Abstract
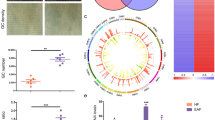
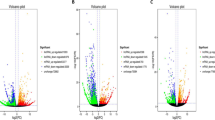
The activation of primordial follicles is critical to ovarian follicle development, which directly influences female fertility and reproductive life span. Several studies have suggested a role for long noncoding RNAs (lncRNAs) in ovarian function. However, the precise involvement of lncRNAs in the initiation of primordial follicles is still unknown.Here, an in vitro culture model was used to investigate the roles of lncRNAs in primordial follicle activation.We found that primordial follicles in day 3 mouse ovaries were activated after culturing for 8 days in vitro, as indicated by ovarian morphology changes, increases in primary follicle number, and downregulation of mammalian Sterile 20-like kinase messenger RNA (mRNA) and upregulation of growth differentiation factor 9 mRNA. We next examined lncRNA expression profiles by RNA sequencing at the transcriptome level and found that among 60 078 lncRNAs, 6541 lncRNA were upregulated and 2135 lncRNA were downregulated in 3-day ovaries cultured for 8 days in vitro compared with ovaries from day 3 mice. We also found that 4171 mRNAs were upregulated and 1795 were downregulated in the cultured ovaries. Gene ontology and pathway analyses showed that the functions of differentially expressed lncRNA targets and mRNAs were closely linked with many processes and pathways related to ovary development, including cell proliferation and differentiation, developmental processes, and other signaling transduction pathways. Additionally, many novel identified lncRNAs showed inducible expression, suggesting that these lncRNAs may be good candidates for investigating mouse primordial follicle activation. This study provides a foundation for further exploring lncRNA-related mechanisms in the initiation of mouse primordial follicles.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Ernst EH, Grondahl ML, Grund S, et al. Dormancy and activation of human oocytes from primordial and primary follicles: molecular clues to oocyte regulation. Hum Reprod. 2017;32(8):1684–1700.
Sun X, Su Y, He Y, et al. New strategy for in vitro activation of primordial follicles with mTOR and PI3K stimulators. Cell Cycle. 2015;14(5):721–731.
Kezele P, Nilsson E, Skinner MK. Cell-cell interactions in primordial follicle assembly and development. Front Biosci. 2002;7:d1990–d1996.
Hu X, Sood AK, Dang CV, Zhang L. The role of long noncoding RNAs in cancer: the dark matter matters. Curr Opin Genet Dev. 2017;48:8–15.
Svoboda P. Long and small noncoding RNAs during oocyte-toembryo transition in mammals. Biochem Soc Trans. 2017;45(5):1117–1124.
Su Y, Wu H, Pavlosky A, et al. Regulatory non-coding RNA: new instruments in the orchestration of cell death. Cell Death Dis. 2016;7(8):e2333.
Hart RP, Goff LA. Long noncoding RNAs: central to nervous system development. Int J Dev Neurosci. 2016;55:109–116.
Nachtergaele S, He C. The emerging biology of RNA posttranscriptional modifications. RNA Biol. 2017;14(2):156–163.
Dykes IM, Emanueli C. Transcriptional and post-transcriptional gene regulation by long non-coding RNA. Genomics Proteomics Bioinformatics. 2017;15(3):177–186.
Wei JW, Huang K, Yang C, Kang CS. Non-coding RNAs as regulators in epigenetics (Review). Oncol Rep. 2017;37(1):3–9.
Wang KC, Chang HY. Molecular mechanisms of long noncoding RNAs. Mol Cell. 2011;43(6):904–914.
Qiu JJ, Wang Y, Ding JX, Jin HY, Yang G, Hua KQ. The long non-coding RNA HOTAIR promotes the proliferation of serous ovarian cancer cells through the regulation of cell cycle arrest and apoptosis. Exp Cell Res. 2015;333(2):238–248.
Gao Y, Meng H, Liu S, et al. LncRNA-HOST2 regulates cell biological behaviors in epithelial ovarian cancer through a mechanism involving microRNA let-7b. Hum Mol Genet. 2015;24(3):841–852.
Qiu JJ, Lin YY, Ding JX, Feng WW, Jin HY, Hua KQ. Long noncoding RNA ANRIL predicts poor prognosis and promotes invasion/ metastasis in serous ovarian cancer. Int J Oncol. 2015;46(6):2497–2505.
Gao Q, Ren H, Chen M, et al. Long non-coding RNAs regulate effects of beta-crystallin B2 on mouse ovary development. Mol Med Rep. 2016;14(5):4223–4231.
