Abstract
Background
Oxidative stress is known to be involved in most of the aetiological factors of nasopharyngeal carcinoma (NPC). Cells that are under oxidative stress may undergo apoptosis. We have previously demonstrated that oxidative stress-induced apoptosis could be a potential mechanism mediating chromosome breakages in nasopharyngeal epithelial cells. Additionally, caspase-activated DNase (CAD) may be the vital player in mediating the chromosomal breakages during oxidative stress-induced apoptosis. Chromosomal breakage occurs during apoptosis and chromosome rearrangement. Chromosomal breakages tend to cluster in certain regions, such as matrix association region/scaffold attachment region (MAR/SAR). We hypothesised that oxidative stress-induced apoptosis may result in chromosome breaks preferentially at the MAR/SAR sites. The AF9 gene at 9p22 was targeted in this study because 9p22 is a deletion site commonly found in NPC.
Results
By using MAR/SAR recognition signature (MRS), potential MAR/SAR sites were predicted in the AF9 gene. The predicted MAR/SAR sites precisely match to the experimentally determined MAR/SARs. Hydrogen peroxide (H2O2) was used to induce apoptosis in normal nasopharyngeal epithelial cells (NP69) and NPC cells (HK1). Nested inverse polymerase chain reaction was employed to identify the AF9 gene cleavages. In the SAR region, the gene cleavage frequency of H2O2-treated cells was significantly higher than that of the non-treated cells. A few chromosomal breakages were detected within the AF9 region which was previously found to be involved in the mixed lineage leukaemia (MLL)-AF9 translocation in an acute lymphoblastic leukaemia patient. As for the non-SAR region, no significant difference in the gene cleavage frequency was found between the untreated control and H2O2-treated cells. Furthermore, H2O2-induced cleavages within the SAR region were reduced by caspase-3 inhibitor, which indirectly inhibits CAD.
Conclusions
These results reaffirm our previous findings that oxidative stress-induced apoptosis could be one of the potential mechanisms underlying chromosome breakages in nasopharyngeal epithelial cells. MAR/SAR may play a vital role in defining the location of chromosomal breakages mediated by oxidative stress-induced apoptosis, where CAD is the major nuclease.
Similar content being viewed by others
Background
Nasopharyngeal carcinoma (NPC) is a type of malignant solid tumour which has been associated with multiple factors. One of the strong risk factors of NPC is Epstein–Barr virus (EBV) infection [1, 2]. A previous study has reported increased levels of immunoglobulin G (IgG) and immunoglobulin A (IgA) antibodies to EBV viral capsid antigen (VCA) and early antigen (EA) in NPC patients [1]. Besides, a case–control study of NPC among Malaysian Chinese has reported that salted fish consumption during childhood was a significant risk factor for developing NPC [3]. Mutagenic activity has been detected in the urine collected from experimental rats that were regularly fed with salted fish [4]. In addition, Chinese salted fish was found to cause nasal cavity tumours in rats [5, 6]. These findings suggested the presence of carcinogenic substances in salted fish. Most studies pointed toward nitrosamines and nitrosamine precursors, which have been recognised as animal carcinogens [7, 8]. In addition, long-term exposures to intense industrial heat, formaldehyde, cigarette smoke and wood dust have also been found to be significantly associated with NPC [3, 9,10,11,12]. Recently, chronic inflammation of sinonasal tract has been increasingly recognised as a risk factor for NPC [13, 14].
It is important to note that, all of these aetiological factors provoke the production of reactive oxygen species (ROS) [15,16,17,18,19,20]. Additionally, ROS was found to be involved in EBV reactivation in NPC cells after treatment with N-methyl-N′-nitro-N-nitroguanidine (MNNG) [21]. The ROS-mediated EBV reactivation was inhibited by apigenin which has been suggested to be a potent ROS scavenger [22]. Increased ROS may cause DNA double-strand breaks and error-prone repair. This may in turn lead to genomic instability [23]. The cancer cells and inflammatory cells in stroma of NPC patients have been found to contain oxidative and nitrative DNA lesions [24]. Oxidative stress may trigger apoptosis, a process of programmed cell death [25]. We have previously demonstrated that oxidative stress-induced apoptosis resulted in chromosomal breakages in normal nasopharyngeal epithelial and NPC cells. In addition, the apoptotic nuclease, caspase-activated DNase (CAD) may be a major player in mediating these chromosomal breakages [26].
Chromosomal breakage is an early event in both apoptotic DNA fragmentation and chromosome rearrangement. Previous studies revealed that chromosome breaks tend to fall within certain regions which contain specific chromatin structural elements, such as the matrix attachment region/scaffold attachment region (MAR/SAR) [27, 28]. MAR/SAR is the DNA sequence where the DNA loop structure binds to nuclear scaffold/matrix proteins [29]. In the early stage of apoptosis, DNA cleavages take place at the base of the DNA loop [30, 31]. We hypothesised that H2O2-induced apoptosis may cause chromosomal breakages at MAR/SAR resulting in chromosome rearrangement in nasopharyngeal epithelial cells.
This study focuses on the AF9 gene which is located at 9p22 because 9p22 is one of the deletion hotspots in NPC [32]. The AF9 gene is 280,880 bp in length. The nucleotide position of its exons and introns are shown in Additional file 1. Strissel et al. have identified two MAR/SARs within the AF9 gene. These two MAR/SARs were designated as SAR1 and SAR2 [28].
In the present study, in silico prediction of MAR/SAR sites was performed in the AF9 gene. It was found that in the region that contains MAR/SAR (SAR region), the gene cleavage frequency of H2O2-treated cells was higher than that of the untreated control. On the contrary, in the region that does not contain MAR/SAR (non-SAR region), there was no significant difference in gene cleavage frequency between untreated and H2O2-treated cells. These observations are true for both normal nasopharyngeal epithelial and NPC cells. Moreover, the oxidative stress-induced chromosome breakages within the SAR region were reduced by caspase-3 inhibitor, which indirectly inhibits CAD. Our results suggested that MAR/SAR may play an important role in defining the location of chromosome breaks mediated by oxidative stress-induced apoptosis, where CAD is the essential nuclease. These chromosomal breakages may in turn lead to chromosome aberrations in nasopharyngeal epithelial cells.
Methods
Cell lines and chemicals
NP69 normal nasopharyngeal epithelial cell line and HK1 NPC cell line were kindly provided by Prof. Tsao Sai Wah (The University of Hong Kong, Hong Kong, China) and Prof. Lo Kwok Wai (The Chinese University of Hong Kong, Hong Kong, China). StemPro ACCUTASE Cell Dissociation Reagent, Keratinocyte-SFM medium, RPMI 1640 medium, penicillin, streptomycin, l-glutamine and fetal bovine serum were purchased from GIBCO, Invitrogen, USA. Camptothecin (CPT) was purchased from Santa Cruz Biotechnology, California, USA. Hydrogen peroxide (H2O2) was bought from MP Biomedicals, USA. Annexin V-Fluorescein isothiocyanate (FITC) Apoptosis Detection Kit I (BD Pharmingen™) and Flow Cytometry Mitochondrial Membrane Potential Detection Kit (BD™MitoScreen) were obtained from Becton–Dickinson Biosciences, USA. Caspase-Glo 3/7 Assay Kit and dNTP mix were purchased from Promega, USA. Caspase-3 inhibitor II (Z-DEVD-FMK) was obtained from Calbiochem, USA. Isoamyl alchohol was procured from Fluka, Switzerland. Sodium dodecyl sulfate (SDS) and phenol were bought from Amresco, USA. Ammonium acetate was from Merck, Germany. Chloroform was obtained from R&M Chemicals, UK. All the restriction enzymes, T4 DNA Ligase and DNA Polymerase I Large (Klenow) Fragment were purchased from New England Biolabs (NEB), USA. QIAquick Gel Extraction Kit and QIAquick Nucleotide Removal Kit were obtained from QIAGEN, Germany. Phusion High-Fidelity DNA Polymerase was obtained from Finnzymes, Finland. PCR primers were bought from First Base Laboratories.
