Abstract
ADB-CHMINACA (MAB-CHMINACA) is a new synthetic cannabinoid with high potency and many reported adverse events and fatalities. The drug is currently scheduled in several countries in Europe and the USA. Analytical methods need to be developed to confirm ADB-CHMINACA intake for clinical and forensic programs. For many synthetic cannabinoids, parent compound is not detectable in biological samples after intake, making the detection of metabolites the only way to prove consumption. Therefore, detection of ADB-CHMINACA metabolites in biological specimens is critical. Since there are currently no published data on ADB-CHMINACA metabolism, we aimed to identify its major metabolites. Cryopreserved human hepatocytes were incubated with 10 μmol/L ADB-CHMINACA for 3 h. Incubations were analyzed with liquid chromatography on a biphenyl column, high resolution tandem mass spectrometry (orbitrap), and metabolite identification software. A reference standard of six commercially available potential metabolites was simultaneously analyzed under the same conditions to allow correct assignment of isomers. We detected ten major metabolites. Biotransformations mainly occurred at the cyclohexylmethyl tail of the compound, as also observed with structural analogs’ metabolism. Minor reactions also occurred at the tert-butyl chain. Only two analytical standards of potential metabolites matched an actual metabolite detected in hepatocyte incubations. We recommend A9 (ADB-CHMINACA hydroxycyclohexylmethyl), A4 (ADB-CHMINACA 4″-hydroxycyclohexyl), and A6 (ADB-CHMINACA hydroxycyclohexylmethyl) as metabolite targets to document ADB-CHMINACA intake in clinical and forensic cases. Additionally, these results will guide analytical standard manufacturers to better provide suitable references for further studies on ADB-CHMINACA metabolism.
Similar content being viewed by others
INTRODUCTION
Synthetic cannabinoids (SC) are novel psychoactive substances (NPS) producing their effects via the endocannabinoid system, as central (CB1) and peripheral (CB2) cannabinoid receptor agonists (1). SC potency and binding affinity for CB1 and CB2 are often much higher than classic cannabinoids, such as Δ9-tetrahydrocannabinol (THC), the main psychoactive constituent of cannabis (1–4). Consequently, SC are actively abused since 2008 as “legal highs” (5). Many countries scheduled SC as illicit substances, but clandestine laboratories continue to produce new molecules to circumvent the law. In Japan, where the scheduling process is the fastest, 858 SC were controlled by April 2015 (6). As the occurrence of consumption and the diversity of SC increased, SC intake is associated with considerable morbidity and mortality (7–10).
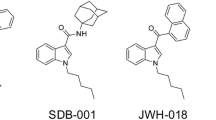
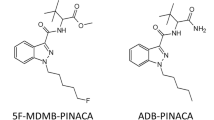
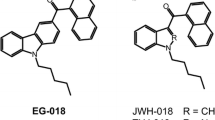
ADB-CHMINACA (N-[1′-(aminocarbonyl)-2′,2′-dimethylpropyl]-1-(cyclohexylmethyl)-1H-indazole-3-carboxamide), also known as MAB-CHMINACA, is an indazole-based SC designed by Pfizer in 2009 (11). The indazole core is substituted at N1 by a cyclohexylmethyl tail and at C3 by a L-tert-leucinamide side chain via a carboxamide link (Fig. 1). Binding affinity for CB1 receptors and half maximal effective concentration were K i = 0.289 nM and EC50 = 0.620 nM, respectively (no data for CB2) (11), making ADB-CHMINACA one of the most potent SC to date (1–4). Its binding affinity and potency are 140 and 270 times higher than those of THC, respectively (1,3). ADB-CHMINACA was first identified in herbal blends labeled “GM sapphire” in Japan in late 2014 (12). According to internet drug forums, ADB-CHMINACA may be marketed in the form of a white powder that can be smoked or ingested. Typical doses range between 0.25 and 0.5 mg and the effect ends between 2 and 3 h after smoking or 8 and 15 h after ingestion. Reported symptoms include euphoria, relaxation, improved mood, increased motivation and creativity, increased appetite, drowsiness, and dry cough (13).
Adverse effects associated with ADB-CHMINACA intake were observed in four patients who smoked a white powder labeled “AM-2201.” Observed effects included vomiting, seizures, limb twisting, muscle tremors, aggression, slurred speech, pressure spikes, wheezing, respiratory failure, and losses of consciousness. Blood concentrations ranged from 1.3 to 14.6 μg/L between 1 and 2 h after intake (13). In 2014, ADB-CHMINACA was involved in a fatal case of SC intoxication. Herbal blends that were ingested contained 133 ± 4.5 mg/g ADB-CHMINACA and 49.2 ± 2.5 mg/g 5F-ADBICA, another SC (12,14). Postmortem femoral vein blood, right heart blood, and left heart blood concentrations of ADB-CHMINACA were 6.1 ± 0.2, 10.6 ± 0.7, and 9.3 ± 0.3 μg/L, respectively (15). Multiple outbreaks of severe intoxications associated with ADB-CHMINACA intake were also reported in the USA (Kansas, Louisiana, Mississippi, Maryland, Texas, and Virginia), including hundreds of hospitalizations and at least three fatalities (7,16). Because of increasing prevalence and safety concerns, ADB-CHMINCA was controlled in Japan in 2014 (17), in Singapore in 2015 (18), and in several European countries. The drug was placed into schedule I in the USA in late 2015 (16).
