R, Yoshinaga K, Kaneki M, Pandey P, Kufe D. Phorbol ester-induced generation of reactive oxygen species is protein kinase cbeta-dependent and required for SAPK activation. J Biol Chem. 2000;275(52):41000–3. https://doi.org/10.1074/jbc.M009322200.
Marengo B, Nitti M, Furfaro AL, Colla R, Ciucis CD, Marinari UM, Pronzato MA, Traverso N, Domenicotti C. Redox homeostasis and cellular antioxidant systems: crucial players in cancer growth and therapy. Oxid Med Cell Longev. 2016;2016:6235641. https://doi.org/10.1155/2016/6235641.
Zhao Y, Hu X, Liu Y, Dong S, Wen Z, He W, Zhang S, Huang Q, Shi M. ROS signaling under metabolic stress: cross-talk between AMPK and AKT pathway. Molecular Cancer. 2017;16(1):79. https://doi.org/10.1186/s12943-017-0648-1.
Lane AN, Higashi RM, Fan TW. Metabolic reprogramming in tumors: contributions of the tumor microenvironment. Genes Dis. 2020;7(2):185–98. https://doi.org/10.1016/j.gendis.2019.10.007.
Warburg O, Wind F, Negelein E. The Metabolism of Tumors in the Body. J Gen Physiol. 1927;8(6):519–30. https://doi.org/10.1085/jgp.8.6.519.
Missiroli S, Perrone M, Genovese I, Pinton P, Giorgi C. Cancer metabolism and mitochondria: finding novel mechanisms to fight tumours. EBioMedicine. 2020;59:102943. https://doi.org/10.1016/j.ebiom.2020.102943.
Wu Q, Ni X. ROS-mediated DNA methylation pattern alterations in carcinogenesis. Curr Drug Targets. 2015;16(1):13–9. https://doi.org/10.2174/1389450116666150113121054.
Colburn NH, Former BF, Nelson KA, Yuspa SH. Tumour promoter induces anchorage independence irreversibly. Nature. 1979;281(5732):589–91. https://doi.org/10.1038/281589a0.
Hosseini M, Dousset L, Mahfouf W, Serrano-Sanchez M, Redonnet-Vernhet I, Mesli S, Kasraian Z, Obre E, Bonneu M, Claverol S, Vlaski M, Ivanovic Z, Rachidi W, Douki T, Taieb A, Bouzier-Sore AK, Rossignol R, Rezvani HR. Energy metabolism rewiring precedes UVB-induced primary skin tumor formation. Cell Reports. 2018;23(12):3621–34. https://doi.org/10.1016/j.celrep.2018.05.060.
Janke R, Dodson AE, Rine J. Metabolism and epigenetics. Annual Review of Cell and Developmental Biology. 2015;31(1):473–96. https://doi.org/10.1146/annurev-cellbio-100814-125544.
Su X, Wellen KE, Rabinowitz JD. Metabolic control of methylation and acetylation. Curr Opin Chem Biol. 2016;30:52–60. https://doi.org/10.1016/j.cbpa.2015.10.030.
Cyr AR, Domann FE. The redox basis of epigenetic modifications: from mechanisms to functional consequences. Antioxid Redox Signal. 2011;15(2):551–89. https://doi.org/10.1089/ars.2010.3492.
Wu R, Li S, Hudlikar R, Wang L, Shannar A, Peter R, Chou PJ, Kuo HCD, Liu Z, Kong AN. Redox signaling, mitochondrial metabolism, epigenetics and redox active phytochemicals. Free Radical Biology and Medicine. 2020. https://doi.org/10.1016/j.freeradbiomed.2020.12.007.
Locato V, Cimini S, De Gara L. ROS and redox balance as multifaceted players of cross-tolerance: epigenetic and retrograde control of gene expression. Journal of experimental botany. 2018;69:3373–91. https://doi.org/10.1093/jxb/ery168.