In silico prediction of MAR/SARs
The whole sequence of the AF9 gene was retrieved from Ensembl database [EMBL:ENSG00000171843]. The locations of experimentally isolated MAR/SAR, which were found within the AF9 gene, were determined from the previous reports [27, 28]. Possible MAR/SAR sites were also identified using MAR/SAR recognition signature (MRS) which was suggested to be strongly associated with MAR/SAR [33]. This MAR/SARs prediction was performed by using DNASTAR software (Lasergene, USA). The MRS comprises two nucleotide motifs that are within 200 bp apart. The first nucleotide motif is an 8 bp degenerate sequence, AATAAYAA while the second nucleotide motif is a 16 bp degenerate sequence, AWWRTAANNWWGNNNC, where Y = C or T; W = A or T; R = A or G; N = A, C, G or T. No mismatch is allowed within the 8 bp sequence whereas one mismatch is allowed within the 16 bp sequence. These two degenerate sequences should be within 200 bp apart. Each sequence can be aligned on either the Watson strand or the Crick strand. Either sequence can precede the other sequence. The sequences may even be overlapping. Clusters of more than one motif of either 8 or 16 bp within 200 bp apart are considered as a single MRS. Moreover, clusters of more than one MRS within close proximity are regarded as a single potential MAR/SAR site. The locations of the presently predicted MAR/SARs were compared with the locations of the experimentally extracted MAR/SARs reported in previous studies [27, 28].
In our in silico prediction of MAR/SAR which had been performed in the Abelson murine leukaemia viral oncogene homolog 1 (ABL) gene, there was only one MAR/SAR site predicted in the experimentally isolated SAR. The distance between 8 bp sequence element and the 16 bp sequence element was 248 bp (data not shown). Therefore, in this study, the maximal distance between 8 bp sequence element and the 16 bp sequence element was set at 250 bp.
Cell cultures
NP69 cells were grown in Keratinocyte-SFM medium supplemented with 4–5 ng/ml recombinant Epidermal Growth Factor (rEGF), 40–50 µg/ml Bovine Pituitary Extract (BPE), 100 U/ml penicillin, 100 µg/ml streptomycin and 2% (v/v) heat-inactivated fetal bovine serum. HK1 cells were cultured in RPMI 1640 medium supplemented with 2 mM l-glutamine, 100 U/ml penicillin, 100 µg/ml streptomycin and 10% (v/v) heat-inactivated fetal bovine serum. Cells were incubated at 37 °C with 5% CO2.
Detection of phosphatidylserine (PS) externalisation
NP69 cells (1.5 × 105) were plated in 150 mm culture dishes and allowed to grow for 48 h. The NP69 cells were incubated with 100 µM of H2O2 for 16 and 24 h. HK1 cells (5.5 × 105) were seeded in 150 mm culture dishes and allowed to grow for 72 h. The HK1 cells were incubated with 50 µM of H2O2 for 4 and 8 h. NP69 and HK1 cells treated with camptothecin (CPT) were used as positive controls. After exposure, the cells were harvested by using StemPro ACCUTASE Cell Dissociation Reagent. Annexin V-FITC Apoptosis Detection Kit I was used to detect PS externalisation in the harvested cells as previously described [26].
Detection of mitochondrial membrane potential (MMP) disruption
NP69 and HK1 cells were treated and harvested as described above. Flow Cytometry Mitochondrial Membrane Potential Detection Kit was used to detect the loss of MMP in the harvested cells as previously described [26].
Nested IPCR detection of oxidative stress-induced chromosome breaks
H2O2 treatment and genomic DNA extraction
NP69 cells (2 × 104) were plated in 60 mm culture dishes and allowed to grow for 48 h. The NP69 cells were incubated with H2O2 at concentration of 100 µM for 16 h. HK1 cells (8 × 104) were seeded in 60 mm culture dishes and allowed to grow for 72 h. The HK1 cells were incubated with H2O2 at concentration of 50 µM for 8 h. After treatment with H2O2, genomic DNA extraction was carried out as previously described [26].
Manipulation of genomic DNA and nested IPCR for the AF9 SAR region
The extracted genomic DNA was manipulated in preparation for nested IPCR as previously described [26]. Additional file 2 shows the simplified manipulation steps. Briefly, BamH I digestion, Klenow fill-in, cyclisation and ethanol precipitation were performed. The DNA was then either digested with Kpn I or Nde I. QIAGEN QIAquick Nucleotide Removal Kit was used to clean up the DNA. Nested IPCR was carried out as previously reported [26].
Manipulation of genomic DNA and nested IPCR for the AF9 non-SAR region
The manipulation steps were similar to the SAR region, except that Hind III (RE2 in Additional file 2) and Xba I (RE3 in Additional file 2) were used for the AF9 non-SAR region instead of Kpn I and Nde I. Cycle condition used in the first round of IPCR was as below: 30 s of 98 °C for 1 cycle (initial denaturation), followed by 30 cycles of 98 °C for 10 s (denaturation), 64 °C for 30 s (annealing), 72 °C for 22 s (extension), followed by 1 cycle of 72 °C for 10 min (final extension). Two microliter of fivefold diluted first round IPCR product was used for second round with similar cycle condition, except that the annealing temperature was 63 °C and the extension time was 15 s. The primers used for the first round of IPCR were 5′-TACCAAACATTTTGAGTCCTACAG-3′ (reverse) and 5′-GGCATTCAGGTGAGTAGTTTATTC-3′ (forward), whereas the primers used in the second round were 5′-AGCAGTAGACTTTTGTAACCTCAC-3′ (reverse) and 5′-AGGGGATGACTTTTCTTCAATC-3′ (forward).
Inhibition of caspase by Z-DEVD-FMK
HK1 cells (8 × 104) were seeded in 60 mm culture dishes and grown until 60–70% confluency. HK1 cells were either left untreated or pretreated with 50 μM of Z-DEVD-FMK for 1 h. The HK1 cells were then either left untreated or co-treated with 50 μM of H2O2 for 8 h. After incubation, genomic DNA was extracted as previously described [26]. Following that, IPCR identification of the chromosome breaks within the AF9 SAR and non-SAR regions was performed as described above.
Visualisation and DNA sequencing of the IPCR products
The IPCR products were loaded on 1% agarose gel. To analyse the IPCR bands, the gel was stained with ethidium bromide and visualised on an ultraviolet (UV) transilluminator (Vilber Lourmat, USA). QIAGEN QIAquick Gel Extraction Kit was used to clean up the IPCR bands which represent the cleaved fragments derived from the AF9 SAR region. The purified IPCR bands were sequenced. By blasting the human genome database (Nucleotide BLAST, http://blast.ncbi.nlm.nih.gov/Blast.cgi), the sequencing results were annotated. To identify the position of the chromosome breaks, the sequencing results were aligned with the AF9 gene sequence accessed from EMBL database [EMBL:ENSG00000171843] by using Seqman DNASTAR software (Lasergene, USA). The breakpoints identified were compared with the location of the experimentally extracted MAR/SARs reported in the previous study [28] and the MRS predicted MAR/SARs. A genomic map was constructed to illustrate the position of the chromosome breaks relative to the location of the MAR/SARs.
Quantification of gene cleavage frequency
One to two sets of nested IPCR assays were performed for each experiment. Each set of IPCR assay consisted of five to eight IPCR replicates per cell sample. The number of IPCR bands which represent the DNA fragments derived from the cleaved AF9 SAR and non-SAR regions was counted. Gene cleavage frequency expresses the median number of chromosome breaks detected within the AF9 SAR region or non-SAR region in two to three independent experiments.
Statistical analysis
The Student’s t test was used to assess the difference between untreated control and treated samples in the detections of PS externalisation and MMP disruption. The Mann–Whitney U test was used to analyse the difference between untreated control and treated samples in the nested IPCR assays. For the detections of PS externalisation and MMP disruption, data are presented as means and standard deviation (SD). For the IPCR assays, data are expressed as median and interquartile range (IQR). Differences were considered statistically significant at p value < 0.05. All statistical tests are two-sided.