Huang X, Hao C, Bao H, Wang M, Dai H. Aberrant expression of long noncoding RNAs in cumulus cells isolated from PCOS patients. J Assist Reprod Genet. 2016;33(1):111–121.
Ren J, Du X, Zeng T, et al. Divergently expressed gene identification and interaction prediction of long noncoding RNA and mRNA involved in duck reproduction. Anim Reprod Sci. 2017;185:8–17.
Xiang C, Li J, Hu L, et al. Hippo signaling pathway reveals a spatio-temporal correlation with the size of primordial follicle pool in mice. Cell Physiol Biochem. 2015;35(3):957–968.
Association APOEaVM. 2000 Report of the AVMA panel on Euthanasia. J Am Vet Med Assoc. 2001;218(5):669–696.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12(4):357–360.
Pertea M, Pertea GM, Antonescu CM, Chang TC, Mendell JT, Salzberg SL. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol. 2015;33(3):290–295.
Kong L, Zhang Y, Ye ZQ, et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 2007;35(Web Server issue):W345–W349.
Sun L, Luo H, Bu D, et al. Utilizing sequence intrinsic composition to classify protein-coding and long non-coding transcripts. Nucleic Acids Res. 2013;41(17):e166.
Punta M, Coggill PC, Eberhardt RY, et al. The Pfam protein families database. Nucleic Acids Res. 2012;40(Database issue):D290–D301.
Trapnell C, Roberts A, Goff L, et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc. 2012;7(3):562–578.
Wang L, Feng Z, Wang X, Wang X, Zhang X. DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics. 2010;26(1):136–138.
Tafer H, Hofacker IL. RNAplex: a fast tool for RNA-RNA interaction search. Bioinformatics. 2008;24(22):2657–2663.
Asadi-Azarbaijani B, Santos RR, Jahnukainen K, et al. Developmental effects of imatinib mesylate on follicle assembly and early activation of primordial follicle pool in postnatal rat ovary. Reprod Biol. 2017;17(1):25–33.
Li J, Kawamura K, Cheng Y, et al. Activation of dormant ovarian follicles to generate mature eggs. Proc Natl Acad Sci U S A. 2010;107(22):10280–10284.
Wang H, Fu Z, Dai C, et al. LncRNAs expression profiling in normal ovary, benign ovarian cyst and malignant epithelial ovarian cancer. Sci Rep. 2016;6:38983.
Liu YD, Li Y, Feng SX, et al. Long noncoding RNAs: potential regulators involved in the pathogenesis of polycystic ovary syndrome. Endocrinology. 2017;158(11):3890–3899.
O’brien MJ, Pendola JK, Eppig JJ. A revised protocol for in vitro development of mouse oocytes from primordial follicles dramatically improves their developmental competence. Biol Reprod. 2003;68(5):1682–1686.
Braw-Tal R, Yossefi S. Studies in vivo and in vitro on the initiation of follicle growth in the bovine ovary. J Reprod Fertil. 1997;109(1):165–171.
Wandji SA, Srsen V, Nathanielsz PW, Eppig JJ, Fortune JE. Initiation of growth of baboon primordial follicles in vitro. Hum Reprod. 1997;12(9):1993–2001.
Hovatta O, Wright C, Krausz T, Hardy K, Winston RM. Human primordial, primary and secondary ovarian follicles in long-term culture: effect of partial isolation. Hum Reprod. 1999;14(10):2519–2524.
Kedem A, Fisch B, Garor R, et al. Growth differentiating factor 9 (GDF9) and bone morphogenetic protein 15 both activate development of human primordial follicles in vitro, with seemingly more beneficial effects of GDF9. J Clin Endocrinol Metab. 2011;96(8):E1246–E1254.
Martins FS, Celestino JJ, Saraiva MV, et al. Growth and differentiation factor-9 stimulates activation of goat primordial follicles in vitro and their progression to secondary follicles. Reprod Fertil Dev. 2008;20(8):916–924.
Xiong Y, Liu T, Wang S, Chi H, Chen C, Zheng J. Cyclophosphamide promotes the proliferation inhibition of mouse ovarian granulosa cells and premature ovarian failure by activating the lncRNA-Meg3-p53-p66Shc pathway. Gene. 2017;596:1–8.
Bassett AR, Akhtar A, Barlow DP, et al. Considerations when investigating lncRNA function in vivo. 2014;3:e03058.
Mihm M, Bleach EC. Endocrine regulation of ovarian antral follicle development in cattle. Anim Reprod Sci. 2003;78(3–4):217–237.