Clinical evaluation of ADB-CHMINACA intake is not possible, due to lack of toxicology data, making analytical toxicology the most efficient way to prove consumption. Urine is the most common matrix for drug testing because of the longer window of detection compared to blood and oral fluid, a higher concentration of metabolites for several days after intake, and a generally larger sample volume. However, SC are highly metabolized and detection of parent SC in urine is rare (19–23). This is the case for AB-CHMINACA, desmethyl analog of ADB-CHMINACA (Fig. 1), for which 15 different metabolites were detected in a urine specimen from an authentic case of abuse, with no detectable parent compound (24). This is also the case of MDMB-CHMICA, another ADB-CHMINACA analog with an indole core and a terminal methyl ester instead of a carboxamide (Fig. 1) (25). In a fatal case of intoxication involving ADB-CHMINACA, the drug was quantified in 14 different postmortem body fluids and tissues (6.1 to 156 μg/L or g) but was not detectable in urine (15). Under these conditions, detection of ADB-CHMINACA urinary metabolites may be critical to document intake. To date, nothing is known about ADB-CHMINACA metabolism, although several theoretical metabolite standards are commercially available (most likely based on in silico predictions or extrapolation from analog metabolism). Therefore, we aimed to investigate human ADB-CHMINACA metabolism to identify specific urinary markers of intake.
Incubation with human liver microsomes (HLM) is the most common in vitro model for metabolite profiling due to low cost, availability, and simplicity of use. However, they may not predict in vivo metabolites or the relative abundance of metabolites, as observed with 5F-AKB48 (26) and AM-2201 (27,28). Human hepatocyte profiles generally match better with authentic urine SC metabolites because they are functioning complete cells and contain comprehensive phase I and II metabolic enzymes and cofactors, uptake and efflux transporters, and drug binding proteins (29–32). Therefore, we incubated ADB-CHMINACA with human hepatocytes and identified metabolites with liquid chromatography-high resolution tandem mass spectrometry (LC-HRMS/MS), following our previously recommended workflow (33). Additionally, we compared our results to commercially available reference standards of theoretical ADB-CHMINACA metabolites.
EXPERIMENTAL
Chemicals and Reagents
ADB-CHMINACA and ADB-CHMINACA M1 (N-(1-amino-3,3-dimethyl-1-oxobutan-2-yl)-1-((4-hydroxycyclohexyl)methyl)-1H-indazole-3-carboxamide), M2 ((S)-2-(1-(cyclohexylmethyl)-1H-indazole-3-carboxamido)-3,3-dimethylbutanoic acid), M3 ((S)-2-(1-((4-hydroxycyclohexyl)methyl)-1H-indazole-3-carboxamido)-3,3-dimethylbutanoic acid), M7 (3-(1-(cyclohexylmethyl)-1H-indazole-3-carboxamido)-2,2-dimethylsuccinic acid), M10 (1-(cyclohexylmethyl)-N-(4,4-dimethyl-2-oxotetrahydrofuran-3-yl)-1H-indazole-3-carboxamide), and M11 (N-(1-amino-4-hydroxy-3,3-dimethyl-1-oxobutan-2-yl)-1-((4-hydroxycyclohexyl)methyl)-1H-indazole-3-carboxamide) standards were purchased from Cayman Chemical (Ann Arbor, MI, USA) (Fig. 2). Diclofenac standard was purchased from Toronto Research Chemicals (Toronto, Canada). M10 was dissolved in acetonitrile to prevent hydrolysis, while other standards were dissolved in methanol. LC-MS grade water, methanol, and formic acid (Optima™ LC/MS) were acquired from Fisher Scientific (Fair Lawn, NJ, USA), and trypan blue and LC-MS grade acetonitrile from Sigma-Aldrich® (St. Louis, MT, USA). Ten-donor-pooled cryopreserved human hepatocytes, InVitroGRO™ CP Medium, and Krebs-Henseleit buffer (KHB) were obtained from BioreclamationIVT (Baltimore, MD, USA).
Incubation with Hepatocytes
Hepatocytes were thawed at 37°C and gently rinsed twice with CP Medium and once with KHB, then centrifuged at room temperature, 100×g, for 5 min. The pellet was solubilized in 2 mL KHB. Cell viability was measured with trypan blue exclusion dye to allow adjusting the buffer volume to a 2 × 106 viable cells/mL concentration. Two hundred fifty microliters suspension was gently mixed with 20 μmol/L ADB-CHMINACA in KHB with 0.7% methanol (10 μmol/L final concentration) and incubated for 0 and 3 h at 37°C in a Forma™ Steri-Cycle™ CO2 incubator (Thermo Scientific; Fremont, CA, USA). Metabolic reactions were quenched with 500 μL acetonitrile. Samples were centrifuged at 4°C, 15,000×g, for 5 min, and stored at −80°C until analysis. A control sample with diclofenac was incubated under the same conditions and 4′-hydroxydiclofenac and acyl-β-D-glucuronide diclofenac were monitored to ensure hepatocyte metabolic activity. Simultaneously, a negative control with ADB-CHMINACA standard without hepatocytes was incubated for 3 h to control for interferences and non-enzymatic reactions.