Results
In silico prediction of MAR/SAR
By using MRS, 29 possible MAR/SAR sites were predicted in the AF9 gene. The nucleotide positions of the MRSs with their sequence composition, relative orientation, distance between the two sequence elements and location of the MRSs in the exon or intron of the AF9 gene are shown in Table 1. Out of the 29 predicted MAR/SAR sites, 14 were found in intron 2 (MAR/SARs 2–15 in Table 1). Intron 2 is the largest intron of the AF9 gene which is approximately 164 kb in length. Five MAR/SAR sites were predicted in each intron 3b (MAR/SARs 17–21 in Table 1) and intron 4 (MAR/SARs 22–26 in Table 1). Intron 7 was found to contain two potential MAR/SAR sites (MAR/SARs 27–28 in Table 1). One MAR/SAR site was predicted in each intron 1 (MAR/SAR 1 in Table 1), intron 3a (MAR/SAR 16 in Table 1) and intron 9 (MAR/SAR 29 in Table 1).
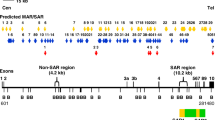
The distribution of the predicted MAR/SAR sites in the AF9 gene is illustrated in Fig. 1. Based on this in silico prediction of MAR/SAR, we determined a SAR region (contains MAR/SAR) and a non-SAR region (does not contain MAR/SAR) as the targeted regions of our study. The AF9 SAR region contains four MRSs (MAR/SARs 24-1 to 24-4 in Table 1). However, they are regarded as a single potential MAR/SAR site (MAR/SAR 24) because they were found in close proximity to each other (< 1.5 kb). Three out of these four MRSs were found within the biochemically defined SAR1 (located in intron 4) [28]. On the contrary, the AF9 non-SAR region is a region which contains neither biochemically defined MAR/SAR nor MRS predicted MAR/SAR.
Distribution of potential MAR/SAR sites predicted in the AF9 gene. The AF9 genomic map from nucleotide positions 601–281,480 is illustrated above [EMBL:ENSG00000171843]. The locations of exons 1 to 10 and BamH I (B) restriction sites are shown. Green boxes indicate the two patient BCRs reported in the previous study. These two patient BCRs were denominated as BCR1 and BCR2 [28]. Yellow boxes indicate the two MAR/SARs that were biochemically identified by Strissel and co-workers. These two MAR/SARs were designated as SAR1 and SAR2 [28]. Yellow arrows represent the potential MAR/SAR sites predicted by MRS in our study. Clusters of more than one MRS within close proximity are regarded as a single potential MAR/SAR site. Three MRSs were found in SAR1 (MAR 24-2, 24-3, 24-4). One MRS (MAR 27) has been predicted next to the SAR2. Based on the in silico prediction of MAR/SAR, a SAR region (contains MAR/SAR) and a non-SAR region (does not contain MAR/SAR) were determined to be the regions of study
Apoptosis detection in H2O2-treated NP69 and HK1 cells
By using flow cytometric analyses of PS externalisation and MMP disruption, significant percentages of apoptosis were detected in H2O2-treated NP69 and HK1 cells. These data have been reported in our previous study [26]. Our findings indicate that H2O2 could induce apoptosis in NP69 and HK1 cells.
IPCR detection of chromosome breaks mediated by H2O2-induced apoptosis in NP69 cells
To detect chromosome breaks within the AF9 SAR and non-SAR regions in cells undergoing H2O2-induced apoptosis, nested IPCR assay was performed. In the SAR region, IPCR primers were designed to detect chromosome breaks within the first breakpoint cluster region of the AF9 gene (BCR1). The AF9 BCR1 is located at the telomeric end of intron 4. It is bordered by two biochemically defined MAR/SARs [27, 28]. The SAR region also contains one MRS predicted MAR/SAR (MAR/SAR 24 in Table 1). The non-SAR region is a region which contains neither biochemically defined MAR/SAR nor MRS predicted MAR/SAR. The intact IPCR band for the AF9 SAR region and non-SAR region are 944 bp (~ 950 bp) and 956 bp (~ 950 bp), respectively. If there is chromosome break within the region of study, for both SAR and non-SAR regions, IPCR band of less than 950 bp will be detected.
AF9 SAR region
As shown in Fig. 2a i, numerous IPCR bands smaller than 950 bp which represent the cleaved AF9 gene were identified in NP69 cells treated with H2O2 (lanes 8–13). The untreated NP69 cells also show a few IPCR bands (lanes 3–7). By using flow cytometric analysis of phosphatidylserine (PS) externalisation, we detected a small amount of apoptotic cells in the untreated sample ([26], Fig. 1). These apoptotic cells might undergo spontaneous DNA breaks and contribute to the background as seen in lanes 3–7. As summarised by the box plot in Fig. 2b, the median AF9 cleavage frequency of H2O2-treated NP69 cells was 2.0-fold higher than that of the untreated control cells (p = 0.008). Our findings clearly indicate that H2O2-induced apoptosis results in cleavages within the AF9 SAR region.
Cleavage frequencies of the AF9 SAR and non-SAR regions in H2O2-treated NP69 cells. a Representative gel pictures showing the IPCR results of the AF9 gene in H2O2-treated NP69 cells: i SAR region, ii Non-SAR region. NP69 cells were either untreated (lanes 2–7) or treated with 100 µM of H2O2 for 16 h (lanes 8–13). The cells were harvested for gDNA extraction and nested IPCR. For each cell sample, six IPCR replicates (R1–R6) were prepared. The side brackets show the IPCR bands derived from the cleavages of the AF9 gene. M: 100 bp DNA ladder. N: Negative control for IPCR. b AF9 cleavage frequency detected in NP69 cells. Data are representative of three independent experiments. Each experiment consisted of 1–2 sets of IPCR. Each set of IPCR was performed in 5–8 IPCR replicates per cell sample. The results are presented as medians with IQRs. *P < 0.05; NS: no significant difference (Mann–Whitney U test)
AF9 non-SAR region
As shown in Fig. 2a ii, numerous IPCR bands of less than 950 bp which represent the cleaved AF9 gene were detected in both untreated NP69 cells (lanes 2–7) and NP69 cells treated with H2O2 (lanes 8–13). However, there was no significant difference between the untreated cells and H2O2-treated cells in the cleavage frequency of the AF9 non-SAR region (p = 0.739) (Fig. 2b).
IPCR detection of chromosome breaks mediated by H2O2-induced apoptosis in HK1 cells
AF9 SAR region
To further strengthen our observation that H2O2 could induce chromosome breaks within the AF9 SAR region, IPCR detection of chromosome breaks was also performed in H2O2-treated HK1 cells. The representative gel picture in Fig. 3a i shows that more IPCR bands were identified in H2O2-treated HK1 cells (lanes 7–11) as compared with the untreated control cells (lanes 2–6). The median AF9 cleavage frequency of H2O2-treated HK1 cells was 4.0-fold higher than that of the untreated control cells (p < 0.001) (Fig. 3b). These findings strengthen the suggestion that oxidative stress-induced apoptosis leads to AF9 gene cleavages within the SAR region.
Cleavage frequencies of the AF9 SAR and non-SAR regions in H2O2-treated HK1 cells. a Representative gel pictures showing the IPCR results of the AF9 gene in H2O2-treated HK1 cells: i SAR region, ii non-SAR region. HK1 cells were either untreated (lanes 2–6) or treated with 50 µM of H2O2 for 8 h (lanes 7–11). The cells were harvested for gDNA extraction and nested IPCR. For each cell sample, five IPCR replicates (R1–R5) were prepared. The side brackets show the IPCR bands derived from the cleavages of the AF9 gene. M: 100 bp DNA ladder. N: Negative control for IPCR. b AF9 cleavage frequency detected in HK1 cells. Data are representative of two independent experiments. Each experiment consisted of 1–2 sets of IPCR. Each set of IPCR was performed in 5–6 IPCR replicates per cell sample. The results are expressed as medians with IQRs. *P < 0.05; NS: no significant difference (Mann–Whitney U test)
AF9 non-SAR region
As shown in Fig. 3a ii, numerous IPCR bands of less than 950 bp which represent the cleaved AF9 gene were detected in both untreated HK1 cells (lanes 2–6) and H2O2-treated HK1 cells (lanes 7–11). However, there was no significant difference between the untreated HK1 cells and H2O2-treated HK1 cells in the cleavage frequency of the AF9 non-SAR region (p = 0.405) (Fig. 3b). Taken together, our findings suggest that MAR/SAR sequence plays a crucial role in defining the location of chromosome breaks in H2O2-induced apoptosis.
