Abstract
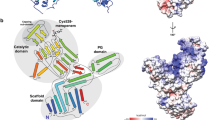
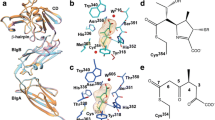
The final step of peptidoglycan (PG) synthesis in all bacteria is the formation of cross-linkage between PG-stems. The cross-linking between amino acids in different PG chains gives the peptidoglycan cell wall a 3-dimensional structure and adds strength and rigidity to it. There are two distinct types of cross-linkages in bacterial cell walls. D,D-transpeptidase (D,D-TPs) generate the classical 4➔3 cross-linkages and the L,D-transpeptidase (L,D-TPs) generate the 3➔3 non-classical peptide cross-linkages. The present study is aimed at understanding the nature of drug resistance associated with L,D-TP and gaining insights for designing novel antibiotics against multi-drug resistant bacteria. Penicillin and cephalosporin classes of β-lactams cannot inhibit L,D-TP function; however, carbapenems inactivate its function. We analyzed the structure of L,D-TP of Mycobacterium tuberculosis in the apo form and in complex with meropenem and imipenem. The periplasmic region of L,D-TP folds into three domains. The catalytic residues are situated in the C-terminal domain. The acylation reaction occurs between carbapenem antibiotics and the catalytic Cys-354 forming a covalent complex. This adduct formation mimics the acylation of L,D-TP with the donor PG-stem. A novel aspect of this study is that in the crystal structures of the apo and the carbapenem complexes, the N-terminal domain has a muropeptide unit non-covalently bound to it. Another interesting observation is that the calcium complex crystallized as a dimer through head and tail interactions between the monomers.









Similar content being viewed by others
References
Rogers HJ, James CA, Morrison PJ, Bradbrook ID. Effect of cimetidine on oral absorption of ampicillin and cotrimoxazole. J Antimicrob Chemother. 1980;6(2):297–300. https://doi.org/10.1093/jac/6.2.297.
van Heijenoort J. Formation of the glycan chains in the synthesis of bacterial peptidoglycan. Glycobiology. 2001;11(3):25R–36R. https://doi.org/10.1093/glycob/11.3.25R.
Ghuysen JM, Goffin C. Lack of cell wall peptidoglycan versus penicillin sensitivity: new insights into the chlamydial anomaly. Antimicrob Agents Chemother. 1999;43(10):2339–44.
Vollmer W, Holtje JV. The architecture of the murein (peptidoglycan) in gram-negative bacteria: vertical scaffold or horizontal layer(s)? J Bacteriol. 2004;186(18):5978–87. https://doi.org/10.1128/JB.186.18.5978-5987.2004.
Matsuhashi M. Biosynthesis in the bacterial cell wall. Tanpakushitsu Kakusan Koso. 1966;11(10):875–86.
Jamin M, Wilkin JM, Frere JM. Bacterial DD-transpeptidases and penicillin. Essays Biochem. 1995;29:1–24.
Goffin C, Ghuysen JM. Biochemistry and comparative genomics of SxxK superfamily acyltransferases offer a clue to the mycobacterial paradox: presence of penicillin-susceptible target proteins versus lack of efficiency of penicillin as therapeutic agent. Microbiol Mol Biol Rev. 2002;66(4):702–38, table of contents. https://doi.org/10.1128/MMBR.66.4.702-738.2002.
Lavollay M, Arthur M, Fourgeaud M, Dubost L, Marie A, Riegel P, et al. The beta-lactam-sensitive D,D-carboxypeptidase activity of Pbp4 controls the L,D and D,D transpeptidation pathways in Corynebacterium jeikeium. Mol Microbiol. 2009;74(3):650–61. https://doi.org/10.1111/j.1365-2958.2009.06887.x.
Schleifer KH, Kandler O. Peptidoglycan types of bacterial cell walls and their taxonomic implications. Bacteriol Rev. 1972;36(4):407–77.
Mainardi JL, Legrand R, Arthur M, Schoot B, van Heijenoort J, Gutmann L. Novel mechanism of beta-lactam resistance due to bypass of DD-transpeptidation in Enterococcus faecium. J Biol Chem. 2000;275(22):16490–6. https://doi.org/10.1074/jbc.M909877199.
Gupta R, Lavollay M, Mainardi JL, Arthur M, Bishai WR, Lamichhane G. The mycobacterium tuberculosis protein LdtMt2 is a nonclassical transpeptidase required for virulence and resistance to amoxicillin. Nat Med. 2010;16(4):466–9. https://doi.org/10.1038/nm.2120.
Peltier J, Courtin P, El Meouche I, Lemee L, Chapot-Chartier MP, Pons JL. Clostridium difficile has an original peptidoglycan structure with a high level of N-acetylglucosamine deacetylation and mainly 3-3 cross-links. J Biol Chem. 2011;286(33):29053–62. https://doi.org/10.1074/jbc.M111.259150.
