Abstract
Providing an accurate count of total leukocytes and specific subsets (such as T-cells and B-cells) within small amounts of whole blood is a rather challenging ordeal due to the lack of techniques that enable the separation of leukocytes from a limited amount of whole blood. In a previous study we designed a microfluidic chip utilizing a micropillar array to isolate T-cells and B-cells from the sub-microliter of whole blood. Due to the variability of cells in size, morphology and color intensity, a Histogram of Oriented Gradients (HOG) based Support Vector Machine (SVM) classifier was proposed with an average accuracy of 94%, specificity of 99% and sensitivity of 90%. The HOG can separate the cells from the background with a high accuracy rate however, some noise is similar in shape and size to the actual cells and this results in misclassification. To alleviate this situation, in this study a convolutional neural network is trained and used to distinguish T-cells and B-cells with an accuracy rate of 98%, a specificity of 99% and a sensitivity of 97%. We also propose an HOG feature based SVM classifier to preselect the detection windows to accelerate the detection to process images in less than 10 min. The proposed on-chip cell detecting and counting method will be useful for numerous applications in diagnosis and for monitoring diseases.
Similar content being viewed by others
Introduction
Over the years with the development of new techniques and emerging technologies, cell analysis has evolved with regards to speed, sensitivity, spatial resolution, cost, etc. Every form of cell analysis represents a compromise. Three dimensional (3D) optical scanning microscopy can achieve great spatial resolution, but scanning takes time [1]. A standard optical microscopy has a variety of modes (transmitted light, scattered light, fluorescence, phase contrast, etc.), each of which provides distinct and complementary information about the cell [2] and is suitable to obtain a two-dimensional (2D) image of the cell. A flow cytometry measures fluorescence intensity and scattering from cells suspended in flow. Only a limited number of signals for each cell is available depending on the optical system, and spatial resolution is lost entirely but it is possible to measure thousands of cells in a second [3].
The form of cell analysis and the method to process the data depends on the application and its time requirements. In case of real time detection and sorting of cells as in flow cytometry, only a fraction of a millisecond per cell is allowed. On the other hand, for confocal microscopy, 3D scans can be analyzed in detail offline. Due to the sheer amount of data from 3D scans or images, storing and processing raw cell data is not always possible and various features were extracted to simplify cell analysis [4]. Extracting simple features such as the size or circularity of a cell is significant in many cases, and the 2D image of the cell obtained via fluorescence optical microscopy will ease the processing and sharing of data. If the feature extracted from the 2D image is to be used for the detection and classification of cells, caution is required to prevent the loss of important information that might be included in an image. Hence, the extraction of features that contain most of the related information was researched and used for applications that require the detection and classification of cells [5]. With the improvement of automation in image analysis and the ease of access to available algorithms, creating large datasets of cells and cell features became possible [6].
There is a tremendous need for an automated, portable point-of-care blood cell counter that could yield results in a matter of minutes from a drop of blood without any trained professionals to operate the instrument. Microfluidic based devices are a proven technology for cellular handling and are easily integrated with the fluorescence microscopy to obtain 2D images of cells. Cell counting methods using microfluidic devices are promising substitutes for conventional flow cytometry systems since they allow for faster analysis, reduced sample volume, and are portable, inexpensive and disposable. For the isolation and detection of T-cells and B-cells from whole blood, we designed a microfluidic chip and implemented a machine learning algorithm with an appropriate feature descriptor [7, 8]. Due to the variability of cells in size, morphology and color intensity, a Histogram of Oriented Gradients (HOG) descriptor, which is a popular choice for object detection among the computer vision community due to its robustness, is used [9]. The HOG encompasses the shape of the cells while operating in localized regions and remaining invariant to geometric and photometric transformations. In other words, the HOG can work even when there are fluctuations in the illumination and shapes of the cells to some extent due to its isolation system design. The HOG can distinguish the cells from the background with a high accuracy rate. However, some noise is similar in shape and size to the actual cells, hence there is a possibility of misclassification. In our previous work, we chose to have a lower rate of misclassification and forwent a higher detection rate. It might be possible to incorporate features other than shape to increase the accuracy but deciding on what features to use and how to combine them remains an issue. A key approach is to employ machine learning methods to automate feature extraction, and directly use raw data for classification [10, 11]. Convolutional neural networks (CNNs), train directly on the labeled raw data, and learn the features to be extracted from the images automatically. CNNs are widely used for applications that require object recognition and computer vision such as self-driving cars and face-recognition [12, 13]. CNNs are also used in applications pertaining to cells [14,15,16]. Despite the attractive qualities of CNNs, they were not widely used for cell detection and classification until recent years, since they require powerful GPUs and large datasets to train. Current GPUs optimized for training CNNs and large datasets containing millions of labeled images such as ImageNet only became available in recent years [17].