In this study, we hypothesise that MAR/SAR is a preferential site of chromosome breaks. Therefore, less or no chromosome break was expected to be detected in this non-SAR region after H2O2 treatment. However, the current results are not as what was expected. There are obviously more cleavage bands detected in the non-SAR region compared with the SAR region (Figs. 2 and 3). The box plot in Fig. 2b shows that in the untreated NP69 cells, the median cleavage frequency of the non-SAR region was 4.0-fold higher than that of the SAR region (p = 0.002). Similarly, in the untreated HK1 cells, the median cleavage frequency of the non-SAR region was 5.5-fold higher than that of the SAR region (p < 0.001) (Fig. 3b). Such a difference might reflect that there are other chromatin structures which may also contribute to DNA fragility. In addition to MAR/SAR sequence, repeat elements have also been well implicated in mediating chromosome breaks [27, 34]. Hence, this prompted us to investigate the possibility of repeat elements in contributing to DNA fragility of the AF9 non-SAR region.
Identification of repeat elements within the AF9 gene
CENSOR program (http://www.girinst.org/censor/) was used to identify repeat elements in the AF9 gene. The repeat elements identified within the SAR and non-SAR regions are shown in Table 2. The locations of repeat elements identified within the SAR and non-SAR regions are illustrated in Fig. 4. There are 18 repeat elements identified within the 10.2 kb SAR region (Table 2). Only one out of these 18 repeat elements is located within the amplified region. The region amplified by the reverse primer (AF9 236211 R) is from coordinates 236,059 to 236,211. This region does not contain any repeat element. The region amplified by the forward primer (AF9 245507 F) is from coordinates 245,507 to 246,292. ERE2_EH (coordinates 245,627–245,728, 102 bp in length) is the only one repeat element identified in this region. It occupies 11% (102 bp) of the amplified SAR region (944 bp).
The repeat elements identified within the AF9 SAR and the non-SAR regions. a The SAR region. The SAR region which is bordered by two BamH I sites is 10.2 kb in length (from coordinates 236,059 to 246,292). Green box represents the previously identified patient BCR which is indicated as BCR1. Yellow box shows the previously experimentally isolated MAR/SAR which is indicated as SAR1 [28]. Yellow arrows represent the potential MAR/SAR sites predicted by MRS in the present study. Green and blue arrows represent the primers used in the first and second rounds of nested IPCR, respectively. Black boxes show the repeat elements predicted by CENSOR program. BamH I (B), Kpn I (K) and Nde I (N) restriction sites are shown. b The non-SAR region. The non-SAR region which is bordered by two BamH I sites is 4.2 kb in length (from coordinates 71,116 to 75,277). Green and blue arrows represent the primers used in the first and second rounds of nested IPCR, respectively. Black boxes represent the repeat elements identified by using CENSOR program. BamH I (B), Hind III (H) and Xba I (X) restriction sites are shown
On the other hand, there are nine repeat elements identified within the 4.2 kb non-SAR region (Table 2). Three out of these nine repeat elements are located within the amplified region. The region amplified by the reverse primer (AF9 71282 R) is from coordinates 71,116 to 71,282. There was no repeat element identified in this region. The region amplified by the forward primer (AF9 74494 F) is from coordinates 74,494 to 75,277. There are three repeat elements located in this region, namely two CHARLIE5 (coordinates 74,895–74,998, 104 bp in length and coordinates 75,006–75,169, 164 bp in length) and one AluJr (coordinates 75,192–75,466, 275 bp in length). These three repeat elements (the first CHARLIE5, 104 bp; the second CHARLIE5, 164 bp and AluJr, 275 bp) occupy 57% (543 bp) of the amplified non-SAR region (956 bp). In brief, given that there is no significant difference in the cleavage frequencies between the untreated and H2O2-treated cells, the chromosome breaks in the non-SAR region were most likely not mediated by H2O2-induced apoptosis. It is most likely that the presence of the repeat elements contribute to the DNA fragility of the non-SAR region.
Inhibition of caspase
SAR region
Figure 5a i, ii show the representative IPCR results of the AF9 SAR region in H2O2-treated HK1 cells without and with caspase inhibitor (CI) pretreatment, respectively. In the absence of CI, the median cleavage frequency of the AF9 gene detected in H2O2-treated HK1 cells was 4.0-fold higher than that of the untreated control cells (p < 0.001) (Fig. 5b). The median cleavage frequency of the AF9 SAR region in H2O2-treated HK1 cells with CI pre-treatment was 4.0-fold lower than that without CI pre-treatment (p = 0.004) (Fig. 5b). These results indicate that H2O2 induces cleavages within the AF9 SAR region in a caspase-3-dependent manner.
Caspase-3 inhibitor abolishes H2O2-induced cleavages within the AF9 SAR region. a Representative gel pictures showing the IPCR analysis of the AF9 SAR region in H2O2-treated HK1 cells: i without CI pre-treatment ii with CI pre-treatment. HK1 cells were left untreated or pre-treated with 50 µM of Z-DEVD-FMK for 1 h. The cells were then either untreated (lanes 2–7) or treated with 50 µM of H2O2 for 8 h (lanes 8–13). Genomic DNA was extracted and modified for nested IPCR. For each cell sample, six IPCR replicates (R1–R6) were prepared. The side brackets show the IPCR bands derived from the cleavages of the AF9 gene. M: 100 bp DNA ladder. N: Negative control for IPCR. b Cleavage frequency of the AF9 SAR region detected in HK1 cells. Data are representative of two independent experiments. Each experiment consisted of 1–2 sets of IPCR. Each set of IPCR was performed in 5–6 IPCR replicates per cell sample. The results are expressed as medians with IQRs. *P < 0.05 (Mann–Whitney U test)
Non-SAR region
Figure 6a i, ii show the representative IPCR results of the AF9 non-SAR region in H2O2-treated HK1 cells without and with CI pre-treatment, respectively. There is no significant difference in the cleavage frequency of the non-SAR region between the untreated control and H2O2-treated HK1 cells (p = 0.405) (Fig. 6b). There is also no significant difference in the cleavage frequency between the H2O2-treated HK1 cells without CI pre-treatment and that with CI pre-treatment (p = 0.390) (Fig. 6b). These findings show that CI has no significant effect on the cleavage frequency within the AF9 non-SAR region. This implies that the cleavages of the non-SAR region are not dependent on caspase-3.
Caspase-3 inhibitor shows no effect on reducing cleavages within the AF9 non-SAR region. a Representative gel pictures showing the IPCR analysis of the AF9 non-SAR region in H2O2-treated HK1 cells: i without CI pre-treatment ii with CI pre-treatment. HK1 cells were left untreated or pre-treated with 50 µM of Z-DEVD-FMK for 1 h. The cells were then either untreated (lanes 2–7) or treated with 50 µM of H2O2 for 8 h (lanes 8–13). Genomic DNA was extracted and modified for nested IPCR. For each cell sample, six IPCR replicates (R1–R6) were prepared. The side brackets show the IPCR bands derived from the cleavages of the AF9 gene. M: 100 bp DNA ladder. N: Negative control for IPCR. b Cleavage frequency of the AF9 non-SAR region detected in HK1 cells. Data are representative of two independent experiments. Each experiment consisted of 5–7 IPCR replicates per cell sample. The results are expressed as medians with IQRs. NS: No significant difference (Mann–Whitney U test)
Sequencing results
Some of the IPCR bands detected within the AF9 SAR region were excised, purified and sequenced. The sequencing results show that these fragments were all derived from the cleaved AF9 gene (Additional file 3). Table 3 shows the position of chromosome breaks identified within the AF9 SAR region in H2O2-treated NP69 and HK1 cells. Intriguingly, three chromosome breaks (at coordinates 245,560, 245,566 and 245,591) are identified within the AF9 region (at coordinate 245,252–245,612) that was previously reported to translocate with the mixed lineage leukaemia (MLL) gene. This reciprocal translocation t(9;11)(p22;q23) resulted in the formation of MLL-AF9 fusion gene in an acute lymphoblastic leukaemia (ALL) patient [GenBank:AM050804]. Seven breakpoints (at coordinates 245,560, 245,566, 245,591, 245,634, 245,645, 245,659 and 245,681) are within a distance of 70 nucleotides from the breakpoint identified in the ALL patient (at coordinate 245,612) [GenBank:AM050804]. A breakpoint mapped at coordinate 245,591 is similar with that identified in cultured normal blood cells treated with etoposide (VP16) (at coordinate 245,593) [35]. A genomic map illustrating the positions of H2O2-induced chromosome breaks in NP69 and HK1 cells relative to the MAR/SAR sequences within the AF9 gene is shown in Fig. 7.