Kuzin AP, Liu H, Kelly JA, Knox JR. Binding of cephalothin and cefotaxime to D-ala-D-ala-peptidase reveals a functional basis of a natural mutation in a low-affinity penicillin-binding protein and in extended-spectrum beta-lactamases. Biochemistry. 1995;34(29):9532–40. https://doi.org/10.1021/bi00029a030.
Bugg TD, Braddick D, Dowson CG, Roper DI. Bacterial cell wall assembly: still an attractive antibacterial target. Trends Biotechnol. 2011;29(4):167–73. https://doi.org/10.1016/j.tibtech.2010.12.006.
Hugonnet JE, Tremblay LW, Boshoff HI, Barry CE 3rd, Blanchard JS. Meropenem-clavulanate is effective against extensively drug-resistant Mycobacterium tuberculosis. Science. 2009;323(5918):1215–8. https://doi.org/10.1126/science.1167498.
Erdemli SB, Gupta R, Bishai WR, Lamichhane G, Amzel LM, Bianchet MA. Targeting the cell wall of Mycobacterium tuberculosis: structure and mechanism of L,D-transpeptidase 2. Structure. 2012;20(12):2103–15. https://doi.org/10.1016/j.str.2012.09.016.
Kim HS, Kim J, Im HN, Yoon JY, An DR, Yoon HJ, et al. Structural basis for the inhibition of Mycobacterium tuberculosis L,D-transpeptidase by meropenem, a drug effective against extensively drug-resistant strains. Acta Crystallogr D Biol Crystallogr. 2013;69(Pt 3):420–31. https://doi.org/10.1107/S0907444912048998.
Both D, Steiner EM, Stadler D, Lindqvist Y, Schnell R, Schneider G. Structure of LdtMt2, an L,D-transpeptidase from Mycobacterium tuberculosis. Acta Crystallogr D Biol Crystallogr. 2013;69(Pt 3):432–41. https://doi.org/10.1107/S0907444912049268.
Correale S, Esposito C, Pirone L, Vitagliano L, Di Gaetano S, Pedone E. A biophysical characterization of the folded domains of KCTD12: insights into interaction with the GABAB2 receptor. J Mol Recognit. 2013;26(10):488–95. https://doi.org/10.1002/jmr.2291.
Lecoq L, Bougault C, Triboulet S, Dubee V, Hugonnet JE, Arthur M, et al. Chemical shift perturbations induced by the acylation of Enterococcus faecium L,D-transpeptidase catalytic cysteine with ertapenem. Biomol NMR Assign. 2014;8(2):339–43. https://doi.org/10.1007/s12104-013-9513-3.
Li WJLD, Hu YL, Zhang XE, Bi LJ, Wang DC. Crystal structure of L,D-transpeptidase LdtMt2 in complex with meropenem reveals the mechanism of carbapenem against Mycobacterium tuberculosis. Cell Res. 2013;23(5):728–31. https://doi.org/10.1038/cr.2013.53.
Van Duyne GD, Standaert RF, Karplus PA, Schreiber SL, Clardy J. Atomic structures of the human immunophilin FKBP-12 complexes with FK506 and rapamycin. J Mol Biol. 1993;229(1):105–24. https://doi.org/10.1006/jmbi.1993.1012.
Otwinowski Z, Minor W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 1997;276:307–26. https://doi.org/10.1016/S0076-6879(97)76066-X.
Cruickshank DW. Remarks about protein structure precision. Acta Crystallogr D Biol Crystallogr. 1999;55(3):583–601. https://doi.org/10.1107/S0907444998012645.
Murshudov GN, Vagin AA, Dodson EJ. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr D Biol Crystallogr. 1997;53(Pt 3):240–55. https://doi.org/10.1107/S0907444996012255.
Cowtan KD, Main P. Phase combination and cross validation in iterated density-modification calculations. Acta Crystallogr D Biol Crystallogr. 1996;52(Pt 1):43–8. https://doi.org/10.1107/S090744499500761X.
Langer G, Cohen SX, Lamzin VS, Perrakis A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nat Protoc. 2008;3(7):1171–9. https://doi.org/10.1038/nprot.2008.91.
Sheldrick GM. A short history of SHELX. Acta Crystallogr A: Found Crystallogr. 2008;64(Pt 1):112–22. https://doi.org/10.1107/S0108767307043930.
Emsley P, Lohkamp B, Scott WG, Cowtan K. Features and development of Coot. Acta Crystallogr D Biol Crystallogr. 2010;66(Pt 4):486–501. https://doi.org/10.1107/S0907444910007493.