Sample Preparation
Incubation Samples
Samples were thawed at room temperature, vortex mixed, and centrifuged at 4°C, 15,000×g, for 5 min. One hundred microliter supernatants was transferred to disposable plastic microtubes with 100 μL acetonitrile, vortex mixed, and centrifuged at 4°C, 15,000×g, for 5 min. Supernatants were transferred to conical glass tubes and evaporated to dryness under nitrogen at 40°C in a TurboVap® LV evaporator (Zymark; Hopkinton, MA, USA). Residues were reconstituted with 150 μL mobile phase A/B 80:20 (v/v), centrifuged at 4°C, 15,000×g for 5 min, and supernatants transferred into autosampler vials with glass inserts. Fifteen microliters was injected onto the chromatographic system.
Reference Standards
Ten microliters ADB-CHMINACA M1, M2, M3, M7, M10, and M11 standards was vortex mixed with 990 μL mobile phase A/B 80:20 (v/v) in individual glass autosampler vials. Fifteen microliters was injected onto the chromatographic system.
LC-HRMS Parameters
LC-MS analysis was performed on a Q Exactive™ Plus mass spectrometer equipped with a heated electrospray in positive-ion mode (Thermo Scientific) and coupled with an Ultimate™ 3000 LC system (Dionex; Sunnyvale, CA, USA). Data were acquired with Xcalibur software (v. 3.0.63, Thermo Scientific).
LC Parameters
Separation was achieved on a 100 × 2.1 mm, 3 μm, Ultra Biphenyl column (Restek®; Bellefonte, PA, USA) and identically packed guard cartridge (10 × 2.1 mm). Elution was performed at 30°C with 0.1% formic acid in water (A) and 0.1% formic acid in acetonitrile (B) at a 0.5 mL/min flow rate. Initial conditions (20% B) were maintained for 0.5 min, then %B was increased to 95% within 10.5 min and held for 2 min; the column was re-equilibrated within 2 min (15 min run time).
HRMS Parameters
Ionization source parameters and collision energies were optimized on parent signal by post-column infusion of ADB-CHMINACA standard in A/B 50:50 (v/v) at 0.5 mL/min. Source parameters were as follows: spray voltage, 4 kV; sheath gas flow rate, 40 a.u.; auxiliary gas flow rate, 5 a.u.; sweep gas flow rate, 2 a.u.; S-lens radio frequency level, 50 a.u.; auxiliary gas heater temperature, 400°C; capillary temperature, 300°C. Each sample was injected twice with a different acquisition method; pros and cons of the two methods were discussed previously (34). On first injection, data were acquired in full-scan MS/data-dependent MS/MS mode (FullMS/ddMS2) with an inclusion list of expected metabolites based on the literature (24,25,34–36) (Table I). FullMS settings were as follows: resolution (full width at half-maximum at m/z 200), 70,000; scan range, m/z 150–650; automatic gain control (AGC) target, 1.0 × 106; and maximum injection time (IT), 200 ms. ddMS2 settings were as follows: TopN, 5; isolation window, m/z 1.5; intensity threshold, 2.0 × 104; resolution, 17,500; AGC target, 1.0 × 105; maximum IT, 50 ms; stepped collision energy, 10, 40, and 60%; apex triggering, 3 to 6 s; and dynamic exclusion, 2 s. On a second injection, data were acquired in full-scan MS/all-ion fragmentation/data-dependent MS/MS mode (FullMS/AIF/ddMS2) with a list of neutral losses based on ADB-CHMINACA fragmentation pattern; sulfate and glucuronide losses were included in the case of phase II metabolism (Table I). FullMS and ddMS2 settings were the same as described above. AIF settings were as follows: TopN, 5; isolation window, m/z 150–650; intensity threshold, 2.0 × 104; resolution, 17,500; AGC target, 1.0 × 105; maximum IT, 50 ms; and stepped collision energy, 10, 40, and 60%.
Metabolite Identification
Raw data from samples and controls were processed with Compound Discoverer™ (Thermo Scientific) and compiled in one analysis. Retention times (RT) of chromatographic peaks present in several data files were aligned to facilitate identification. RT alignment used the adaptive curve model with a 5-ppm mass tolerance and a 0.1-min maximum time shift. Peaks were automatically compared to a list of potential metabolites generated by a combination of suggested biotransformations (Table II). Mass tolerance for MS identification was 5 ppm, minimum MS peak intensity was 2.5 × 102, and intensity tolerance for isotope search was 50%. Percentage match between experimental and theoretical MS/MS spectra was calculated based on presence/absence of theoretical fragments in the experimental MS/MS spectra. Mass tolerance for fragment identification was 10 ppm; signal/noise ratio threshold was 3.