The positions of H2O2-induced chromosome breaks within the AF9 SAR region. a The AF9 genomic map from nucleotide positions 601–281,480 [EMBL:ENSG00000171843] [26]. Black vertical lines represent the locations of exons 1–10. Green boxes show the two previously identified patient BCRs, namely BCR1 and BCR2 [28]. Yellow boxes show the two MAR/SARs which were extracted experimentally in the previous study. These two MAR/SARs were indicated as SAR1 and SAR2 [28]. BamH I (B) restriction sites are shown. Yellow arrows represent the potential MAR/SAR sites predicted by MRS in our study. b The AF9 SAR region (10.2 kb). BamH I (B), Kpn I (K) and Nde I (N) restriction sites are shown. Green and blue arrows represent the primers used in the first and second rounds of nested IPCR, respectively. c Breakpoints mapped in H2O2-treated cells. Red and green vertical lines show the breakpoints identified in H2O2-treated NP69 and HK1 cells, respectively. All the chromosome breaks were mapped within BCR1 which is bordered SAR1 and SAR2. Blue box represents the AF9 region (at coordinates 245,252–245,612) that was previously reported to translocate with the MLL gene resulting in the formation of the MLL-AF9 fusion gene in an ALL patient [GenBank:AM050804]
Discussion
Much effort had been employed to identify tumor suppressor genes and oncogenes associated with NPC (reviewed in [36, 37]). However, the underlying mechanism of NPC chromosome rearrangement remains elusive. Oxidative stress has been well implicated in carcinogenesis [38]. Most of the aetiological factors of NPC are known to induce oxidative stress [15,16,17,18,19,20]. In addition, oxidative stress is also a potent apoptotic inducer [39]. Although apoptosis has long been recognised as a programmed cell death process [40], the perception that cells undergoing apoptosis are destined to die has been challenged [41]. It was shown that cells have the potential to recover from the execution phase of apoptosis through DNA repair. However, surviving cells that have undergone compromised DNA repair may carry chromosome rearrangements [41, 42].
In order to test the apoptotic effect of H2O2 in NP69 and HK1 cells, we analysed the H2O2-treated NP69 and HK1 cells by flow cytometric analyses of PS externalisation and MMP loss. Our observations showed that H2O2 could induce apoptosis in both NP69 and HK1 cells. These data have been published in our previous report [26]. Chromosomal breakage resulting from chromosome loop excision is an initial event in both apoptotic DNA fragmentation and chromosome rearrangement. It was found that chromosome breaks tend to fall in certain regions containing specific chromatin structural elements such as MAR/SAR [27, 28]. MAR/SAR sequences possess unwinding properties which facilitate the entry of protein factors involved in apoptosis, replication, transcription and chromosome condensation [43, 44]. The unwinding properties of MAR/SAR sequences also render them to be more susceptible to cleavage [44, 45]. In our previous report, we demonstrated that high cell density and EBV latent membrane protein 1 (LMP1) expression triggered apoptosis in NPC cells. This in turn caused cleavages of the MLL BCR at the MAR/SAR sequence. These findings implied that MAR/SAR may play an essential role in defining the cleavage sites during high cell density or LMP1-induced apoptosis [46]. In this study, we intended to investigate if MAR/SAR is a preferential site of chromosome breaks mediated by oxidative stress-induced apoptosis.
The human AF9 gene at 9p22 was targeted in this study for two reasons. Firstly, this gene is one of the most common fusion partner genes of the MLL gene at 11q23 [28]. The t(9;11)(p22;q23) has been strongly associated with acute myelogenous leukaemia (AML), less common with therapy-related AML (t-AML), with ALL and myelodysplastic syndromes (MDS) [28, 47]. Secondly, 9p22 is a common chromosomal deletion site in NPC [32]. There were two MAR/SARs isolated experimentally in the AF9 gene. They were designated as SAR1 and SAR2. SAR1 is found in intron 4, whereas SAR2 spans from exons 5 to 7. Two patient breakpoint cluster regions (BCR) have been identified in the AF9 gene, namely, BCR1 and BCR2. BCR1 is located in intron 4, whereas BCR2 encompasses introns 7 to 8. These two BCRs are bordered by SAR1 and SAR2 [27, 28].
In the present study, in silico prediction of MAR/SAR was performed by using MRS. MRS is a bipartite sequence element that is specific for a large group of MAR/SARs. MRS consists of two individual sequence elements that are approximately 200 bp apart. However, when the DNA is wrapped around the histones, these two sequence elements are located at a position near the dyad axis of the nucleosome. Therefore, they can be aligned together in MAR/SAR after the nucleosomes are positioned. The close proximity between these two sequence elements on the positioned nucleosome enables them to create a protein binding site in MAR/SAR. In the study of van Drunen and co-workers, more than 300 kb of DNA sequence from several eukaryotic organisms were analysed. Their findings showed that all the MRSs that have been identified map specifically to the biochemically identified MAR/SARs [33]. MRS has been widely used in previous studies. MRS has been used to predict MAR/SAR in the human LMP/TAP gene region. All of the five predicted MAR/SARs in the analysed region match to the experimentally defined MAR/SARs [48]. Besides, MRS has also been used to identify the positions of MAR/SARs in human β-globin locus [49] and wheat high-molecular-weight glutenin 1Dy10 gene promoter [50].
The potential MAR/SAR sites predicted by MRS in the present study were compared with the location of biochemically identified MAR/SAR reported in previous studies [27, 28]. Strissel et al. have analysed 61 kb of the AF9 region for MAR/SAR. Their region of study encompassed exons 4 to 10. In their region of study, two MAR/SARs were identified through experimental extraction. These two MAR/SARs were designated as SAR1 and SAR2. SAR1 is a 6.2 kb MAR/SAR located in intron 4 whereas SAR2 is a 4.6 kb MAR/SAR spans through parts of introns 5 to 7. To the extent of our knowledge, no analysis on MAR/SAR was reported for the AF9 region from exon 1 to intron 3.
Within the AF9 gene of 280,880 bp in length, 29 possible MAR/SAR sites were predicted in our study. Four MRSs (MAR/SARs 24-1 to 24-4 in Table 1 and Fig. 1) are associated with SAR1. However, these four MRSs are regarded as a single potential MAR/SAR site (MAR/SAR 24) because they cluster within close proximity to each other (< 1.5 kb). Three out of these four MRSs fall within SAR1 (MAR/SARs 24-2 to 24-4 in Table 1 and Fig. 1). One of the MRSs was found in a region < 1 kb centromeric to SAR1 (MAR/SARs 24-1 in Table 1 and Fig. 1). In addition, one MAR/SAR site (MAR/SAR 27 in Table 1 and Fig. 1) has been predicted in a region < 1.5 kb telomeric to SAR2.