In this study we propose to use a CNN to detect T-cells and B-cells from 2D fluorescence microscopy images. A pretrained convolutional neural network (CNN) is fine tuned to detect T-cells and B-cells from the background. A preselection method for detection windows using the HOG feature based SVM classifier is introduced to accelerate the detection process. In the testing phase, images are scanned with a sliding window and each detection window in the image is classified by the trained CNN. The comparison of the performance between the trained CNN and our previous work which utilized the HOG feature based SVM classifier is demonstrated. Furthermore, the impact of the preselection method in performance is also discussed.
Sample preparation and experiment
Human blood samples were collected from healthy donors at the National Hospital Organization Nagoya Medical Center, and all participants provided written informed consent to participate in the study. This study was approved by the institutional review board of the National Hospital Organization Nagoya Medical Center. For the detection of T-cells and B-cells, the blood sample was mixed with a two-color direct immunofluorescence reagent (BD Simultest™ CD3-FITC(Ex 494 nm, Em 520 nm)/CD19-PE(Ex 496 nm, Em 578 nm), BD Bioscience, San Jose, CA) and incubated at room temperature for 15 min.
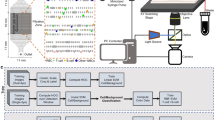
Figure 1 demonstrates the overview of the experiment setup. We used a pillar-based microfluidic chip that is designed to isolate leukocytes from peripheral blood with high efficiency and without clogging [7]. This microfluidic chip uses gradual size-based filtration with gap sizes ranging from 3–6 μm and 8–15 μm. The larger micro-pillar gaps are intended to capture bigger cells such as lymphocytes, which are mostly spherical in shape and vary from 7 μm to 30 μm in diameter. We introduced 1 μL peripheral blood into the chip and collected the fluid from the chip into a tube. After the entire blood sample flowed through the chip, a 30 μL sheath liquid (PBS with 5 mM EDTA) was introduced to remove non-trapped cells. Following the sample introduction, fluorescence of T-cells and B-cells were excited (488 nm) and the filtration zone on the chip was scanned automatically. The images were captured by a CMOS-camera (ASI178 MC, Zhen Wang Optical Company, China, 3096 × 2080, pixel size 2.4 µm) installed on the eyepiece body tube of the microscope. The whole chip was scanned frame by frame, and after an image was acquired the chip was moved using motorized stages. The total area of the filtration zone can be covered by 420 images and the scanning time is 14 min.
The dataset
There is no standard dataset for fluorescent cell images, so we created a custom dataset in our previous work [8]. Previously a total of 6200 cells and 35,000 background images were gathered semi-automatically. We augmented this dataset by using the horizontal reflections of the cell images, then a random subset of background images were chosen for the sake of a better-balanced dataset.
Figure 2 illustrates the steps of creating the final dataset and the training of CNNs. The emission spectrum crosstalk between the dyes prevents us from manually annotating the datasets to be used as training data. To alleviate the influence of fluorescence emission spectrum crosstalk, we performed the same experiment by staining the cells with a single dye. Cell images from these experiments were automatically extracted by the proposed CNN and added to the final dataset. These cells were used to calibrate the color distribution and a separate CNN—which will be further elaborated on in the following chapters—was trained to distinguish T-cells from B-cells. Then using the trained CNN, previously un-annotated cells are annotated. The dataset for training the CNN is finalized with three labels; T cell, B-cell and background as Fig. 3 demonstrates. The dataset for testing is kept as it is, for the sake of performance comparison.