RESULTS AND DISCUSSION
ADB-CHMINACA Fragmentation
ADB-CHMINACA eluted at 7.16 min ([M+H]+, m/z 371.2438) and yielded characteristic fragments shown in Fig. 3a. As observed in structural analogs ADB-FUBINACA (36), ADB-PINACA, and 5F-ADB-PINACA (34), major fragments were produced by neutral loss of amine (m/z 354.2169), terminal carboxamide (m/z 326.2219), dimethylbutanamide (m/z 259.1439), and aminodimethylbutanamide (m/z 241.1330) groups. Fragment m/z 97.1011 was produced by cyclohexylmethylium formation, as observed in AB-CHMINACA (24) and MDMD-CHMICA (35) fragmentation. Subsequent cyclohexylmethyl loss from m/z 241.1330 and 259.1439 generated fragments m/z 145.0395 and 163.0499, respectively, characteristic of the indazole core (34,36). ADB-CHMINACA fragments m/z 145, 241, and 326 were previously identified by MS/MS after ESI (12,15) and MS after electronic impact ionization (12).
ADB-CHMINACA Metabolites in Human Hepatocytes
ADB-CHMINACA MS peak area signal decreased to 15% after 3 h incubation with human hepatocytes (extracted ion chromatogram). Ten different metabolites were identified at T3h and listed from A1 to A10 by ascending retention time (RT) (Fig. 4). Metabolites were absent from control samples, excluding interferences and non-enzymatic reactions. Metabolic reactions, accurate mass molecular ion, RT, elemental composition, nominal mass product ions, and MS peak areas of ADB-CHMINACA and metabolites are reported in Table III. MS/MS spectra and fragmentation patterns are shown in Fig. 3 and supplementary electronic data S1. The dominant reaction was the hydroxylation of the cyclohexylmethyl tail (A4, A5, A6, A8, A9) as observed in structural analogs AB-CHMINACA (24) and MDMB-CHMICA (25,35) (although the hydrolysis of the terminal ester also was a major transformation of MDMB-CHMICA). Cyclohexylmethyl ketolization (A7), tert-butyl hydroxylation (A10), and dihydroxylation (A1, A2, A3) also occurred to a lesser extent. No other transformation was detected at the indazole core, the carboxamide linker, or the dimethylbutanamide side chain, as opposed to analogs ADB-FUBINACA (36), ADB-PINACA, and 5F-ADB-PINACA (34), confirming the cyclohexylmethyl tail as an extensively metabolized group that is the primary focus of ADB-CHMINACA biotransformation. We currently have no explanation for the absence of hydroxyindazole metabolites. No phase II reaction was detected, although glucuronidation is a common transformation for SC in humans (29–32). In AB-CHMINACA metabolism, glucuronidation occurred at the side chain after amide hydrolysis. However, no phase II transformation was detected after hydroxylation of the L-valinamide side chain or the cyclohexylmethyl tail (24). Similarly, no phase II reaction was detected after hydroxylation of the dimethylbutanamide side chain of ADB-CHMINACA structural analogs (34,36). The absence of phase II biotransformation in ADB-CHMINACA metabolism is consistent with these observations, as oxidations at the dimethylbutanamide side chain and the cyclohexylmethyl tail were the only detectable phase I reactions. The size of the cyclohexylmethyl structure may be a hindrance for enzymes that glucuronidate the hydroxyl groups. Other phase II reactions, such as sulfations and cysteine conjugations, are rarely reported for synthetic cannabinoids in humans. We propose a metabolic pathway for ADB-CHMINACA in Fig. 5.
Cyclohexylmethyl Hydroxylation and Further Ketolization
Hydroxylation (+O) occurred in A4, A5, A6, A8, and A9, as indicated by the +15.9948 Da mass shift from parent. The five MS/MS spectra displayed the same fragments with characteristic neutral loss of amine (m/z 370.2119), carboxamide (m/z 342.2172), dimethylbutanamide (m/z 275.1386), and aminodimethylbutanamide (m/z 257.1282) groups indicating that neither the carboxamide linker nor the dimethylbutanamide side chain carried the transformation; the reaction did not occur either at the indazole core, as indicated by fragments m/z 145.0395 and 163.0500 also observed in parent, suggesting that hydroxylation occurred at the cyclohexylmethyl tail. Fragment m/z 95.0855 produced by the hydroxylated cyclohexylmethylium ion followed by water loss confirmed the cyclohexylmethyl group as the site of reaction. Interestingly, fragments m/z 239.1175 (water loss from m/z 257.1282), 257.1282, and 275.1386 presented with different intensity ratios in A4, A5, A6, A8, and A9: m/z 239.1175 signal was high in A5 but low or absent from other metabolites; m/z 257.1282 had the most intense signal in A4 and A9, while m/z 275.1386 was dominating in A6 and A8. As previously mentioned, cyclohexylmethyl hydroxylation was expected, as it is a major metabolic reaction of structural analogs (24,25,35). However, the MS/MS spectra do not allow accurate location of the hydroxyl group on the tail. The transformation may be generated by cytochrome P450 3A4 (CYP3) as it was identified as the main metabolic enzyme responsible for cyclohexylmethyl hydroxylation in AB-CHMINACA in vitro metabolism (24).