Bielnicki J, Devedjiev Y, Derewenda U, Dauter Z, Joachimiak A, Derewenda ZS. B. subtilis ykuD protein at 2.0 a resolution: insights into the structure and function of a novel, ubiquitous family of bacterial enzymes. Proteins. 2006;62(1):144–51. https://doi.org/10.1002/prot.20702.
Tan TC, Mijts BN, Swaminathan K, Patel BK, Divne C. Crystal structure of the polyextremophilic alpha-amylase AmyB from Halothermothrix orenii: details of a productive enzyme-substrate complex and an N domain with a role in binding raw starch. J Mol Biol. 2008;378(4):852–70. https://doi.org/10.1016/j.jmb.2008.02.041.
LaPorte SL, Juo ZS, Vaclavikova J, Colf LA, Qi X, Heller NM, et al. Molecular and structural basis of cytokine receptor pleiotropy in the interleukin-4/13 system. Cell. 2008;132(2):259–72. https://doi.org/10.1016/j.cell.2007.12.030.
Matthews DA, Alden RA, Birktoft JJ, Freer T, Kraut J. Re-examination of the charge relay system in subtilisin comparison with other serine proteases. J Biol Chem. 1977;252(24):8875–83.
Robertus JD, Kraut J, Alden RA, Birktoft JJ. Subtilisin; a stereochemical mechanism involving transition-state stabilization. Biochemistry. 1972;11(23):4293–303. https://doi.org/10.1021/bi00773a016.
Betts JC. Transcriptomics and proteomics: tools for the identification of novel drug targets and vaccine candidates for tuberculosis. IUBMB Life. 2002;53(4–5):239–42. https://doi.org/10.1080/15216540212651.
Keren I, Minami S, Rubin E, Lewis K. Characterization and transcriptome analysis of Mycobacterium tuberculosis persisters. MBio. 2011;2(3):e00100–11. https://doi.org/10.1128/mBio.00100-11.
Mainardi JL, Hugonnet JE, Rusconi F, Fourgeaud M, Dubost L, Moumi AN, et al. Unexpected inhibition of peptidoglycan LD-transpeptidase from enterococcus faecium by the beta-lactam imipenem. J Biol Chem. 2007;282(42):30414–22. https://doi.org/10.1074/jbc.M704286200.
Biarrotte-Sorin S, Hugonnet JE, Delfosse V, Mainardi JL, Gutmann L, Arthur M, et al. Crystal structure of a novel beta-lactam-insensitive peptidoglycan transpeptidase. J Mol Biol. 2006;359(3):533–8. https://doi.org/10.1016/j.jmb.2006.03.014.
Triboulet S, Arthur M, Mainardi JL, Veckerle C, Dubee V, Nguekam-Moumi A, et al. Inactivation kinetics of a new target of beta-lactam antibiotics. J Biol Chem. 2011;286(26):22777–84. https://doi.org/10.1074/jbc.M111.239988.
Triboulet S, Dubee V, Lecoq L, Bougault C, Mainardi JL, Rice LB, et al. Kinetic features of L,D-transpeptidase inactivation critical for beta-lactam antibacterial activity. PLoS One. 2013;8(7):e67831. https://doi.org/10.1371/journal.pone.0067831.
Fedarovich A, Nicholas RA, Davies C. Unusual conformation of the SxN motif in the crystal structure of penicillin-binding protein A from Mycobacterium tuberculosis. J Mol Biol. 2010;398(1):54–65. https://doi.org/10.1016/j.jmb.2010.02.046.
Das R, Pandey GK. Expressional analysis and role of calcium regulated kinases in abiotic stress signaling. Curr Genomics. 2010;11(1):2–13. https://doi.org/10.2174/138920210790217981.
Wagner D, Maser J, Lai B, Cai Z, Barry CE 3rd, Honer Zu Bentrup K, et al. Elemental analysis of Mycobacterium avium-, Mycobacterium tuberculosis-, and mycobacterium smegmatis-containing phagosomes indicates pathogen-induced microenvironments within the host cell’s endosomal system. J Immunol. 2005;174(3):1491–500. https://doi.org/10.4049/jimmunol.174.3.1491.
Acknowledgements
The authors thank Dr. Henry Bellamy, CAMD, for SAD data collection, and SSRL for high resolution data collection. Dr. Kuppan Gokulan is supported by the FDA Commissioner Fellowship Program. The authors would like to thank Dr. Kevin Young (UAMS), and Dr. Sutherland (NCTR) for their helpful review and critique of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Guest Editors: Shraddha Thakkar and Cesar M. Compadre
Rights and permissions
About this article
Cite this article
Gokulan, K., Khare, S., Cerniglia, C.E. et al. Structure and Inhibitor Specificity of L,D-Transpeptidase (LdtMt2) from Mycobacterium tuberculosis and Antibiotic Resistance: Calcium Binding Promotes Dimer Formation. AAPS J 20, 44 (2018). https://doi.org/10.1208/s12248-018-0193-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12248-018-0193-x