Wang LJ, Brannstrom M, Cui KH, et al. Localisation of mRNA for interleukin-1 receptor and interleukin-1 receptor antagonist in the rat ovary. J Endocrinol. 1997;152(1):11–17.
Fuller EA, Sominsky L, Sutherland JM, et al. Neonatal immune activation depletes the ovarian follicle reserve and alters ovarian acute inflammatory mediators in neonatal rats. Biol Reprod. 2017;97(5):719–730.
Kezele PR, Ague JM, Nilsson E, Skinner MK. Alterations in the ovarian transcriptome during primordial follicle assembly and development. Biol Reprod. 2005;72(1):241–255.
Pepling ME. Follicular assembly: mechanisms of action. Reproduction. 2012;143(2):139–149.
Tagler D, Makanji Y, Tu T, et al. Promoting extracellular matrix remodeling via ascorbic acid enhances the survival of primary ovarian follicles encapsulated in alginate hydrogels. Biotechnol Bioeng. 2014;111(7):1417–1429.
Skinner MK. Regulation of primordial follicle assembly and development. Hum Reprod Update. 2005;11(5):461–471.
Du XY, Huang J, Xu LQ, et al. The proto-oncogene c-src is involved in primordial follicle activation through the PI3K, PKC and MAPK signaling pathways. Reprod Biol Endocrinol. 2012;10:58.
Li-Ping Z, Da-Lei Z, Jian H, et al. Proto-oncogene c-erbB2 initiates rat primordial follicle growth via PKC and MAPK pathways. Reprod Biol Endocrinol. 2010;8:66.
Chen CL, Fu XF, Wang LQ, et al. Primordial follicle assembly was regulated by Notch signaling pathway in the mice. Mol Biol Rep. 2014;41(3):1891–1899.
Vanorny DA, Prasasya RD, Chalpe AJ, Kilen SM, Mayo KE. Notch signaling regulates ovarian follicle formation and coordinates follicular growth. Mol Endocrinol. 2014;28(4):499–511.
Hussein MR. Apoptosis in the ovary: molecular mechanisms. Hum Reprod Update. 2005;11(2):162–177.
Marcinkiewicz JL, Balchak SK, Morrison LJ. The involvement of tumor necrosis factor-alpha (TNF) as an intraovarian regulator of oocyte apoptosis in the neonatal rat. Front Biosci. 2002;7:d1997–d2005.
Chen YC, Chang HM, Cheng JC, Tsai HD, Wu CH, Leung PC. Transforming growth factor-beta1 up-regulates connexin43 expression in human granulosa cells. Hum Reprod. 2015;30(9):2190–2201.
Cheng JC, Chang HM, Fang L, Sun YP, Leung PC. TGF-beta1 up-regulates connective tissue growth factor expression in human granulosa cells through Smad and ERK1/2 signaling pathways. PLoS One. 2015;10(5):e0126532.
Nomura M, Sakamoto R, Morinaga H, Wang L, Mukasa C, Takayanagi R. Activin stimulates CYP19A gene expression in human ovarian granulosa cell-like KGN cells via the Smad2 signaling pathway. Biochem Biophys Res Commun. 2013;436(3):443–448.
Nilsson EE, Skinner MK. Bone morphogenetic protein-4 acts as an ovarian follicle survival factor and promotes primordial follicle development. Biol Reprod. 2003;69(4):1265–1272.
Nilsson EE, Schindler R, Savenkova MI, Skinner MK. Inhibitory actions of anti-Mullerian hormone (AMH) on ovarian primordial follicle assembly. PLoS One. 2011;6(5):e20087.
Liu Z, Ren YA, Pangas SA, et al. FOXO1/3 and PTEN depletion in granulosa cells promotes ovarian granulosa cell tumor development. Mol Endocrinol. 2015;29(7):1006–1024.
Lan ZJ, Krause MS, Redding SD, et al. Selective deletion of Pten in theca-interstitial cells leads to androgen excess and ovarian dysfunction in mice. Mol Cell Endocrinol. 2017;444:26–37.
Acknowledgments
The authors thank Edanz Group (www.edanzediting.com/ac) for editing a draft of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Zheng, L., Luo, R., Su, T. et al. Differentially Expressed lncRNAs After the Activation of Primordial Follicles in Mouse. Reproductive Sciences 26, 1094–1104 (2019). https://doi.org/10.1177/1933719118805869
Published:
Issue Date:
DOI: https://doi.org/10.1177/1933719118805869