Flow chart of training AlexCAN. Firstly, a CNN < Cell/Background > is re-trained using the original dataset with the labels of cell and background. Then, training images dyed with a single dye are exhaustively searched, and the trained CNN < Cell/Background > is used to create a new dataset with labels T-cell and B-cell. After that, another CNN < T-cell/B-cell > is trained from the newly created dataset, which can distinguish T-cells from B-cells. Then, cells in the original dataset are annotated by using the newly trained CNN < T-cell/B-cell >. We obtain the extended dataset by merging the newly created dataset and annotated original dataset. Lastly, AlexCAN is re-trained from the extended dataset as our final classifier, that can separate cells from background and can also annotate them as T-cell and B-cell
Training of convolutional neural network
Training a CNN from scratch requires a tremendous amount of data. Although it is possible to re-train a pre-trained CNN using a small dataset which is called transfer learning [18]. It is not always possible to have enough data for a certain application but using data which is in another domain can greatly improve the performance of learning. Transfer training makes use of the already existing CNNs and fine tunes them to the new datasets with a short training time. Many pre-trained CNNs exist like AlexNet [19], VGG [20] and GoogleNet [21]. In this study we use AlexNet since it is lightweight.
Transfer learning can be accomplished by removing the last layer of the AlexNet and using the rest of the parameters. We replaced the last layer of AlexNet with a fully connected layer according to the number of labels we have. Then using the new labelled data, we re-trained the pre-trained CNN. Matlab’s pretrained AlexNet was chosen as a starting point and a stochastic gradient descent with momentum algorithm (with hyperparameters: momentum 0.9, initial learn rate 0.001, max epochs 100, mini batch size 128) was used for re-training. In this study, our dataset consists of a few thousand training images and we only have three labels: T-cell, B-cell and background. Thus, our transfer learned CNN (from here on called AlexCAN: AlexNet based Cell Analyzer Network) has 3 neurons in its last layer.
Our initial dataset from previous work did not have annotations for T-cells and B-cells so it was only labeled as cells and background. We re-trained a CNN which can detect cells from the background as Fig. 2 shows. Training images were scanned exhaustively by employing the sliding window method to detect cells. We used 3 different scales for the detection window as 64 × 64 pixels, 80 × 80 pixels and 100 × 100 pixels. Since AlexNet requires an input size of 227 × 227 pixels, image patches must be resized before they can be classified. Following the classification of the whole image, multiple detection windows for single cells are combined and weak detections are eliminated.
Calibrating color distribution
In our previous work, after the detection of the cells, a color feature-based detector was used to identify cells as T-cells or B-cells respectively. The emission spectrum crosstalk between the dyes prevented us from annotating the cells manually, hence we dyed cells with a single dye, and introduced them to the same isolation system. As they were stained with only one fluorescent dye, the detected cells’ type could be determined.
Images from the single dye experiments are used to create a color dataset as Fig. 2 illustrates. Using the sliding windows method and trained CNN for detection, we created a dataset of 2300 cells annotated as T-cells and B-cells from 6 experiments and 2500 images. By using this new dataset, another CNN is re-trained which can separate cells into T-cells and B-cells. A fivefold cross validation is used to assess the performance. Although this process is straightforward and can be achieved with a simple color feature, we achieved a 99% accuracy rate compared to a 96% accuracy rate from our previous work which used a simpler color feature detector [8].
Due to the high purity of the detections from the trained CNN and overall accuracy of the automatic annotation, we could add this new dataset to the initial dataset. We also annotated the cells in the initial dataset as T-cells and B-cells using the newly re-trained CNN. This finalized our dataset with three labels as Fig. 2 illustrates. Using the final dataset AlexCAN is re-trained to detect the cells from the background and this time also annotate them as T-cells and B-cells. Cross validation proved that using the final dataset for AlexCAN can achieve the same 99% accuracy rate for classifying T-cells from B-cells using the color dataset. Moreover, its accuracy in distinguishing cells from the background is also equal to that of the previously re-trained CNN which could detect cells from the background without the type of the cell. Therefore, AlexCAN simplifies the two layer detection approach in our last work, which first detected cells from the background and then annotated them as T-cells and B-cells [8].
Speeding-up by preselecting windows using HOG and SVM
AlexCAN has improved performance in terms of identifying T-cells and B-cells and distinguishing them from noise and background. But processing all the images from a single experiment is still timewise infeasible even with the utilization of GPU processing (CPU i7-8700K 3.7 GHz, Nvidia GTX 1080Ti, 64 GB RAM) and efficient implementation (MATLAB—pretrained AlexNet). This is due to the number of detection windows that needs to be classified (in the order of 105 for a single image). Moreover every detection window has to be resized to the input size of the AlexCAN. However, even if a smaller CNN with an input size of our detection window is trained, it will still not be fast enough to be used in our application. Hence, we have to reduce the number of detection windows to meet the time requirement of our application.