A7 was a minor metabolite produced by oxidation or hydroxylation followed by dehydrogenation, resulting in a ketone formation (+O –2H). Neutral loss of amine (m/z 368.1963), carboxamide (m/z 340.2023), and aminodimethylbutanamide (m/z 255.1125) groups and indazole fragments m/z 145.0396 and 163.0504 indicated a ketolization at the cyclohexylmethyl tail. Cyclohexylmethyl ketolization was previously reported in MDMB-CHMICA in vitro and in vivo metabolism with a low signal intensity (25). It was not detected in AB-CHMINACA metabolism (24).
tert-Butyl Hydroxylation
A10 also was formed by hydroxylation (+O), as suggested by a +15.9952 Da mass shift from ADB-CHMINACA. However, A10 shared most of its fragments with parent such as m/z 97.1015 (cyclohexylmethyl tail), 145.0394 and 163.0498 (indazole core), and 241.1331 and 259.1443 (cyclohexylmethyl + indazole + carboxamide linker) designating the tert-butyl group of the side chain as the site of attack. It is noteworthy that the carboxamide loss was followed by subsequent hydroxymethyl loss (m/z 312.2064) and did not appear in A10 spectrum, as observed in the fragmentation of ADB-FUBINACA (36), ADB-PINACA, and 5F-ADB-PINACA (34) metabolites with the same transformation. tert-Butyl hydroxylation also occurred in MDMB-CHMICA (25,35) and AB-CHMINACA (24) metabolism, in which CYP3A3 was identified as the main enzyme responsible for the reaction (24).
Dihydroxylation
Second generation metabolites A1, A2, and A3 were produced by dihydroxylation (+2O), as indicated by a 31.9900-Da mass shift from parent. A1 and A3 followed a similar fragmentation pattern to A4, A5, A6, A8, and A9, yielding fragments by neutral loss of amine (m/z 386.2072), carboxamide (m/z 358.2115), dimethylbutanamide (m/z 291.1330), and aminodimethylbutanamide (m/z 273.1228) groups and characteristic fragments from the indazole core (m/z 145.0394 and 163.0500). This pattern suggested a dihydroxylation at the cyclohexylmethyl tail. Fragment m/z 93.0698 generated by dihydroxylation of the cyclohexylmethylium ion followed by two water losses further designated the tail as the site of attack. The signal of the aminodimethylbutanamide loss dominated in the A3 spectrum, as observed in major metabolites A4 and A9. However, this fragmentation was followed by two successive water losses (m/z 255.1122 and 237.1010) in A1.
A2 was dihydroxylated at the cyclohexylmethyl tail and at the tert-butyl group of the dimethylbutanamide side chain. As observed in A4, A5, A6, A8, and A9 MS/MS spectra, fragments m/z 95.0855, 145.0396, 163.0501, 239.1183, 257.1283, and 275.1385 indicated a hydroxycyclohexylmethyl metabolite. As shown for A10, amine (m/z 386.2069) and carboxamide (m/z 368.1961) neutral losses and subsequent hydroxymethyl loss (m/z 328.2008) indicated that the second hydroxylation occurred at the tert-butyl chain.
Comparison to Reference Standards
Analytical standards of six different theoretical metabolites were injected to allow correct assignment of positional isomers in hepatocyte incubations (Fig. 2). ADB-CHMINACA M2 (RT, 9.68 min), M3 (two stereoisomers at 5.09 and 5.20 min), M7 (6.46 min), and M10 (8.00 min) were not detected in the incubations. All four compounds were hydrolyzed at the terminal carboxamide, which is a common metabolic transformation for synthetic cannabinoids with a dimethylbutanamide chain (34,36), and even more common for their desmethyl analogs (29,30). Amide hydrolysis also was detected in AB-CHMINACA metabolism but as a minor transformation, since the cyclohexylmethyl tail appeared to be a more critical site of transformation. The absence of M2, M3, M7, and M10 in hepatocyte incubations with ADB-CHMINACA was therefore not surprising. M1 matched A4 RT and MS/MS spectrum (Fig. 6) implying that A4 hydroxylation occurred at C4″ of the cyclohexyl ring. Similarly, M11 matched A2 RT and fragmentation (Fig. 6), demonstrating that hydroxylation occurred at C4″ of the cyclohexyl ring and at C3′ of the tert-butyl chain. Consequently, although it was not confirmed with a reference standard, A10 also may be hydroxylated at C3′ of the tert-butyl chain. The exact position of the transformations occurring for other metabolites cannot be determined with LC-HRMS/MS data and commercially available standards alone. Nuclear magnetic resonance could better identify the exact position of these hydroxylations.
Extracted ion chromatogram and tandem mass spectrometry spectrum of M11 (ADB-CHMINACA 3′-hydroxypropyl-4″-hydroxycyclohexyl) (a and b, respectively) and M1 (ADB-CHMINACA 4″-hydroxycyclohexyl) (c and d, respectively) reference standards versus ADB-CHMINACA metabolites identified after 3 h hepatocyte incubation. Mass tolerance, 5 ppm; RT, retention time
Optimal Targets for ADB-CHMINACA Intake
We recommend major metabolites A4 (ADB-CHMINACA 4″-hydroxycyclohexyl), A6 (ADB-CHMINACA hydroxycyclohexylmethyl), and A9 (ADB-CHMINACA hydroxycyclohexylmethyl) as biological markers for ADB-CHMINACA consumption. All three compounds are the result of a simple hydroxylation and carry ADB-CHMINACA indazole core substituted with a cyclohexylmethyl tail and a dimethylbutanamide side chain. Consequently, all three compounds specifically identify ADB-CHMINACA intake and none were reported as a metabolite from closely related analogs. MS/MS analysis does not allow precise location of A6 and A9 hydroxylation and requires comparison with reference standards that are not commercially available. These data inform manufacturers on their synthesis efforts to provide suitable standards. Currently, A4 is the most convenient target for ADB-CHMINACA intake, as its standard is available for purchase. Remarkably, although it is recommended for synthetic cannabinoids analysis (25,37–41), for ADB-CHMINACA identification, it is not necessary to hydrolyze urine samples as no phase II ADB-CHMINACA metabolite was detected.