In the present study, both the normal transformed nasopharyngeal epithelial cells (NP69) and nasopharyngeal carcinoma cells (HK1) were used. In both of these cell lines, oxidative stress-induced apoptosis results in cleavages within the AF9 SAR sequences. To relate the position of H2O2-induced chromosome breaks with the MAR/SAR sites, the IPCR bands that represent cleavages within the AF9 SAR region were sequenced. The sequencing results revealed that all the chromosome breaks were mapped within BCR1 which is bordered by SAR1 and SAR2 (Fig. 7). Intriguingly, a few chromosome breaks were mapped within the region of AF9 that was previously reported being involved in the formation of the MLL-AF9 fusion gene in an ALL patient [GenBank:AM050804]. Similar chromatin structural elements have been identified in the BCRs of the AF9 and MLL genes. These include DNase I hypersensitive (HS) cleavage sites, MAR/SAR sequences and topoisomerase II cleavage sites. The similarity in the structural elements is suggested to cause the AF9 and MLL BCRs to be the recombination hotspots resulting in MLL-AF9 translocations in leukaemia [28]. Taken together, our results are consistent with those of the other studies which found that MAR/SAR may be a preferential site of chromosome breaks in apoptosis [51] and chromosome rearrangements [27,28,29]. Considering the observations in leukaemic cells and nasopharyngeal epithelial cells, it is plausible that regardless of the cancer type, the chromatin structure could be playing a vital role in determining the site of chromosome rearrangement.
In order to further examine the involvement of MAR/SAR in stress-induced chromosome breaks formation, the cleavage frequency of the AF9 SAR region was compared with that of the AF9 non-SAR region. We hypothesised that MAR/SAR is a preferential site of chromosome breaks, thus we expected to see less or no chromosome breaks detected in the non-SAR region after H2O2 treatment. However, to our surprise, in both untreated NP69 and HK1 cells, the cleavage frequencies of the non-SAR region were significantly higher than those of the SAR region. These findings imply that there are other chromatin structures which may also contribute to DNA fragility. In addition to MAR/SAR sequence, repeat elements have also been strongly implicated in mediating chromosome breaks [27, 34]. Thus, the possibility of repeat elements in contributing to DNA fragility of the AF9 non-SAR region was explored.
By using CENSOR program, three repeat elements (the first CHARLIE5, 104 bp; the second CHARLIE5, 164 bp and AluJr, 275 bp) were identified in the amplified non-SAR region of the AF9 gene. These repeat elements occupy 57% (543 bp) of the amplified non-SAR region (956 bp). On the contrary, ERE2_EH (102 bp in length) is the only one repeat element identified in the amplified SAR region. It occupies 11% (102 bp) of the amplified SAR region (944 bp).
It is noteworthy that, although the cleavage frequency of the non-SAR region detected in the untreated cells was higher than that of the SAR region, there was no significant difference between the H2O2-treated cells and untreated cells in the cleavage frequency of the AF9 non-SAR region. This is true for both NP69 and HK1 cells. Hence, it can be suggested that the cleavages identified in the non-SAR region were not stress-induced or stress-mediated. It is likely that the presence of repeat elements renders the chromosome to be more prone to cleavage. Previous studies have reported that common fragile sites, including FRA3B, FRA7G, FRA7H, FRA16D and FRAXB have all been shown to contain a high proportion of repeat elements, such as interspersed repeat elements, long terminal repeats (LTR), transposable elements, Mirs, L1 elements, L2 elements and Alu elements. These repetitive elements have been associated with the fragility of these fragile sites [52, 53]. The findings of our study conclude that MAR/SAR may be a preferential site of chromosome breaks during oxidative stress-induced apoptosis and may play an important role in oxidative stress-induced chromosome rearrangement.
We have previously demonstrated that H2O2 induces apoptosis in NP69 and HK1 cells in a caspase-3-dependent manner. By using Caspase-Glo 3/7, a luminescence-based assay, activation of caspase-3/7 was detected in H2O2-treated NP69 and HK1 cells. Pretreatment with Z-DEVD-FMK inhibits the activity of caspase-3/7 in H2O2-treated cells [26]. In the cytoplasm of healthy cells, CAD exists naturally as a heterodimer with its chaperone, inhibitor of CAD (ICAD). ICAD possesses two caspase-3 cleavage sites. Upon caspase-3-mediated cleavage of ICAD, CAD is released from ICAD. Subsequently, CAD enters the nucleus and cleaves DNA by generating double-strand breaks [54, 55]. Given that ICAD is primarily inactivated by DEVD-cleaving caspase-3 [56], inhibiting caspase-3 by using Z-DEVD-FMK is the most effective way of inactivating CAD. Therefore, if CAD is responsible for mediating chromosome breaks in H2O2-induced apoptosis, the chromosome breaks in H2O2-treated cells will be reduced or eliminated when caspase-3 is inhibited.
For the AF9 SAR region, inhibition of caspase by Z-DEVD-FMK significantly reduced the AF9 cleavages in H2O2-treated HK1 cells. Our findings suggest that, H2O2 induces chromosome breaks through caspase-3 activation. This study confirms the claims made in previous researches where H2O2 induces DNA fragmentation in a caspase-3-dependent manner [39]. Given that activated caspase-3 can stimulate CAD which is responsible for apoptotic DNA fragmentation, CAD is most likely the major player responsible for H2O2-induced chromosome breaks within the AF9 SAR region. Indeed, our previous study had demonstrated that, overexpression of ICAD resulted in expression of CAD and also inhibited H2O2-induced MLL gene cleavages. The observations of our previous study suggested a role for CAD in mediating H2O2-induced chromosome breaks [57].
In addition, our findings were supported by other research that CAD preferentially binds to the nuclear matrix of cells undergoing apoptosis. CAD/ICAD complex is freely moving in dividing cells. However, once apoptosis is induced, the mobility of the activated CAD becomes gradually restricted. The immobilisation of CAD is due to its association with the nuclear matrix [51]. Nuclear matrix is the binding site for the organisation of DNA loop structure [58]. DNA interacts with the nuclear matrix through MAR/SAR sequences [59]. When CAD binds to the nuclear matrix during apoptosis [51], it is in close proximity to the MAR/SAR sequences of the DNA loops. Hence, CAD potentially cleaves the DNA at the MAR/SAR sequences when it is being associated with the nuclear matrix. The reduction of cleavages within the SAR region by inhibiting CAD thus supports our hypothesis that CAD cleaves the DNA preferentially at the MAR/SAR sites during oxidative stress.
By contrast, Z-DEVD-FMK shows no effect on reducing cleavages within the AF9 non-SAR region. This indicates that the cleavages within the AF9 non-SAR region are neither dependent on caspase-3 nor CAD. Since H2O2 induces apoptosis and chromosome breaks in a caspase-3-dependent manner, these findings therefore strengthen the evidence that the cleavages within the AF9 non-SAR region are not mediated by H2O2-induced apoptosis.
In the current study, there are some limitations in using in silico prediction of MAR/SAR. The length and exact location of MAR/SAR could not be determined by using MRS-prediction. In order to study MAR/SAR in a more comprehensive way, biochemical isolation of MAR/SAR may be carried out simultaneously with in silico prediction in future works. This may be done by using a Southern blot-based SAR mapping assay [28]. Nevertheless, the positions of MRS-predicted MAR/SAR may serve as a guide for designing suitable probes to identify biochemically isolated MAR/SAR. As for the comparison of SAR region and non-SAR region, more non-SAR regions may be studied. These could help further elucidate the roles of MAR/SAR in stress-induced chromosome breakages and rearrangements.
Conclusions
Our results clearly demonstrate that oxidative stress-induced apoptosis results in the AF9 gene cleavages within the region that contains MAR/SAR. This implies that MAR/SAR may play an important role in defining the location of chromosomal cleavages during oxidative stress-induced apoptosis. In addition, the apoptotic nuclease CAD may be closely associated with MAR/SAR in mediating these oxidative stress-induced chromosomal cleavages. By investigating the role of MAR/SAR and its association with CAD, our findings provide deeper insights into the potential role of oxidative stress-induced apoptosis in mediating the chromosome rearrangements in nasopharyngeal epithelial cells.