Due to the design of the chip, cells are sparse on the images. Thus it is possible to reduce the number of detection windows by preselecting them by employing a simpler and faster method. In our previous work, we used a HOG features based SVM classifier to detect cells from noise and background. By using HOG features, it is possible to differentiate not only cells but also noise from the background. Hence, we prepared a dataset by coupling cells and noise images together as a positive dataset and used background images as a negative dataset. Figure 4 illustrates samples from this new dataset. Using this dataset, the HOG features based SVM classifier is trained.
We propose a two-layer classification scheme. At the first layer the HOG features based SVM classifier is used to detect cells and noise from background. The detection windows classified as positive by the HOG features based SVM constitute the preselected windows in our algorithm, and the remaining detection windows will be ignored. In other words, detections from the first classifier are used as input for the second classifier which is AlexCAN. We define the success rate of the HOG features based SVM classifier using the number of cells that are missed. When a cell is not within any preselected window, we count it as a miss. Figure 5 demonstrates the miss rate of cells by the first classifier versus the percentage of the preselected windows i.e. the ratio of preselected windows to all the detection windows. The miss rate decreases as the percentage of preselected windows increases. The processing time of AlexCAN increases linearly with the increasing of the percentage of the preselected windows, hence the miss rate has a lower bound depending on our time requirement.
Figure 6 illustrates this process flow, with an example image. Input images are first preprocessed by the HOG features based SVM classifier. Using the sliding windows approach, the images are scanned using three different scales as 64 × 64 pixels, 80 × 80 pixels and 100 × 100 pixels. Figure 6b shows the preselected windows which are the output of the first classification layer. The preselected windows are resized to the input size of AlexCAN and then each preselected window is classified as T-cell, B-cell and background. The detections from T-cells and B-cells are grouped separately, and weak windows are eliminated.
a Original image. b Green rectangles show preselected windows by the HOG features based SVM classifier, c classification results by AlexCAN; green rectangles show detection windows that are classified as T-cells and orange rectangles show detection windows that are classified as B-cells and d final result after grouping the detection windows
Results
We used the same test set from our previous work, in order to compare performance. 500 positive cell images were cropped, centered and scaled from the annotated test images. 50 complex examples containing noise and 450 randomly selected background examples were used as negative test set. To quantify the performance of the detectors we plotted the receiver operating characteristics (ROC’s), i.e. true positive rate (\(\frac{TruePos}{TruePos + FalseNeg}\)) versus false positive rate (\(\frac{FalsePos}{FalsePos + TrueNeg}\)). Also using this test set we calculated accuracy, specificity and sensitivity values as:
In addition to this set of 1000 image patches, negative images from the test data were exhaustively searched and around 4,500,000 negative images were added to the plot Detection Error Tradeoff (DET) curve on a log–log scale i.e. miss rate (\(\frac{FalseNeg}{TruePos + FalseNeg}\)) versus false positive per window (\(\frac{FalsePos}{Total\;Population}\)). The DET curves present the same information as the ROC’s but small differences in probabilities are easier to distinguish. We presented the performance of three different cases. In the first case AlexCAN was used to classify the image by employing the sliding window method. In the second case detection windows were preselected using the HOG features based SVM classifier to be used with AlexCAN. The last case was our previous work to provide a performance comparison between our previous work and AlexCAN and AlexCAN using preselected windows.
Figure 7 presents the ROC’s and DET curves for the three different cases. The different points in the ROC’s and DET curves correspond to different rates of accuracy, specificity and sensitivity. We achieved an accuracy of 98%, sensitivity of 97% and specificity of 99% in both cases using AlexCAN and AlexCAN using preselected windows, compared to the accuracy of 94%, sensitivity of 90% and specifity of 99% of our previous work [8].
In one experiment, a total of 420 images were scanned for the filtration zone on the microfluidic chip. Preselecting windows via the HOG feature based SVM classifier takes about 20 secs for 420 images. Classifying preselected windows with AlexCAN takes less than 10 min with our experiment PC. We used OpenCV libraries for the HOG features to perform GPU processing and Matlab for AlexCAN.
Discussion
The imaging technique and the methods to be used for the analysis is decided by the type of application. In the case of counting cells using standard microscopy, several methods exist for counting the cells in an image [22, 23]. If the cells are well separated and have uniform intensity, simple thresholding and watershed algorithms are popular. If the cells are not well separated, algorithms which account for cell shape and size are preferred [24]. In our application, fluorescence intensity is heterogenous throughout the microfluidic chip, and cells vary in shape due to deformation by micropillars. The intensity, shape and size features could not be utilized directly to detect cells in our pillar-based microfluidic chip system. In this study we adopted a supervised machine learning based approach, in which a model is automatically learned through examples of cells and background.