In previous experiments involving human metabolism, hepatocyte incubations appeared successful to predict in vivo urinary metabolites (29–32,42,43). Ideally, a confirmation of our results with urine specimens from authentic cases of ADB-CHMINACA intake would be advantageous (33); however, despite our efforts, we were unable to obtain such specimens. It is noteworthy that structural analogs AB-CHMINACA (24) and MDMB-CHMICA (25,35) in vivo metabolism corroborates our results: the most intense urinary metabolites are mono-hydroxylated at the cyclohexylmethyl tail, as observed in ADB-CHMINACA hepatocyte incubations.
CONCLUSIONS
For the first time, we determined ADB-CHMINACA (MAB-CHMINACA) human metabolism using hepatocyte incubations and LC-HRMS analysis with metabolite identification data-mining software. We detected 10 different phase I metabolites, most of the transformations occurring at the cyclohexylmethyl tail of the compound. We suggest A9 (ADB-CHMINACA hydroxycyclohexylmethyl), A4 (ADB-CHMINACA 4″-hydroxycyclohexyl), and A6 (ADB-CHMINACA hydroxycyclohexylmethyl) as targets for documenting ADB-CHMINACA intake in clinical and forensic specimens. These data also will provide support for analytical standard manufacturers to produce suitable metabolites for further pharmacodynamics and pharmacokinetic studies.
REFERENCES
Gurney SM, Scott KS, Kacinko SL, Presley BC, Logan BK. Pharmacology, toxicology, and adverse effects of synthetic cannabinoid drugs. Forensic Sci Rev. 2014;26(1):53–78.
Banister SD, Stuart J, Kevin RC, Edington A, Longworth M, Wilkinson SM, et al. Effects of bioisosteric fluorine in synthetic cannabinoid designer drugs JWH-018, AM-2201, UR-144, XLR-11, PB-22, 5F-PB-22, APICA, and STS-135. ACS Chem Neurosci. 2015;6(8):1445–58. doi:10.1021/acschemneuro.5b00107.
Banister SD, Moir M, Stuart J, Kevin RC, Wood KE, Longworth M, et al. Pharmacology of indole and indazole synthetic cannabinoid designer drugs AB-FUBINACA, ADB-FUBINACA, AB-PINACA, ADB-PINACA, 5F-AB-PINACA, 5F-ADB-PINACA, ADBICA, and 5F-ADBICA. ACS Chem Neurosci. 2015;6(9):1546–59. doi:10.1021/acschemneuro.5b00112.
Banister SD, Longworth M, Kevin R, Sachdev S, Santiago M, Stuart J, et al. Pharmacology of valinate and tert-leucinate synthetic cannabinoids 5F-AMBICA, 5F-AMB, 5F-ADB, AMB-FUBINACA, MDMB-FUBINACA, MDMB-CHMICA, and their analogues. ACS Chem Neurosci. 2016;7(9):1241–54. doi:10.1021/acschemneuro.6b00137.
Auwarter V, Dresen S, Weinmann W, Muller M, Putz M, Ferreiros N. ‘Spice’ and other herbal blends: harmless incense or cannabinoid designer drugs? J Mass Spectrom: JMS. 2009;44(5):832–7. doi:10.1002/jms.1558.
Uchiyama N, Asakawa K, Kikura-Hanajiri R, Tsutsumi T, Hakamatsuka T. A new pyrazole-carboxamide type synthetic cannabinoid AB-CHFUPYCA [N-(1-amino-3-methyl-1-oxobutan-2-yl)-1-(cyclohexylmethyl)-3-(4-fluorophenyl)-1H-pyrazole-5-carboxamide] identified in illegal products. Forensic Toxicol. 2015;33(2):367–73. doi:10.1007/s11419-015-0283-8.
Trecki J, Gerona RR, Schwartz MD. Synthetic cannabinoid-related illnesses and deaths. N Engl J Med. 2015;373(2):103–7. doi:10.1056/NEJMp1505328.
Law R, Schier J, Martin C, Chang A, Wolkin A, Centers for Disease Control and Prevention. Notes from the field: increase in reported adverse health effects related to synthetic cannabinoid use—United States, January–May 2015. MMWR Morb Mortal Wkly Rep. 2015;64(22):618–9.
Kamijo Y, Takai M, Fujita Y, Sakamoto T. A multicenter retrospective survey of poisoning after consumption of products containing novel psychoactive substances from, to 2014 in Japan. Am J Drug Alcohol Abuse. 2013;2016:1–7. doi:10.1080/00952990.2016.1176177.