Baird L, Yamamoto M. The molecular mechanisms regulating the KEAP1-NRF2 pathway. Mol Cell Biol. 2020;40(13). https://doi.org/10.1128/MCB.00099-20.
Hu R, Saw CL, Yu R, Kong AN. Regulation of NF-E2-related factor 2 signaling for cancer chemoprevention: antioxidant coupled with antiinflammatory. Antioxid Redox Signal. 2010;13(11):1679–98. https://doi.org/10.1089/ars.2010.3276.
Suzuki T, Motohashi H, Yamamoto M. Toward clinical application of the Keap1-Nrf2 pathway. Trends Pharmacol Sci. 2013;34(6):340–6. https://doi.org/10.1016/j.tips.2013.04.005.
Saw CL, Huang MT, Liu Y, Khor TO, Conney AH, Kong AN. Impact of Nrf2 on UVB-induced skin inflammation/photoprotection and photoprotective effect of sulforaphane. Mol Carcinog. 2011;50(6):479–86. https://doi.org/10.1002/mc.20725.
Khor TO, Huang MT, Prawan A, Liu Y, Hao X, Yu S, et al. Increased susceptibility of Nrf2 knockout mice to colitis-associated colorectal cancer. Cancer Prev Res (Phila). 2008;1(3):187–91. https://doi.org/10.1158/1940-6207.CAPR-08-0028.
Pelicano H, Martin DS, Xu RH, Huang P. Glycolysis inhibition for anticancer treatment. Oncogene. 2006;25(34):4633–46. https://doi.org/10.1038/sj.onc.1209597.
Saw CL, Yang AY, Huang MT, Liu Y, Lee JH, Khor TO, et al. Nrf2 null enhances UVB-induced skin inflammation and extracellular matrix damages. Cell Biosci. 2014;4:39. https://doi.org/10.1186/2045-3701-4-39.
Lin T-Y, Cantley LC, GMJAMRoOS-TTFN DN. NRF2 rewires cellular metabolism to support the antioxidant response; 2016. p. 107–8.
Naranjo Arcos M, Bauer P. Iron nutrition, oxidative stress, and pathogen defense. 2016.
Su ZY, Zhang C, Lee JH, Shu L, Wu TY, Khor TO, et al. Requirement and epigenetics reprogramming of Nrf2 in suppression of tumor promoter TPA-induced mouse skin cell transformation by sulforaphane. Cancer Prev Res (Phila). 2014;7(3):319–29. https://doi.org/10.1158/1940-6207.CAPR-13-0313-T.
Yang Y, Yin R, Wu R, Ramirez CN, Sargsyan D, Li S, Wang L, Cheng D, Wang C, Hudlikar R, Kuo HC, Lu Y, Kong AN. DNA methylome and transcriptome alterations and cancer prevention by triterpenoid ursolic acid in UVB-induced skin tumor in mice. Mol Carcinog. 2019;58(10):1738–53. https://doi.org/10.1002/mc.23046.
Bott A, Shen J, Tonelli C, Zhan L, Sivaram N, Jiang Y-P, et al. Glutamine anabolism plays a critical role in pancreatic cancer by coupling carbon and nitrogen metabolism. Cell reports. 2019;29:1287–98.e6. https://doi.org/10.1016/j.celrep.2019.09.056.
Dragan M, Nguyen M-U, Guzman S, Goertzen C, Brackstone M, Dhillo WS, Bech PR, Clarke S, Abbara A, Tuck AB, Hess DA, Pine SR, Zong WX, Wondisford FE, Su X, Babwah AV, Bhattacharya M. G protein-coupled kisspeptin receptor induces metabolic reprograming and tumorigenesis in estrogen receptor-negative breast cancer. Cell Death & Disease. 2020;11(2):106. https://doi.org/10.1038/s41419-020-2305-7.