In our previous work, the HOG features were utilized to distinguish cells from background and noise. However, the HOG features only use shape information, and other features such as color and texture are omitted. In our application, the estimation of the ratio of T-cells and B-cells have utmost priority, so we chose to have a specificity rate as high as 99% and hence sacrificing the accuracy and sensitivity rate which were 94% and 90% respectively. This is a result of the HOG features’ inability to distinguish between noise and cells effectively.
We trained a pre-trained CNN employing transfer learning in order to address the low sensitivity. Owing to optimized feature kernels that are trained using an image dataset in the order of millions, the pre-trained AlexNet enables a great improvement in performance compared to the HOG features based SVM classifier. Transfer learning makes it possible to apply a pre-trained network to a new domain such as T-cell and B-cell detection with a dataset only in the order of thousands. With AlexCAN we achieved an accuracy of 98%, sensitivity of 97% and specificity of 99%. Even though recent advances in GPU architecture rendered it possible to train CNNs as large as ours, using the sliding window method and classifying all the windows in every image from an experiment is time consuming.
Increasing the speed for our application is crucial. However using a better GPU or training a smaller and faster CNN is unfortunately not the answer. We need an increase in speed in the order of hundreds, and we proposed a method to preselect detection windows. By reducing the number of windows to be classified, the total time for detection was reduced to 10 min. This method might miss some cells, but looking at our results, the total time required for detection is down to feasible levels. As Fig. 6d shows, AlexCAN using preselected windows can detect all the cells despite their varying morphology and size.
Conclusion
A deep convolutional neural network classifier is trained to detect and classify T-cells and B-cells isolated by a microfluidic chip. The experiments performed on various image datasets, have produced satisfactory detection results that prove the effectiveness of our proposed approach. We achieved a high accuracy of 98%, specificity of 99% and sensitivity of 97%. For T-cells and B-cells detection we achieved a 99% cross-validation accuracy rate. AlexCAN’s performance is found to be better compared to our previous work which utilized a HOG features based SVM classifier (accuracy of 94%, specificity of 99% and sensitivity of 90%). However, CNNs require more time compared to the simple HOG features based SVM classifier, and it is infeasible for practical use. Hence, we proposed a machine learning method to preselect windows to be classified by AlexCAN, and thus speeding up the process. The proposed method and system could also be applied to all other specific leukocytes using different stain agents.
References
Hulspas R, Bauman JG (1992) The use of fluorescent in situ hybridization for the analysis of nuclear architecture by confocal microscopy. Cell Biol Int Rep 16(8):739–747
Basiji DA, Ortyn WE, Liang L, Venkatachalam V, Morrissey P (2007) Cellular image analysis and imaging by flow cytometry. Clin Lab Med 27(3):653–670
Orchard JA, Ibbotson RE, Davis Z, Wiestner A, Rosenwald A, Thomas PW, Hamblin TJ, Staudt LM, Oscier DG (2004) ZAP-70 expression and prognosis in chronic lymphocytic leukaemia. The Lancet. 363(9403):105–111
Brasko C, Smith K, Molnar C, Farago N, Hegedus L, Balind A, Balassa T, Szkalisity A, Sukosd F, Kocsis K, Balint B (2018) Intelligent image-based in situ single-cell isolation. Nat Commun 9(1):226
Anselmetti D (ed) (2009) Single cell analysis: technologies and applications. John Wiley & Sons, Hoboken
Caicedo JC, Cooper S, Heigwer F, Warchal S, Qiu P, Molnar C, Vasilevich AS, Barry JD, Bansal HS, Kraus O, Wawer M (2017) Data-analysis strategies for image-based cell profiling. Nat Methods 14(9):849
Noor AM, Masuda T, Lei W, Horio K, Miyata Y, Namatame M, Hayase Y, Saito TI, Arai F (2018) A microfluidic chip for capturing, imaging and counting CD3+ T-lymphocytes and CD19+ B-lymphocytes from whole blood. Sens Actuat B Chem 276:107–113