European Monitoring Centre for Drugs and Drug Addiction. Perspectives on drugs—Synthetic cannabinoids in Europe. 2016.
Buchler IP, Hayes MJ, Hegde SG, Hockerman SL, Jones DE, Kortum SW, et al. Indazole derivatives. Google Patents; 2009.
Wurita A, Hasegawa K, Minakata K, Gonmori K, Nozawa H, Yamagishi I, et al. Identification and quantitation of 5-fluoro-ADB-PINACA and MAB-CHMINACA in dubious herbal products. Forensic Toxicol. 2015;33(2):213–20. doi:10.1007/s11419-015-0264-y.
Adamowicz P, Gieron J. Acute intoxication of four individuals following use of the synthetic cannabinoid MAB-CHMINACA. Clin Toxicol. 2016;54(8):650–4. doi:10.1080/15563650.2016.1190016.
Hasegawa K, Wurita A, Minakata K, Gonmori K, Yamagishi I, Nozawa H, et al. Identification and quantitation of 5-fluoro-ADB, one of the most dangerous synthetic cannabinoids, in the stomach contents and solid tissues of a human cadaver and in some herbal products. Forensic Toxicol. 2015;33(1):112–21. doi:10.1007/s11419-014-0259-0.
Hasegawa K, Wurita A, Minakata K, Gonmori K, Nozawa H, Yamagishi I, et al. Postmortem distribution of MAB-CHMINACA in body fluids and solid tissues of a human cadaver. Forensic Toxicol. 2015;33(2):380–7. doi:10.1007/s11419-015-0272-y.
Drug Enforcement Administration, Department of Justice. Schedules of controlled substances: temporary placement of the synthetic cannabinoid MAB-CHMINACA into schedule I. Final order. Fed Regist. 2016;81(24):6171–5.
National Institute of Health Sciences. Data search system for new psychoactive substances. 2016 [cited 2016 November]; Available from: http://npsdb.nihs.go.jp/Search/Search/AllCompounds_e.aspx.
Central Narcotics Bureau. Misuse of drugs act. CNB news release. 2015.
Adamowicz P, Zuba D, Sekula K. Analysis of UR-144 and its pyrolysis product in blood and their metabolites in urine. Forensic Sci Int. 2013;233(1–3):320–7. doi:10.1016/j.forsciint.2013.10.005.
Jang M, Yang W, Choi H, Chang H, Lee S, Kim E, et al. Monitoring of urinary metabolites of JWH-018 and JWH-073 in legal cases. Forensic Sci Int. 2013;231(1–3):13–9. doi:10.1016/j.forsciint.2013.03.053.
Jang M, Yang W, Shin I, Choi H, Chang H, Kim E. Determination of AM-2201 metabolites in urine and comparison with JWH-018 abuse. Int J Legal Med. 2014;128(2):285–94. doi:10.1007/s00414-013-0884-x.
Bertol E, Vaiano F, Di Milia MG, Mari F. In vivo detection of the new psychoactive substance AM-694 and its metabolites. Forensic Sci Int. 2015;256:21–7. doi:10.1016/j.forsciint.2015.07.018.
Jang M, Kim IS, Park YN, Kim J, Han I, Baeck S, et al. Determination of urinary metabolites of XLR-11 by liquid chromatography-quadrupole time-of-flight mass spectrometry. Anal Bioanal Chem. 2016;408(2):503–16. doi:10.1007/s00216-015-9116-1.
Erratico C, Negreira N, Norouzizadeh H, Covaci A, Neels H, Maudens K, et al. In vitro and in vivo human metabolism of the synthetic cannabinoid AB-CHMINACA. Drug Test Anal. 2015;7(10):866–76. doi:10.1002/dta.1796.
Franz F, Angerer V, Moosmann B, Auwarter V. Phase I metabolism of the highly potent synthetic cannabinoid MDMB-CHMICA and detection in human urine samples. Drug Test Anal. 2016. doi:10.1002/dta.2049.
Holm NB, Pedersen AJ, Dalsgaard PW, Linnet K. Metabolites of 5F-AKB-48, a synthetic cannabinoid receptor agonist, identified in human urine and liver microsomal preparations using liquid chromatography high-resolution mass spectrometry. Drug Test Anal. 2015;7(3):199–206. doi:10.1002/dta.1663.
Sobolevsky T, Prasolov I, Rodchenkov G. Detection of urinary metabolites of AM-2201 and UR-144, two novel synthetic cannabinoids. Drug Test Anal. 2012;4(10):745–53. doi:10.1002/dta.1418.
Hutter M, Moosmann B, Kneisel S, Auwarter V. Characteristics of the designer drug and synthetic cannabinoid receptor agonist AM-2201 regarding its chemistry and metabolism. J Mass Spectrom: JMS. 2013;48(7):885–94. doi:10.1002/jms.3229.
Wohlfarth A, Castaneto MS, Zhu M, Pang S, Scheidweiler KB, Kronstrand R, et al. Pentylindole/pentylindazole synthetic cannabinoids and their 5-fluoro analogs produce different primary metabolites: metabolite profiling for AB-PINACA and 5F-AB-PINACA. AAPS J. 2015;17(3):660–77. doi:10.1208/s12248-015-9721-0.