Abbreviations
- NPC:
-
nasopharyngeal carcinoma
- CAD:
-
caspase-activated deoxyribonuclease
- MAR/SAR:
-
matrix association region/scaffold attachment region
- MRS:
-
MAR/SAR recognition signature
- H2O2 :
-
hydrogen peroxide
- PS:
-
phosphatidylserine
- MMP:
-
mitochondrial membrane potential
- IPCR:
-
inverse polymerase chain reaction
- BCR:
-
breakpoint cluster region
- MLL:
-
mixed lineage leukaemia
- ALL:
-
acute lymphoblastic leukaemia
- EBV:
-
Epstein–Barr virus
- ROS:
-
reactive oxygen species
- ICAD:
-
inhibitor of caspase-activated deoxyribonuclease
- CPT:
-
camptothecin
- VP16:
-
etoposide
- PI:
-
propidium iodide
- CI:
-
caspase inhibitor
References
Henle G, Henle W. Epstein–Barr virus-specific IgA serum antibodies as an outstanding feature of nasopharyngeal carcinoma. Int J Cancer. 1976;17:1–7.
Raab-Traub N. Epstein–Barr virus and nasopharyngeal carcinoma. Semin Cancer Biol. 1992;3:297–307.
Armstrong RW, Armstrong MJ, Yu MC, Henderson BE. Salted fish and inhalants as risk factors for nasopharyngeal carcinoma in Malaysian Chinese. Cancer Res. 1983;43:2967–70.
Fong LY, Ho JH, Huang DP. Preserved foods as possible cancer hazards: WA rats fed salted fish have mutagenic urine. Int J Cancer. 1979;23:542–6.
Huang DP, Ho JH, Saw D, Teoh TB. Carcinoma of the nasal and paranasal regions in rats fed Cantonese salted marine fish. IARC Sci Publ. 1978;20:315–28.
Yu MC, Nichols PW, Zou XN, Estes J, Henderson BE. Induction of malignant nasal cavity tumours in Wistar rats fed Chinese salted fish. Br J Cancer. 1989;60:198–201.
Fong YY, Chan WC. Nitrate, nitrite, dimethylnitrosamine and N-nitrosopyrrolidine in some Chinese food products. Food Cosmet Toxicol. 1977;15:143–5.
Poirier S, Ohshima H, de The G, Hubert A, Bourgade MC, Bartsch H. Volatile nitrosamine levels in common foods from Tunisia, south China and Greenland, high-risk areas for nasopharyngeal carcinoma (NPC). Int J Cancer. 1987;39:293–6.
Cheng YJ, Hildesheim A, Hsu MM, Chen IH, Brinton LA, Levine PH, et al. Cigarette smoking, alcohol consumption and risk of nasopharyngeal carcinoma in Taiwan. Cancer Causes Control. 1999;10:201–7.
Armstrong RW, Imrey PB, Lye MS, Armstrong MJ, Yu MC, Sani S. Nasopharyngeal carcinoma in Malaysian Chinese: occupational exposures to particles, formaldehyde and heat. Int J Epidemiol. 2000;29:991–8.
Vaughan TL, Stewart PA, Teschke K, Lynch CF, Swanson GM, Lyon JL, et al. Occupational exposure to formaldehyde and wood dust and nasopharyngeal carcinoma. Occup Environ Med. 2000;57:376–84.
Hildesheim A, Dosemeci M, Chan CC, Chen CJ, Cheng YJ, Hsu MM, et al. Occupational exposure to wood, formaldehyde, and solvents and risk of nasopharyngeal carcinoma. Cancer Epidemiol Biomark Prev. 2001;10:1145.
Hung SH, Chen PY, Lin HC, Ting J, Chung SD. Association of rhinosinusitis with nasopharyngeal carcinoma: a population-based study. Laryngoscope. 2014;124:1515–20.
Tsou YA, Lin CC, Tai CJ, Tsai MH, Tsai TC, Chen CM. Chronic rhinosinusitis and the risk of nasopharyngeal cancer in a Taiwanese health study. Am J Rhinol Allergy. 2014;28:168–72.
Ahotupa M, Bussacchini-Griot V, Bereziat JC, Camus AM, Bartsch H. Rapid oxidative stress induced by N-nitrosamines. Biochem Biophys Res Commun. 1987;146:1047–54.
Carnevali S, Petruzzelli S, Longoni B, Vanacore R, Barale R, Cipollini M, et al. Cigarette smoke extract induces oxidative stress and apoptosis in human lung fibroblasts. Am J Physiol Lung Cell Mol Physiol. 2003;284:L955–63.
Federico A, Morgillo F, Tuccillo C, Ciardiello F, Loguercio C. Chronic inflammation and oxidative stress in human carcinogenesis. Int J Cancer. 2007;121:2381–6.
Kum C, Kiral F, Sekkin S, Seyrek K, Boyacioglu M. Effects of xylene and formaldehyde inhalations on oxidative stress in adult and developing rats livers. Exp Anim. 2007;56:35–42.
Gruhne B, Sompallae R, Marescotti D, Kamranvar SA, Gastaldello S, Masucci MG. The Epstein–Barr virus nuclear antigen-1 promotes genomic instability via induction of reactive oxygen species. Proc Natl Acad Sci USA. 2009;106:2313–8.
Pylkkanen L, Stockmann-Juvala H, Alenius H, Husgafvel-Pursiainen K, Savolainen K. Wood dusts induce the production of reactive oxygen species and caspase-3 activity in human bronchial epithelial cells. Toxicology. 2009;262:265–70.
Huang SY, Fang CY, Tsai CH, Chang Y, Takada K, Hsuc TY, et al. N-methyl-N′-nitro-N-nitrosoguanidine induces and cooperates with 12-O-tetradecanoylphorbol-1,3-acetate/sodium butyrate to enhance Epstein–Barr virus reactivation and genome instability in nasopharyngeal carcinoma cells. Chem Biol Interact. 2014;188:623–34.
Wu CC, Fang CY, Cheng YJ, Hsu HY, Chou SP, Huang SY, et al. Inhibition of Epstein–Barr virus reactivation by the flavonoid apigenin. J Biomed Sci. 2017;24:2.
Sallmyr A, Fan J, Rassool FV. Genomic instability in myeloid malignancies: increased reactive oxygen species (ROS), DNA double strand breaks (DSBs) and error-prone repair. Cancer Lett. 2008;270:1–9.
Ma N, Kawanishi M, Hiraku Y, Murata M, Huang GW, Huang Y, et al. Reactive nitrogen species-dependent DNA damage in EBV-associated nasopharyngeal carcinoma: the relation to STAT3 activation and EGFR expression. Int J Cancer. 2008;122:2517–25.
Zhao F, Wang Q. The protective effect of peroxiredoxin II on oxidative stress induced apoptosis in pancreatic beta-cells. Cell Biosci. 2012;2:22.
Tan SN, Sim SP, Khoo AS. Potential role of oxidative stress-induced apoptosis in mediating chromosomal rearrangements in nasopharyngeal carcinoma. Cell Biosci. 2016;6:35.
Strick R, Zhang Y, Emmanuel N, Strissel PL. Common chromatin structures at breakpoint cluster regions may lead to chromosomal translocations found in chronic and acute leukemias. Hum Genet. 2006;119:479–95.
Strissel PL, Strick R, Tomek RJ, Roe BA, Rowley JD, Zeleznik L. DNA structural properties of AF9 are similar to MLL and could act as recombination hot spots resulting in MLL/AF9 translocations and leukemogenesis. Hum Mol Genet. 2000;9:1671–9.
Broeker PL, Super HG, Thirman MJ, Pomykala H, Yonebayashi Y, Tanabe S, et al. Distribution of 11q23 breakpoints within the MLL breakpoint cluster region in de novo acute leukemia and in treatment-related acute myeloid leukemia: correlation with scaffold attachment regions and topoisomerase II consensus binding sites. Blood. 1996;87:1912–22.
Oberhammer F, Wilson JW, Dive C, Morris ID, Hickman JA, Wakeling AE, et al. Apoptotic death in epithelial cells: cleavage of DNA to 300 and/or 50 kb fragments prior to or in the absence of internucleosomal fragmentation. EMBO J. 1993;12:3679–84.
Berezney R, Mortillaro MJ, Ma H, Wei X, Samarabandu J. The nuclear matrix: a structural milieu for genomic function. Int Rev Cytol. 1995;162A:1–65.