Guo Y, Su ZY, Zhang C, Gaspar JM, Wang R, Hart RP, Verzi MP, Kong ANT. Mechanisms of colitis-accelerated colon carcinogenesis and its prevention with the combination of aspirin and curcumin: transcriptomic analysis using RNA-seq. Biochem Pharmacol. 2017;135:22–34. https://doi.org/10.1016/j.bcp.2017.02.021.
Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. J EMBnet.journal. 2011;17(1):3. https://doi.org/10.14806/ej.17.1.200.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nature Methods. 2015;12(4):357–60. https://doi.org/10.1038/nmeth.3317.
Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014;30(7):923–30. https://doi.org/10.1093/bioinformatics/btt656.
Wang L, Feng Z, Wang X, Wang X, Zhang X. DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics. 2010;26(1):136–8. https://doi.org/10.1093/bioinformatics/btp612.
Gaspar JM, Hart RP. DMRfinder: efficiently identifying differentially methylated regions from MethylC-seq data. BMC Bioinformatics. 2017;18(1):528. https://doi.org/10.1186/s12859-017-1909-0.
Yu G, Wang L-G, He Q-Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics. 2015;31(14):2382–3. https://doi.org/10.1093/bioinformatics/btv145%JBioinformatics.
Zitka O, Skalickova S, Gumulec J, Masarik M, Adam V, Hubalek J, et al. Redox status expressed as GSH:GSSG ratio as a marker for oxidative stress in paediatric tumour patients. Oncol Lett. 2012;4(6):1247–53. https://doi.org/10.3892/ol.2012.931.
Wong CC, Qian Y, Yu J. Interplay between epigenetics and metabolism in oncogenesis: mechanisms and therapeutic approaches. Oncogene. 2017;36(24):3359–74. https://doi.org/10.1038/onc.2016.485.
Bostick M, Kim JK, Esteve PO, Clark A, Pradhan S, Jacobsen SE. UHRF1 plays a role in maintaining DNA methylation in mammalian cells. Science. 2007;317(5845):1760–4. https://doi.org/10.1126/science.1147939.
Nishiyama A, Mulholland CB, Bultmann S, Kori S, Endo A, Saeki Y, Qin W, Trummer C, Chiba Y, Yokoyama H, Kumamoto S, Kawakami T, Hojo H, Nagae G, Aburatani H, Tanaka K, Arita K, Leonhardt H, Nakanishi M. Two distinct modes of DNMT1 recruitment ensure stable maintenance DNA methylation. Nature Communications. 2020;11(1):1222. https://doi.org/10.1038/s41467-020-15006-4.
Brenet F, Moh M, Funk P, Feierstein E, Viale AJ, Socci ND, Scandura JM. DNA methylation of the first exon is tightly linked to transcriptional silencing. PLoS One. 2011;6(1):e14524. https://doi.org/10.1371/journal.pone.0014524.
Mihaylova MM, Shaw RJ. The AMPK signalling pathway coordinates cell growth, autophagy and metabolism. Nature Cell Biology. 2011;13(9):1016–23. https://doi.org/10.1038/ncb2329.
Jang M, Kim SS, Lee J. Cancer cell metabolism: implications for therapeutic targets. Exp Mol Med. 2013;45:e45. https://doi.org/10.1038/emm.2013.85.
Zhang Y, Reinberg D. Transcription regulation by histone methylation: interplay between different covalent modifications of the core histone tails. Genes Dev. 2001;15(18):2343–60. https://doi.org/10.1101/gad.927301.
Ferrari KJ, Scelfo A, Jammula S, Cuomo A, Barozzi I, Stutzer A, et al. Polycomb-dependent H3K27me1 and H3K27me2 regulate active transcription and enhancer fidelity. Mol Cell. 2014;53(1):49–62. https://doi.org/10.1016/j.molcel.2013.10.030.
Huang MT, Smart RC, Wong CQ, Conney AH. Inhibitory effect of curcumin, chlorogenic acid, caffeic acid, and ferulic acid on tumor promotion in mouse skin by 12-O-tetradecanoylphorbol-13-acetate. Cancer Res. 1988;48(21):5941–6.