Castaneto M, Wohlfarth A, Pang S, Zhu M, Scheidweiler KB, Kronstrand R, et al. Identification of AB-FUBINACA metabolites in human hepatocytes and urine using high-resolution mass spectrometry. Forensic Toxicol. 2015;33(2):295–310. doi:10.1007/s11419-015-0275-8.
Diao X, Scheidweiler KB, Wohlfarth A, Pang S, Kronstrand R, Huestis MA. In vitro and in vivo human metabolism of synthetic cannabinoids FDU-PB-22 and FUB-PB-22. AAPS J. 2016;18(2):455–64. doi:10.1208/s12248-016-9867-4.
Diao X, Carlier J, Zhu M, Pang S, Kronstrand R, Scheidweiler KB, et al. In vitro and in vivo human metabolism of a new synthetic cannabinoid NM-2201 (CBL-2201). Forensic Toxicol. 2017; 35(1):20–32. doi:10.1007/s11419-016-0326-9.
Diao X, Huestis MA. Approaches, challenges and advances in metabolism of new synthetic cannabinoids and identification of optimal urinary marker metabolites. Clin Pharmacol Ther. 2016. doi:10.1002/cpt.534.
Carlier J, Diao X, Scheidweiler KB, Huestis MA. Distinguishing intake of new synthetic cannabinoids, ADB-PINACA and 5F-ADB-PINACA, with human hepatocyte metabolites and high resolution mass spectrometry. Clin Chem. 2017. doi:10.1373/clinchem.2016.267575.
Grigoryev A, Kavanagh P, Pechnikov A. Human urinary metabolite pattern of a new synthetic cannabimimetic, methyl 2-(1-(cyclohexylmethyl)-1H-indole-3-carboxamido)-3,3-dimethylbutanoate. Forensic Toxicol. 2016;34(2):316–28. doi:10.1007/s11419-016-0319-8.
Carlier J, Diao X, Wohlfarth A, Scheidweiler KB, Huestis MA. In vitro metabolic profiling of ADB-FUBINACA, a new synthetic cannabinoid. Curr Neuropharmacol. 2017. doi:10.2174/1570159X15666161108123419.
Freijo Jr TD, Harris SE, Kala SV. A rapid quantitative method for the analysis of synthetic cannabinoids by liquid chromatography-tandem mass spectrometry. J Anal Toxicol. 2014;38(8):466–78. doi:10.1093/jat/bku092.
Scheidweiler KB, Jarvis MJ, Huestis MA. Nontargeted SWATH acquisition for identifying 47 synthetic cannabinoid metabolites in human urine by liquid chromatography-high-resolution tandem mass spectrometry. Anal Bioanal Chem. 2015;407(3):883–97. doi:10.1007/s00216-014-8118-8.
Berg T, Kaur L, Risnes A, Havig SM, Karinen R. Determination of a selection of synthetic cannabinoids and metabolites in urine by UHPSFC-MS/MS and by UHPLC-MS/MS. Drug Test Anal. 2016;8(7):708–22. doi:10.1002/dta.1844.
Neifeld JR, Regester LE, Holler JM, Vorce SP, Magluilo Jr J, Ramos G, et al. Ultrafast screening of synthetic cannabinoids and synthetic cathinones in urine by RapidFire-tandem mass spectrometry. J Anal Toxicol. 2016;40(5):379–87. doi:10.1093/jat/bkw025.
Borg D, Tverdovsky A, Stripp R. A fast and comprehensive analysis of 32 synthetic cannabinoids using Agilent triple quadrupole LC-MS-MS. J Anal Toxicol. 2016. doi:10.1093/jat/bkw104.
Swortwood MJ, Carlier J, Ellefsen KN, Wohlfarth A, Diao X, Concheiro-Guisan M, et al. In vitro, in vivo and in silico metabolic profiling of alpha-pyrrolidinopentiothiophenone, a novel thiophene stimulant. Bioanalysis. 2016;8(1):65–82. doi:10.4155/bio.15.237.
Swortwood MJ, Ellefsen KN, Wohlfarth A, Diao X, Concheiro-Guisan M, Kronstrand R, et al. First metabolic profile of PV8, a novel synthetic cathinone, in human hepatocytes and urine by high-resolution mass spectrometry. Anal Bioanal Chem. 2016;408(18):4845–56. doi:10.1007/s00216-016-9599-4.
Acknowledgements
The authors would like to thank Tim Moeller from BioreclamationIVT for his assistance with the incubations, and Caroline Ding, Helen Sun, and Thermo Scientific for providing LC-HRMS instrumentation, Compound Discoverer® software and training via a National Institute on Drug Abuse Materials Transfer Agreement. This research was supported by the Intramural Research Program of the National Institute on Drug Abuse, National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Additional information
Jeremy Carlier and Xingxing Diao contributed equally to this work.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 509 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Carlier, J., Diao, X., Sempio, C. et al. Identification of New Synthetic Cannabinoid ADB-CHMINACA (MAB-CHMINACA) Metabolites in Human Hepatocytes. AAPS J 19, 568–577 (2017). https://doi.org/10.1208/s12248-016-0037-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1208/s12248-016-0037-5