Shao JY, Wang HY, Huang XM, Feng QS, Huang P, Feng BJ, et al. Genome-wide allelotype analysis of sporadic primary nasopharyngeal carcinoma from southern China. Int J Oncol. 2000;17:1267–75.
van Drunen CM, Sewalt RG, Oosterling RW, Weisbeek PJ, Smeekens SC, van Driel R. A bipartite sequence element associated with matrix/scaffold attachment regions. Nucleic Acids Res. 1999;27:2924–30.
Jeffs AR, Benjes SM, Smith TL, Sowerby SJ, Morris CM. The BCR gene recombines preferentially with Alu elements in complex BCR-ABL translocations of chronic myeloid leukaemia. Hum Mol Genet. 1998;7:767–76.
Cynthia PN, Sim SP. Etoposide-induced apoptosis results in chromosome breaks within the AF9 gene: its implication in chromosome rearrangement in leukaemia. Adv Biosci Biotechnol. 2012;3:686–94.
Lo KW, Huang DP. Genetic and epigenetic changes in nasopharyngeal carcinoma. Semin Cancer Biol. 2002;12:451–62.
Liu MJ, Wang XY, Peng Y, Shen SR, Li GY. Egr-1 regulates the transcription of NGX6 gene through a Sp1/Egr-1 overlapping site in the promoter. BMC Mol Biol. 2014;15:14.
Wiseman H, Halliwell B. Damage to DNA by reactive oxygen and nitrogen species: role in inflammatory disease and progression to cancer. Biochem J. 1996;313(Pt 1):17–29.
Kim DK, Cho ES, Um HD. Caspase-dependent and -independent events in apoptosis induced by hydrogen peroxide. Exp Cell Res. 2000;257:82–8.
Kerr JF, Wyllie AH, Currie AR. Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. Br J Cancer. 1972;26:239–57.
Vaughan AT, Betti CJ, Villalobos MJ. Surviving apoptosis. Apoptosis. 2002;7:173–7.
Betti CJ, Villalobos MJ, Diaz MO, Vaughan AT. Apoptotic triggers initiate translocations within the MLL gene involving the nonhomologous end joining repair system. Cancer Res. 2001;61:4550–5.
Gohring F, Schwab BL, Nicotera P, Leist M, Fackelmayer FO. The novel SAR-binding domain of scaffold attachment factor A (SAF-A) is a target in apoptotic nuclear breakdown. EMBO J. 1997;16:7361–71.
Bode J, Benham C, Ernst E, Knopp A, Marschalek R, Strick R, et al. Fatal connections: when DNA ends meet on the nuclear matrix. J Cell Biochem. 2000;73(Suppl 35):3–22.
Legault J, Tremblay A, Mirault ME. Preferential localization of DNA damage induced by depurination and bleomycin in a plasmid containing a scaffold-associated region. Biochem Cell Biol. 1997;75:369–75.
Yee PH, Sim SP. High cell density and latent membrane protein 1 expression induce cleavage of the mixed lineage leukemia gene at 11q23 in nasopharyngeal carcinoma cell line. J Biomed Sci. 2010;17:77.
Swansbury GJ, Slater R, Bain BJ, Moorman AV, Secker-Walker LM. Hematological malignancies with t(9;11)(p21–22;q23)—a laboratory and clinical study of 125 cases. European 11q23 Workshop participants. Leukemia. 1998;12:792–800.
Donev R, Horton R, Beck S, Doneva T, Vatcheva R, Bowen WR, et al. Recruitment of heterogeneous nuclear ribonucleoprotein A1 in vivo to the LMP/TAP region of the major histocompatibility complex. J Biol Chem. 2003;278:5214–26.
Ostermeier GC, Liu Z, Martins RP, Bharadwaj RR, Ellis J, Draghici S, et al. Nuclear matrix association of the human b-globin locus utilizing a novel approach to quantitative real-time PCR. Nucleic Acids Res. 2003;31:3257–66.
Thilmony R, Guttman ME, Lin JW, Blechl AE. The wheat HMW-glutenin 1Dy10 gene promoter controls endosperm expression in Brachypodium distachyon. GM Crops Food. 2014;5:36–43.
Lechardeur D, Xu M, Lukacs GL. Contrasting nuclear dynamics of the caspase-activated DNase (CAD) in dividing and apoptotic cells. J Cell Biol. 2004;167:851–62.
Mimori K, Druck T, Inoue H, Alder H, Berk L, Mori M, et al. Cancer-specific chromosome alterations in the constitutive fragile region FRA3B. Proc Natl Acad Sci USA. 1999;96:7456–61.
Arlt MF, Miller DE, Beer DG, Glover TW. Molecular characterization of FRAXB and comparative common fragile site instability in cancer cells. Genes Chromosom Cancer. 2002;33:82–92.
Enari M, Sakahira H, Yokoyama H, Okawa K, Iwamatsu A, Nagata S. A caspase-activated DNase that degrades DNA during apoptosis, and its inhibitor ICAD. Nature. 1998;391:43–50.
Sakahira H, Enari M, Nagata S. Cleavage of CAD inhibitor in CAD activation and DNA degradation during apoptosis. Nature. 1998;391:96–9.
Wolf BB, Schuler M, Echeverri F, Green DR. Caspase-3 is the primary activator of apoptotic DNA fragmentation via DNA fragmentation factor-45/inhibitor of caspase-activated DNase inactivation. J Biol Chem. 1999;274:30651–6.
Boon SS, Sim SP. Inhibitor of caspase-activated DNase expression enhances caspase-activated DNase expression and inhibits oxidative stress-induced chromosome breaks at the mixed lineage leukaemia gene in nasopharyngeal carcinoma cells. Cancer Cell Int. 2015;15:54.
Mirkovitch J, Gasser SM, Laemmli UK. Scaffold attachment of DNA loops in metaphase chromosomes. J Mol Biol. 1988;200:101–9.
Cockerill PN, Garrard WT. Chromosomal loop anchorage of the kappa immunoglobulin gene occurs next to the enhancer in a region containing topoisomerase II sites. Cell. 1986;44:273–82.
Authors’ contributions
SNT carried out the in silico prediction of MAR/SARs and repeat elements, conducted all experimentation, constructed the genomic maps, performed statistical analyses and wrote the manuscript. SPS was the principal investigator of the project, conceived the study, participated in the experimental design, interpreted the data and edited the manuscript. ASBK provided help in the intellectual formulation of the project. All authors read and approved the final manuscript.
Acknowledgements
We would like to thank the Director-General of Health Malaysia for his permission to publish this article and the Director of Institute for Medical Research Malaysia for her support. We would like to acknowledge Prof. Tsao Sai Wah (The University of Hong Kong, Hong Kong, China) and Prof. Lo Kwok Wai (The Chinese University of Hong Kong, Hong Kong, China) for providing the normal nasopharyngeal epithelial cell line (NP69) and NPC cell line (HK1). We are deeply grateful to Ms. Chin Mei Yieng (formerly from Universiti Malaysia Sarawak) for intellectual discussion. We express our gratitude to Universiti Malaysia Sarawak for all infrastructure support. This is also another project by the members of Malaysian Nasopharyngeal Carcinoma Group.
Competing interests
The authors declare that they have no competing interests.
Availability of data and materials
All data generated or analysed during this study are included in this published article.
Consent for publication
Consent granted from the funding agency.
Ethics approval and consent to participate
Not applicable.
Funding
This project was funded by Ministry of Health Malaysia (Grant Number: 06-065).
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Author information
Authors and Affiliations
Corresponding author
Additional files
Additional file 1.
Description of exons and introns in the AF9 gene.
Additional file 2.
DNA manipulation steps in preparation for nested IPCR.
Additional file 3.
DNA sequencing data.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Tan, SN., Sim, SP. & Khoo, A.S.B. Matrix association region/scaffold attachment region (MAR/SAR) sequence: its vital role in mediating chromosome breakages in nasopharyngeal epithelial cells via oxidative stress-induced apoptosis. BMC Molecular Biol 19, 15 (2018). https://doi.org/10.1186/s12867-018-0116-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12867-018-0116-5