Conney AH, Wang ZY, Huang MT, Ho CT, Yang CS. Inhibitory effect of green tea on tumorigenesis by chemicals and ultraviolet light. Prev Med. 1992;21(3):361–9. https://doi.org/10.1016/0091-7435(92)90043-h.
Locato V, Cimini S, De Gara L. ROS and redox balance as multifaceted players of cross-tolerance: epigenetic and retrograde control of gene expression. J Exp Bot. 2018;69(14):3373–91. https://doi.org/10.1093/jxb/ery168.
Elstrom RL, Bauer DE, Buzzai M, Karnauskas R, Harris MH, Plas DR, Zhuang H, Cinalli RM, Alavi A, Rudin CM, Thompson CB. Akt stimulates aerobic glycolysis in cancer cells. Cancer Res. 2004;64(11):3892–9. https://doi.org/10.1158/0008-5472.CAN-03-2904.
Zhong H, Yuan P, Li Y, Batonon-Alavo D, Deschamps C, Feng B, Zhang X, Che L, Lin Y, Xu S, Li J, Zhuo Y, Tian G, Tang J, Jiang X, Huang L, Wu C, Wu, Fang Z. Methionine protects mammary cells against oxidative stress through producing S-adenosylmethionine to maintain mTORC1 signaling activity. Oxidative Medicine and Cellular Longevity. 2021;2021:5550196. https://doi.org/10.1155/2021/5550196.
Wallace DC, Fan W. Energetics, epigenetics, mitochondrial genetics. Mitochondrion. 2010;10(1):12–31. https://doi.org/10.1016/j.mito.2009.09.006.
Hudlikar R, Wang L, Wu R, Li S, Peter R, Shannar A, Chou PJ, Liu X, Liu Z, Kuo HD, Kong AN. Epigenetics/epigenomics and prevention of early stages of cancer by isothiocyanates. Cancer Prev Res (Phila). 2020;14:151–64. https://doi.org/10.1158/1940-6207.CAPR-20-0217.
Mentch SJ, Mehrmohamadi M, Huang L, Liu X, Gupta D, Mattocks D, Gómez Padilla P, Ables G, Bamman MM, Thalacker-Mercer AE, Nichenametla SN, Locasale JW. Histone methylation dynamics and gene regulation occur through the sensing of one-carbon metabolism. Cell Metab. 2015;22(5):861–73. https://doi.org/10.1016/j.cmet.2015.08.024.
Gut P, Verdin E. The nexus of chromatin regulation and intermediary metabolism. Nature. 2013;502(7472):489–98. https://doi.org/10.1038/nature12752.
Shyh-Chang N, Locasale JW, Lyssiotis CA, Zheng Y, Teo RY, Ratanasirintrawoot S, Zhang J, onder T, Unternaehrer JJ, Zhu H, Asara JM, Daley GQ, Cantley LC. Influence of threonine metabolism on S-adenosylmethionine and histone methylation. Science. 2013;339(6116):222–6. https://doi.org/10.1126/science.1226603.
Bashtrykov P, Jankevicius G, Jurkowska RZ, Ragozin S, Jeltsch A. The UHRF1 protein stimulates the activity and specificity of the maintenance DNA methyltransferase DNMT1 by an allosteric mechanism. J Biol Chem. 2014;289(7):4106–15. https://doi.org/10.1074/jbc.M113.528893.
Bronner C, Alhosin M, Hamiche A, Mousli M. Coordinated dialogue between UHRF1 and DNMT1 to ensure faithful inheritance of methylated DNA patterns. Genes (Basel). 2019;10(1). https://doi.org/10.3390/genes10010065.
Vousden KH, Lane DP. p53 in health and disease. Nat Rev Mol Cell Biol. 2007;8(4):275–83. https://doi.org/10.1038/nrm2147.