Turan B, Masuda T, Lei W, Noor AM, Horio K, Saito TI, Miyata Y, Arai F (2018) A Pillar-based microfluidic chip for T-cells and B-cells isolation and detection with machine learning algorithm. Robomech J 5:27. https://doi.org/10.1186/s40648-018-0124-8
Dalal N, Triggs B (2005) Histograms of oriented gradients for human detection. In: Proceedings of computer vision and pattern recognition, p 886–893
Nitta N, Sugimura T, Isozaki A, Mikami H, Hiraki K, Sakuma S, Iino T, Arai F, Endo T, Fujiwaki Y, Fukuzawa H (2018) Intelligent image-activated cell sorting. Cell 175(1):266–276
Ota S, Horisaki R, Kawamura Y, Ugawa M, Sato I, Hashimoto K, Kamesawa R, Setoyama K, Yamaguchi S, Fujiu K, Waki K (2018) Ghost cytometry. Science 360(6394):1246–1251
Bojarski M, Del Testa D, Dworakowski D, Firner B, Flepp B, Goyal P, Jackel LD, Monfort M, Muller U, Zhang J, Zhang X. End to end learning for self-driving cars. arXiv preprint. arXiv:1604.07316. Accessed 25 Apr 2016
Parkhi OM, Vedaldi A, Zisserman A (2015) Deep face recognition. In: International conference on the British machine vision conference (BMVC), 7 Sept 2015, vol 1, p 6
Cruz-Roa AA, Ovalle JE, Madabhushi A, Osorio FA (2013) A deep learning architecture for image representation, visual interpretability and automated basal-cell carcinoma cancer detection. In: International conference on medical image computing and computer-assisted intervention. Springer, Berlin, p 403–410
Chen CL, Mahjoubfar A, Tai LC, Blaby IK, Huang A, Niazi KR, Jalali B (2016) Deep learning in label-free cell classification. Sci Rep 15(6):21471
Saltz J, Gupta R, Hou L, Kurc T, Singh P, Nguyen V, Samaras D, Shroyer KR, Zhao T, Batiste R, Van Arnam J (2018) Spatial organization and molecular correlation of tumor-infiltrating lymphocytes using deep learning on pathology images. Cell Rep 23(1):181
Deng J, Dong W, Socher R, Li LJ, Li K, Fei-Fei L. Imagenet: A large-scale hierarchical image database. InComputer Vision and Pattern Recognition, 2009. CVPR 2009. IEEE Conference on 2009 Jun 20 (pp. 248-255). Ieee
Pan SJ, Yang Q (2010) A survey on transfer learning. IEEE Trans Knowl Data Eng 22(10):1345–1359
Krizhevsky A, Sutskever I, Hinton GE (2012) Imagenet classification with deep convolutional neural networks. In: International conference on advances in neural information processing systems (NIPS), Lake Tahoe, Nevada, 3 Dec 2012, pp 1097–1105
Simonyan K, Zisserman A. Very deep convolutional networks for large-scale image recognition. arXiv preprint. arXiv:1409.1556. Accessed 4 Sep 2014
Szegedy C, Liu W, Jia Y, Sermanet P, Reed S, Anguelov D, Erhan D, Vanhoucke V, Rabinovich A (2015) Going deeper with convolutions. In: Proceedings of the IEEE conference on computer vision and pattern recognition, p 1–9
Kamentsky L, Jones TR, Fraser A, Bray MA, Logan DJ, Madden KL, Ljosa V, Rueden C, Eliceiri KW, Carpenter AE (2011) Improved structure, function and compatibility for Cell Profiler: modular high-throughput image analysis software. Bioinformatics 27(8):1179–1180
Wiesmann V, Franz D, Held C, Münzenmayer C, Palmisano R, Wittenberg T (2015) Review of free software tools for image analysis of fluorescence cell micrographs. J Microsc 257(1):39–53
Xie W, Noble JA, Zisserman A (2018) Microscopy cell counting and detection with fully convolutional regression networks. Comput Methods Biomech Biomed Eng Imaging Visual 6(3):283–292
Authors’ contributions
TM, BT, KH and AMN performed experiments and analysis. TM and BT prepared Figs. TM, BT and FA contributed to write the manuscript text. TM, TIS, YM and FA designed the study. All authors read and approved the final manuscript.
Acknowledgements
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Funding
This work was supported in part by START Program from Japan Science and Technology Agency, JST.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Turan, B., Masuda, T., Noor, A.M. et al. High accuracy detection for T-cells and B-cells using deep convolutional neural networks. Robomech J 5, 29 (2018). https://doi.org/10.1186/s40648-018-0128-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s40648-018-0128-4