Stegh AH. Targeting the p53 signaling pathway in cancer therapy - the promises, challenges and perils. Expert Opin Ther Targets. 2012;16(1):67–83. https://doi.org/10.1517/14728222.2011.643299.
Rivlin N, BroshR Oren M, RotterV,. Mutations in the p53 tumor suppressor gene: Important milestones at the various steps of tumorigenesis. Genes and Cancer. 2011;2(4):466–74. https://doi.org/10.1177/1947601911408889.
Zhang W, Liu HT. MAPK signal pathways in the regulation of cell proliferation in mammalian cells. Cell Research. 2002;12(1):9–18. https://doi.org/10.1038/sj.cr.7290105.
Braicu C, Buse M, Busuioc C, Drula R, Gulei D, Raduly L, et al. A comprehensive review on MAPK: a promising therapeutic target in cancer. Cancers (Basel). 2019;11(10). https://doi.org/10.3390/cancers11101618.
Faubert B, Boily G, Izreig S, Griss T, Samborska B, Dong Z, Dupuy F, Chambers C, Fuerth BJ, Viollet B, Mamer OA, Avizonis D, DeBerardinis RJ, Siegel PM, Jones RG. AMPK is a negative regulator of the Warburg effect and suppresses tumor growth in vivo. Cell Metab. 2013;17(1):113–24. https://doi.org/10.1016/j.cmet.2012.12.001.
Vander Heiden MG, Cantley LC, Thompson CB. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science. 2009;324(5930):1029–33. https://doi.org/10.1126/science.1160809.
Acknowledgements
We thank all the members of Professor Ah-Ng Kong’s laboratory for their invaluable discussion and technical support for preparation of this manuscript.
Funding
This work was supported in part by institutional funds and by R01 AT009152 from the National Center for Complementary and Integrative Health (NCCIH), R01 CA200129 from the National Cancer Institute (NCI), and P30 ES005022 from the National Institute of Environmental Health (NIEHS).
Author information
Authors and Affiliations
Contributions
Participated in overall research study design: Lujing Wang and Ah-Ng Kong
Conducted experiments: Lujing Wang, Ameya Phadnis, and Shan Su
Performed data analysis: Lujing Wang, Renyi Wu, Davit Sargsyan, Shanyi Li, Hsiao-Chen Kuo, Pochung Chou, Yujue Wang, and Xiaoyang Su
Wrote the manuscript: Lujing Wang, Md Shahid Sarwar, and Ah-Ng Kong
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
Nfe2l2 KDmediated metabolites and metabolism pathway regulation in JB6 cells. Top 50 modulated metabolites with their corresponding P-value after Nfe2l2 KD are presented in Tab 1; the pathway analysis produced the significantly regulated metabolism pathways after Nfe2l2 KD are presented in Tab 2; and the pathway enrichment analysis produced significantly regulated pathways after Nfe2l2 KD are presented in Tab 3. (XLSX 13 kb)
ESM 2
DMRs and DEGs which are negatively regulated by TPA and FX. Tab 1: DMRs which are negatively regulated in TPA vs Control and TPA+FX vs TPA comparison groups; Tab 2: DEGs which are negatively regulated in TPA vs Control and TPA+FX vs TPA comparison groups. (XLSX 18 kb)
ESM 3
Correlated genes which possess negatively DNA methylation and gene expression patterns in TPA vs Control and TPA+FX vs TPA comparison groups filtered by integration of RNA-seq and Methyl-seq Analysis (XLSX 12 kb)
ESM 4
(DOCX 15 kb)
Rights and permissions
About this article
Cite this article
Wang, L., Wu, R., Sargsyan, D. et al. Nfe2l2 Regulates Metabolic Rewiring and Epigenetic Reprogramming in Mediating Cancer Protective Effect by Fucoxanthin. AAPS J 24, 30 (2022). https://doi.org/10.1208/s12248-022-00679-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12248-022-00679-0